Table of Contents |
guest 2026-03-14 |
Use Sample Manager with LabKey Server
Use Sample Manager with Studies
LabKey ELN
ELN: Frequently Asked Questions
LabKey LIMS
LIMS: Downloadable Templates
LIMS: Samples
Print Labels with BarTender
LIMS: Assay Data
LIMS: Charts
LIMS: Storage Management
LIMS: Workflow
Biologics LIMS
Introduction to LabKey Biologics
Release Notes: Biologics
Biologics: Navigate
Biologics: Projects and Folders
Biologics: Bioregistry
Create Registry Sources
Register Nucleotide Sequences
Register Protein Sequences
Register Leaders, Linkers, and Tags
Vectors, Constructs, Cell Lines, and Expression Systems
Registry Reclassification
Biologics: Terminology
Protein Sequence Annotations
CoreAb Sequence Classification
Biologics: Chain and Structure Formats
Molecules, Sets, and Molecular Species
Register Molecules
Molecular Physical Property Calculator
Compounds and SMILES Lookups
Entity Lineage
Customize the Bioregistry
Bulk Registration of Entities
Use the Registry API
Biologics: Plates
Biologics: Assay Data
Biologics: Specialty Assays
Biologics: Assay Integration
Biologics: Upload Assay Data
Biologics: Assay Batches and QC
Biologics: Media Registration
Managing Ingredients and Raw Materials
Registering Mixtures (Recipes)
Registering Batches
Biologics Administration
Biologics: Detail Pages and Entry Forms
Biologics: Protect Sequence Fields
Manage Notebook Tags
Biologics Admin: URL Properties
Sample Manager
Sample Manager Resources
LabKey Sample Manager is a sample management application designed to be easy-to-use while providing powerful lab sample tracking and workflow features.Using Sample Manager with LabKey Server
Premium Editions of LabKey Server include the option to use the Sample Manager application within a project and integrate with LabKey Studies and other resources.Using Electronic Lab Notebooks (ELN) in Sample Manager
The Professional Edition of LabKey Sample Manager includes a user-friendly ELN (Electronic Lab Notebook) designed to help scientists efficiently document their experiments and collaborate. This data-connected ELN is seamlessly integrated with other laboratory data in the application, including lab samples, assay data and other registered data.Learn more in this section:Use Sample Manager with LabKey Server
- Sample Manager in a Project
- Using Sample Manager in a Folder
- Export and Import of Sample Manager
- Sample Manager User Interface
- Shared Sample Types and Source Types
- Sample Management Users and Permissions
- Use Naming Prefixes
- Configure Storage Location Options
- Storage Management API
Sample Manager in a Project
Sample Manager is designed to be contained in a project on LabKey Server. This provides advantages in scoping permissions specific to sample management. For instance, many studies in other projects may need to be linked to sample data, but a single group doing the actual sample administration and storage management for all studies would have elevated permissions on the Sample Manager folder instead of needing them on all studies.Resources including Sample Types and Assay Designs that should be usable in both the Sample Manager project and other containers on the server should be placed in the Shared project.- Create a new project, selecting the folder type Sample Manager.
- Configure permissions now, or leave it visible to only your own user account until you have set things up, then open permissions to your users.
Using Sample Manager in a Folder
Sample Manager is designed to be rooted at the top-level project level of LabKey Server. In some limited use cases, it may work to use Sample Manager in a subfolder, however not all features are supported. If you do not expect to share any Sample Types, Assays, or other data with other containers, and will only use a single set of container-based permissions, using Sample Manager in a subfolder may support integration with LabKey Server features and reports. Note that you cannot use Sample Manager Folders unless it is rooted at the project level.If you choose either of the following subfolder options, you will be able to use the product selection menu to switch between Sample Manager and the traditional LabKey Server interface.Folder of "Sample Manager" Type
You can create a new folder and select the "Sample Manager" folder type. The application will launch every time you navigate to this folder, as if it were at the project level.Enable "SampleManagement" Module
If you want to use Sample Manager resources in a folder, but do not want to auto-navigate to the application, you can create a folder of another type (such as Study) and then enable the "SampleManagement" module via > Folder > Management > Folder Type.You will not be navigated directly to the application in this folder, but can still reach it by editing the URL to replace "project-begin.view?" with "sampleManager-app.view?".Export and Import of Sample Manager
Sample Manager projects and folders do not support full fidelity migration to other containers using folder export and reimport available in the LabKey Server interface. Some of the contents of Sample Manager can be migrated in this way, in some cases requiring manually migrating some data in a specific order. Others, including but not limited to Workflow and Notebook contents, cannot be moved in this way.If you need to migrate any Sample Manager data or structures between containers, please work with your Account Manager to identify whether this is feasible in your scenario. Note that it is never possible to "promote" a folder to the project level within LabKey Server.Sample Manager User Interface
When using Sample Manager with a Premium Edition of LabKey Server, you may see different features and options when viewing the "same" data in the two different interfaces. A few examples of differences are listed here, but this is not a comprehensive list.Learn about switching between the interfaces in this topic:Grid Charts are Available
When using Sample Manager with a Premium Edition of LabKey Server, you can define and use chart visualizations within the application. Learn more about charts in this topic for Biologics LIMS:Conditional Formatting is not Available
Conditional formatting of values is not supported in Sample Manager, though you will see the controls for adding such formats in the field editor.In order to use conditional formatting, you will need to define the formats and view your data in the LabKey Server Interface.Shared Sample Types and Source Types
Any Sample Types and Sources defined in the /Shared project will also be available for use in LabKey Sample Manager. You will see them listed on the menu and dashboards alongside local definitions.From within the Sample Manager application, users editing (or deleting) a shared Sample Type or Source Type will see a banner indicating that changes could affect other folders.Sample Management Users and Permission Roles
An administrator in the Sample Manager or LabKey Biologics applications can access a grid of active users using the Administration option on the user avatar menu. You'll see all the users with any access to the container in which the application is enabled.Sample Manager and Biologics both use a subset of LabKey's role-based permissions. The "Reader", "Editor", "Editor without Delete", "Folder Administrator", and "Project Administrator" roles all map to the corresponding container-scoped roles. Learn more about those permissions in this topic: permissionLevels. In addition, "Storage Editor" and "Storage Designer" roles are added to the Sample Manager and Biologics applications and described below.Reader, Editor, and Editor without Delete
- When a user is granted the "Reader", "Editor", or "Editor without Delete" role, either in the LabKey Server folder permissions interface or the Sample Manager > Administration > Permissions tab, they will have that role in both interfaces.
- Assigning a user to a role in either place, or revoking that role in either place will apply to both the Sample Manager and LabKey Server resources in that container.
- If Sample Manager is defined in a folder, and that folder inherits permissions from a parent container, you will not be able to change role assignments within the application.
Administrator Roles
In the stand-alone Sample Manager or application, a single "Administrator" role is provided that maps to the site role "Application Admin". This also means that any "Application Admin" on the site will appear in Sample Manager as an Administrator. The Sample Manager Documentation means these site-level roles when it identifies tasks available to an "Administrator".When using Sample Manager within LabKey Server, administrator permissions work differently. Users with the "Folder Administrator" or "Project Administrator" role on the project or folder containing Sample Manager have those roles carry into the application. These roles may also be assigned from within the application (by users with sufficient permission). The difference between these two roles in Sample Manager is that a Project Administrator can add new user accounts, but a Folder Administrator cannot. Both admin roles can assign the various roles to existing users and perform other administrative tasks, with some exceptions listed below.As in any LabKey Server Installation, "Site Admin" and "Application Admin" roles are assigned at the site level and grant administrative permission to Sample Manager, but are not shown in the Administrator listing of the application.Most actions are available to any administrator role in Sample Manager with some exceptions including:- Only "Site Admin" and "Application Admin" users can manage sample statuses.
Specific Roles for Storage Management
Two roles specific to managing storage of physical samples let Sample Manager and LabKey Biologics administrators independently grant users control over the physical storage details for samples.- Storage Editor: confers the ability to read, add, edit, and delete data related to items in storage, picklists, and jobs.
- Storage Designer: confers the ability to read, add, edit, and delete data related to storage locations and storage units.
Workflow Editor Role
The Workflow Editor role lets a user create and edit sample picklists as well as workflow jobs and tasks, though not workflow templates. Workflow editors also require the "Reader" role or higher in order to accomplish these tasks.Administrators can also perform the tasks available to users with the Workflow Editor role.Use Naming Prefixes
You can apply a container-specific naming prefix that will be added to naming patterns for all Sample Types and Source Types to assist integration of data from multiple locations while maintaining a clear association with the original source of that data.When you have more than one Sample Manager project or folder, or are using both Sample Manager and LabKey Biologics on the same LabKey Server, it can be helpful to assign unique prefixes to each container.Learn more about using prefixes here:Configure Storage Location Options
Freezers and other storage systems can be assigned physical locations to make it easier for users to find them in a larger institution or campus. Learn more about defining and using storage locations in this topic: Note that this is different from configuring location hierarchies within freezers or other storage systems.Storage Management API
In situations where you have a large number of storage systems to define, it may be more convenient to define them programmatically instead of using the UI. Learn more in the API documentation:- Java Storage API
- JavaScript Storage API
- Python Storage API
- Rlabkey Package API: See labkey.storage.create/delete/update.
Related Topics
Use Sample Manager with Studies
Include Participant and Visit Information in Sample Types
Fields of specific types can be included in a Sample Type, providing participant (Subject/Participant type) and visit information (either VisitDate or VisitID/VisitLabel depending on the timepoint type of the target study). These fields can be defined either:- On the type of the samples you want to link
- On the type of a parent sample of the samples you want to link.
Link Samples to Studies from Sample Manager
When the Sample Manager application is used on a Premium Edition of LabKey Server, you gain the ability to link samples to studies directly from within the application.From a Samples grid, select the samples of interest using the checkboxes, then select Edit > Link to LabKey Study.- You must have the necessary alignment fields defined.
- The name of your Sample Type cannot be the same as the name of any table or dataset in the study you are linking to.
Related Topics
LabKey ELN
Related Topics
- Manage Notebook Tags (Available in Biologics LIMS only)
- ELN: Frequently Asked Questions
ELN: Frequently Asked Questions
How can I use Notebooks for my work?
Notebooks are available in the Biologics and Sample Manager applications.- Already using Biologics? You'll already have Notebooks, provided you are on a current version.
- Already using Sample Manager? You'll need to be using the Professional Edition of Sample Manager, or the Enterprise Edition of LabKey Server to access Notebooks.
- Not using either of the above? Please contact us to learn more.
Who can see my Notebooks?
Every user with "Read" access to your folder can also see Notebooks created there.What can I reference from a Notebook?
You can reference anything in your application, including data, experiments, specific samples, and other notebooks. Use a direct reference selector to place a color coded reference directly in your Notebook text. Add a single reference at a time, or add multiple references in bulk.The combined list of all elements referenced from a notebook is maintained in an Overview panel.Learn more in this topic: Add a ReferenceHow are Notebooks locked and protected once signed?
After the notebook has been approved we create a signed snapshot. The snapshot will include all notebook data including:- Notebook metadata and text
- Any data referenced in the notebook (samples, entities, experiments, assay runs, etc.)
- Attached files
What happens to my Notebooks when I upgrade?
- Once you create a Notebook, it will be preserved (and unaltered) by upgrades of LabKey.
What are team templates?
"Team templates" is a term that was in use in earlier versions for templates that are now called Shared Templates.When you view the Notebook Templates Dashboard, all the templates you created are listed on the Your Templates tab. Templates created by yourself or other users and shared (i.e. team templates) are listed on the Shared Templates tab.Related Topics
LabKey LIMS
- Generate customizable reports with ease.
- Add automation with transformation scripts for assay data.
- Customize the downloadable templates for data structures like samples, sources, and assays.
LIMS: Downloadable Templates
- Add Custom Templates (Administrator)
- Download Templates
Add Custom Templates (Administrator)
- Open the Sample Type, Source Type, or Assay Design for which you want to include customized templates.
- Select Manage > Manage Templates.
- You'll see the Default Template and can click to download it as a starting place if desired.
- Click Add a Template to add your own additional ones.
- Give the new template a name that will help your users know when to use it.
- Select or drag and drop the new template file into the target area.
- Click Save.
Download Templates
When a data structure has more than the default template defined, users will be able to select a template by name.Related Topics
LIMS: Samples
- Sample Types:
- Samples:
- Add Samples
- Aliquots, Derivatives, and Sample Pooling
- Sample Details: Lineage, Aliquots, Assays, Jobs, Timeline
- Sample Status
- Sample Picklists
- Sample Grids Containing Samples of Several Types
- Multi-tab Excel Exports
- Sample Finder
Related Topics
- Entity Lineage
- sampleSets: General information about Sample Types
- sampleIDs
- labkeyDataStructures
- useSampleSets
- deriveSamples
- exampleSampleSets
- uniqueStorageIds
- dataClass
- barTender
Print Labels with BarTender
This topic describes how to use LabKey applications, including Sample Manager and Biologics LIMS, with BarTender for printing labels for your samples. Note that an administrator must first complete the one-time steps to configure BarTender Automation. Once configured any user may send labels to the web service for printing.
Configuration steps and management of label templates:Print BarTender Labels for Samples
Before you can print labels, an administrator must have completed the one-time setup steps in this topic, and configured LabKey to print labels. The admin can also specify a folder-specific default label file. When printing, the user can specify a different variant to use.
After configuring BarTender, all users will see the options to print labels in the user interface.
Print Single Sample Label
Open the Sample Type from the main menu, then open details for a sample by clicking the SampleID.
Select Print Labels from the Manage menu.
In the popup, specify (or accept the defaults):
- Number of copies: Default is 1.
- Label template: Select the template to use among those configured by an admin. If the admin has set a default template, it will be preselected here, but you can use the menu or type ahead to search for another.
- Click Yes, Print to send the print request to BarTender.
Print Multiple Sample Labels
From the Sample Type listing, use checkboxes to select the desired samples, then select Print Label from the (Export) menu.
In the popup, you have the following options:
- Number of copies: Specify the number of labels you want for each sample. Default is 1.
- Selected samples to print: Review the samples you selected; you can use the Xs to delete one or more of your selections or open the dropdown menu to add more samples to your selection here.
- Label template: Select the template to use among those configured by an admin. You can type ahead to search. The default label template file can be configured by an admin.
- Click Yes, Print to send the print request to BarTender.
Download BarTender Template
To obtain a BarTender template in CSV format, select > Download Template.

Troubleshooting
If you have trouble printing to BarTender from Chrome (or other Chromium-based browser), try again using Firefox.
If you are not seeing values populate in your labels, confirm that you have configured BarTender to have the necessary data source connection.
Error Reporting
If there is a problem with your configuration or template, you will see a message in the popup interface. You can try again using a different browser, such as Firefox, or contact an administrator to resolve the configuration of BarTender printing.
Related Topics
LIMS: Assay Data
| SampleID | Expression Run | SampleDate | InjVol | Vial | Cal CurveID | Dilution | ResultID | MAb |
|---|---|---|---|---|---|---|---|---|
| 2016_01_22-C3-12-B9-01 | ER-005 | 2016-01-23 | 30 | 2:A,19 | 46188 | 1.0000 | 47836 | 0.2000 |
| 2016_04_21-C7-73-B6-02 | ER-005 | 2016-04-22 | 30 | 2:B,19 | 46188 | 1.0000 | 47835 | 0.2300 |
| 2015_07_12-C5-39-B4-02 | ER-004 | 2015-07-13 | 30 | 2:C,19 | 46188 | 1.0000 | 47834 | 0.3000 |
Documentation
Learn more about using the general assay framework in the Sample Manager documentation here:Additional Features with LabKey LIMS
Additional Features with Biologics LIMS
LIMS: Charts
Add a Chart
To add a chart:- On a data grid, click the Charts menu and choose Create Chart.
- In the pop up dialog, enter the following fields and options:
- Enter a Name and select whether to:
- Make this chart available to all users
- Make this chart available in child folders
- Select a Chart Type:
- Bar
- Box
- Line
- Pie
- Scatter
- Depending on the chart type, you will see selectors for X-axis, Y-axis, and various other settings needed for the chart. Required fields are marked with an asterisk.
- Once you have made selections, you will see a preview of the chart.
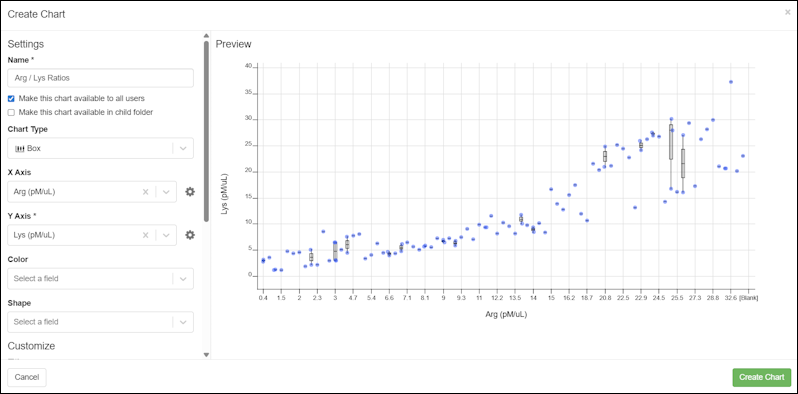
- Click Create Chart to save it.
Trendline Options
View a Chart
Any charts available on a data grid will be listed on the Charts menu above the grid.Selecting a chart will render it above the grid as shown here.
Edit a Chart
To edit a chart, open it from the menu and click the .Error Bars and Aggregation Methods
For Bar and Line charts, aggregation and error bar options are available by clicking the Y-axis gear icon.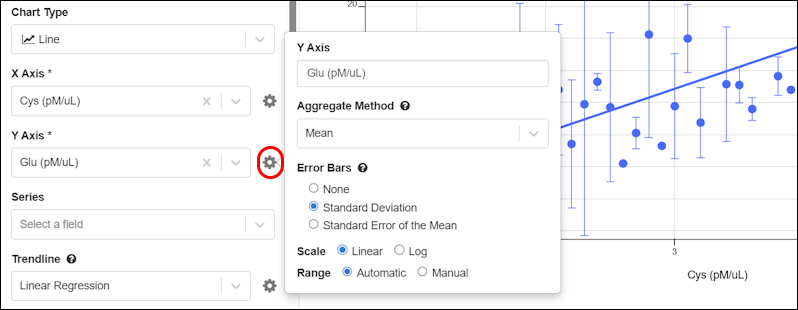 When the aggregation method is set to Mean, then the options for error bars are shown: None, Standard Deviation, Standard Error of the Mean.
When the aggregation method is set to Mean, then the options for error bars are shown: None, Standard Deviation, Standard Error of the Mean.
Export a Chart as PDF or PNG
Click the (Download) button to choose either PNG or PDF format for your export.Related Topics
LIMS: Storage Management
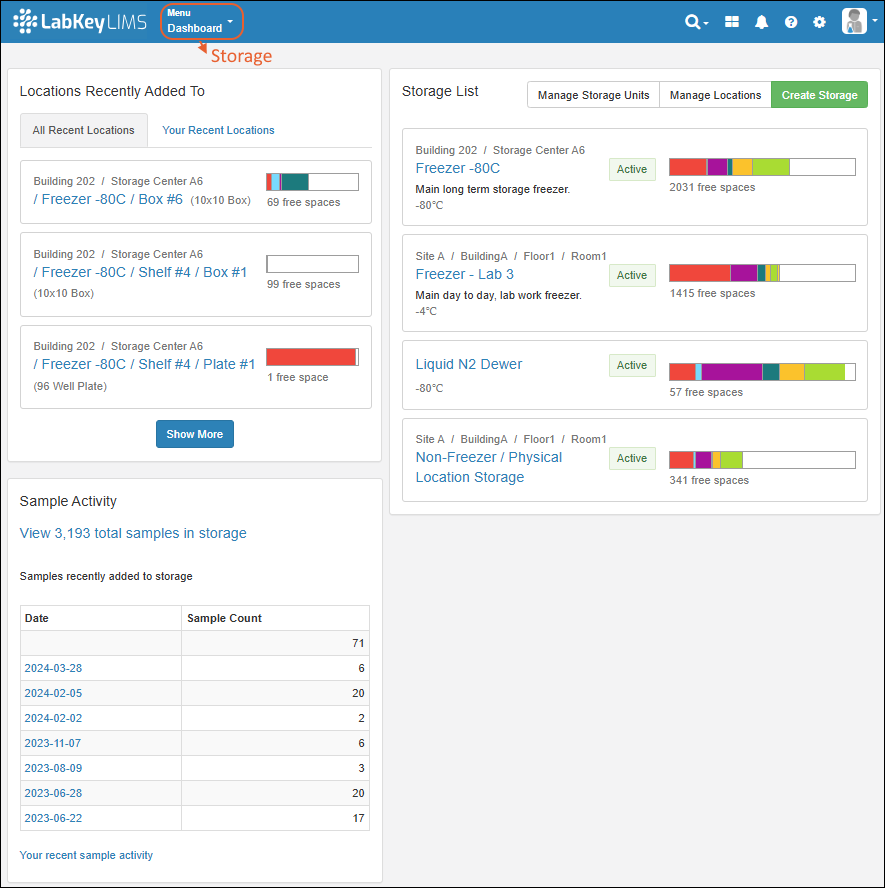
Topics
Learn more in the Sample Manager documentation here:- Storage Management
- Create Storage
- Store Samples
- Manage Storage
- Manage Storage Unit Types
- View Storage Details
- Edit Storage Definition
- Manage Storage Locations
- Move Stored Samples
- Manage Stored Samples
- Check Out and Check In
- View Storage Activity
- Migrate Storage Data into LabKey
- freezerRoles
- Storage Management API
- Shared Storage Across Folders
LIMS: Workflow
- Use workflow templates to standardize common task sequences
- Create jobs to track and prioritize sequential tasks
- Assign work to the right users
- Track progress toward completion
Biologics LIMS
- Record and document your ongoing work in a data-aware Electronic Lab Notebook.
- Easily and uniquely register all biological entities individually or in bulk.
- Track and query the lineage of samples throughout many generations of derivation.
- Integrate biological entity and sample information with downstream assay data used to evaluate therapeutic properties and provide a holistic view of experiment results.
- Manage and monitor the execution of laboratory tasks and requests, supporting efficient collaboration across teams.
- Manage the contents of your freezers and other storage systems using a digital match for your physical storage.
Biologics Release Notes
Biologics Overview
The Biologics LIMS product builds upon the same application core as Sample Manager. As you get started with Biologics, you may find the introductory Sample Manager documentation helpful in learning the basics of using the application.Basic Navigation and Features
Bioregistry
- Biologics: Bioregistry
- Create Registry Sources
- Register Nucleotide Sequences
- Register Protein Sequences
- Register Leaders, Linkers, and Tags
- Register Molecules
- Protein Sequence Annotations
- CoreAb Sequence Classification
- Biologics: Chain and Structure Formats
- Molecules, Sets, and Molecular Species
- Vectors, Constructs, Cell Lines, and Expression Systems
- Entity Lineage
- Bulk Registration of Entities
- Registry API
- Configuring Columns Seen in Grids and Detail Views
- Biologics: Protect Sequence Fields
Samples
Plates
Assay Management
- Biologics: Assay Data
- Biologics: Assay Batches and QC
- Biologics: Upload Assay Data
- Biologics: Specialty Assays
Media Registration
- Biologics: Media Registration
- Managing Ingredients and Raw Materials
- Registering Mixtures (Recipes)
- Registering Batches
Workflow
ELN
Storage Management
Related Topics
Introduction to LabKey Biologics
- A consolidated Bioregistry knowledge base to store and organize all your data and entity relationships
- Sample and Storage management tracking production batches, inventories, and all contents of your physical storage systems
- A data-integrated electronic lab notebook (ELN) for recording your research and coordinating review
- Tools promoting collaboration among teams for automated workflows, smooth handoffs, and reproducible procedures
- Media management for clear tracking of ingredients and mixtures in the Enterprise Edition
Knowledge Base
Bioregistry
The Bioregistry forms the team's shared knowledge base, containing detailed information on your sequences, cell lines, expression systems, and other research assets. The application is prepopulated with common Registry Source Types.
- Cell Lines
- Compounds
- Constructs
- Expression System
- Molecules
- Molecule Sets
- Molecule Species
- Nucleotide Sequences
- Protein Sequences
- Vectors
Entity Details
Click an entity in a grid to see its details page. Each details page shows the entity's properties and relationships to other entities.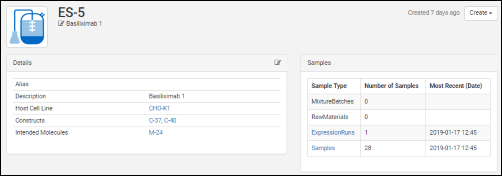 All details pages can be configured to show the most relevant data. For example, the details page for a protein sequence shows the chain format, average mass, extinction coefficient, the number of S-S bonds, etc., while the details page for a expression system shows the cell lines, constructs, and target molecule, as well as the samples drawn from it.
All details pages can be configured to show the most relevant data. For example, the details page for a protein sequence shows the chain format, average mass, extinction coefficient, the number of S-S bonds, etc., while the details page for a expression system shows the cell lines, constructs, and target molecule, as well as the samples drawn from it.
Entity Relationships
Each details page contains a panel of relationships to other entities. For example, for a given protein sequence, the relationship panel shows: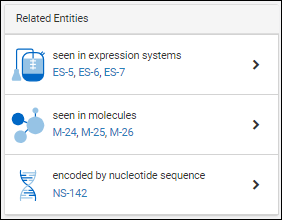
- which expression systems it is included in
- which molecules it is a part of
- which nucleotide sequence encodes for it
Sample Management and Data Integration
Data about your samples, experiments, and instrument assay results can be integrated to provide a full data landscape. Design customized methods for representing and linking your information. Using Biologics LIMS follows the same interface and has all the capabilities of the other Sample Management applications.Analytics and Visualizations
Analytics and visualizations can be included throughout LabKey Biologics to provide key insights into the large molecule research process.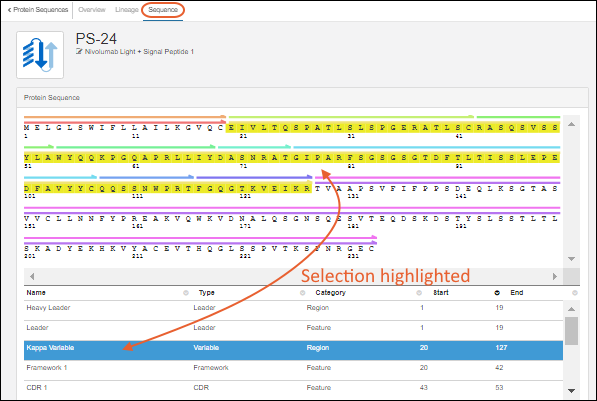
Protein Classification Engine
When new protein sequences are added to the Registry, they are passed to the classification and annotation engine, which calculates their properties and identifies key regions on the protein chain, such as leader sequences, variable regions, and constant regions. (Researchers can always override the results of the classification engine, if desired.) Results are displayed on the Sequence tab of the protein's detail page. Identified regions are displayed as multicolored bars, which, when clicked, highlight the annotation details in the scrollable section below the sequence display.Workflow Collaboration
A fully customizable task-based workflow system is included in LabKey Biologics. Individual users can all be working from the same knowledge base with personalized task queues and priorities. Several example jobs with representative tasks are included in your trial, you can also create your own.Media Management (Enterprise Edition feature)
In the Media section of the main menu, you will find sections for tracking:- Batches
- Ingredients
- Mixtures
- Raw Materials
Electronic Lab Notebook
Our data-integrated ELN (electronic lab notebook) helps you record results and support collaboration and publication of your research. Directly link to relevant registry entities, specific result runs, and other elements of your biologics research. Submit for review and track feedback and responses within the same application.Learn more in this section:Storage/Freezer Management
When you use Storage Management tools within LabKey Biologics, you can directly track the locations of samples and media in all the freezers and other storage systems your team uses. Create an exact digital match of your physical storage and record storage details and movements accurately. Sample data is stored independently of storage data, so that all data is retained even after a sample is consumed or removed from storage.Learn more in the Sample Manager documentation here:Release Notes: Biologics
Biologics LIMS offers strong support for antibody discovery operations. Learn more in our webinar:
Biologics LIMS includes all features available in LabKey LIMS, plus:
- Bioregistry and classification engine
- Media management
- Molecule property calculator
- Plate and plate set support
- Antibody screening and characterization
Each new release of Biologics LIMS includes all the feature updates covered in the Sample Manager and LIMS Release Notes, plus additional features listed on this page.
Release 26.2, February 2026
- GenBank import improved: nearly all information GenBank files is captured on import, including the original file.
- Improved Molecule creation: Select protein sequences to kick off the molecule creation process.
- Column widths now adjust dynamically, allowing more columns to be visible at once with less horizontal scrolling. (docs)
- Configured URL links can now be opened in a new browser tab for easier comparison and multitasking. (docs)
- Entities you don't have access to in lineage views are now shown as restricted rather than being omitted, preserving full context without exposing details. (docs)
- Sample Status is available as a filter for "All Sample Types" in Sample Finder. (docs)
Release 26.1, January 2026
- Support for multiple unit types provides improved inventory and material management. (docs)
- Move workflow jobs to different folders to better reflect changes in projects or organization. (docs)
- Workflow tasks now support sample filters, allowing you to control which samples are included at each step. (docs)
- Improved plot customization with new layout, axis, size, color, and per-series line controls. (docs)
- Client APIs can query and update samples using the RowId value; using the LSID value is no longer required.
- Sample names (SampleId) can be updated via a file, when RowId is provided.
Release 25.12, December 2025
- Amount and Units Fields - Improvements have been made to ensure that the Amounts & Units fields function as paired fields. (docs)
- Negative Amount Values Disallowed - Sample Manager now enforces that the Amounts field cannot have a negative value.(docs)
- Identifying Fields - Identifying fields are now shown in more assay import scenarios. (docs)
Release 25.11, November 2025
- Audit log captures the method used to insert, update, and delete records. (docs)
- When an ELN notebook is recalled by an administrator, the author will now receive an email notification, improving visibility and timely follow-up.
- The Customize Grid View and Filter pop-up dialogs now list fields alphabetically, making it faster and more intuitive to find and select fields.
- Error bars are available on Bar and Line charts. (docs)
- Multiple charts can be displayed above data grids. Select up to 5 charts to display. (docs)
Release 25.3, March 2025
- Rapidly find the plates and experiments in which samples have been used, and vice versa.
- Automatically generate analytics like regressions and statistics to accelerate your work
Release 25.2, February 2025
- Support for advanced plate layouts using using dilutions.
- Use the "Replicate Group" column to denote a plate well as a replicate instead of setting the well's type to "Replicate".
- Replicate wells have a type of "Sample" and the "Replicate Group" will need to be filled in.
- Add Samples to an existing Plate Set.
- Navigate from a plate set to any notebooks that reference it.
- Edits to outlier exclusions will result in the rerunning of any transform scripts that are configured to run on update.
Release 25.1, January 2025
- Users can now specify hit selection filter criteria on Assay fields. When a run is imported/edited the hit selections for the assay results will be recomputed and automatically applied based on these criteria.
- Navigate from a sample to the plate(s) it has appeared on.
- Perform many types of linear regression analysis and chart them.
- Exclude outlier plate-based assay data points and have that reflected in calculations and charts.
Release 24.12, December 2024
- Plate sets can be referenced from an Electronic Lab Notebook.
Release 24.11, November 2024
Major antibody discovery and characterization updates including:
- Campaign modeling with plate set hierarchy support.
- Plan plates easier with graphical plate design and templating.
- Automate routine analyses from raw data collected.
- Perform hit selection from multiple, integrated results across plates and data types.
- Generate instructions for liquid handlers and other instruments.
- Automatically integrate multi-plate results including interplate replicate aggregation.
- Dive deeper into plated materials to understand their characteristics and relationships from plates.
Release 24.10, October 2024
- Charts are added to LabKey LIMS, making them an "inherited" feature set from other product tiers. (docs)
Release 24.7, July 2024
- A new menu has been added for exporting a chart from a grid. (docs)
Release 23.12, December 2023
- The Molecule physical property calculator offers additional selection options and improved accuracy and ease of use. (docs)
Release 23.11, November 2023
Release 23.9, September 2023
- Charts, when available, are now rendered above grids instead of within a popup window. (docs)
Release 23.4, April 2023
- Molecular Physical Property Calculator is available for confirming and updating Molecule variations. (docs)
- Lineage relationships among custom registry sources can be represented. (docs)
- Users of the Enterprise Edition can track amounts and units for raw materials and mixture batches. (docs | docs)
Release 23.3, March 2023
- Potential Backwards Compatibility Issue: In 23.3, we added the materialExpDate field to support expiration dates for all samples. If you happen to have a custom field by that name, you should rename it prior to upgrading to avoid loss of data in that field.
- Note that the built in "expirationDate" field on Raw Materials and Batches will be renamed "MaterialExpDate". This change will be transparent to users as the new fields will still be labelled "Expiration Date".
Release 23.2, February 2023
- Protein Sequences can be reclassified and reannotated in cases where the original classification was incorrect or the system has evolved. (docs)
- Lookup views allow you to customize what users will see when selecting a value for a lookup field. (docs)
- Users of the Enterprise Edition may want to use this feature to enhance details shown to users in the "Raw Materials Used" dropdown for creating media batches. (docs)
Release 23.1, January 2023
- Heatmap and card views of the bioregistry, sample types, and assays have been removed.
- The term "Registry Source Types" is now used for categories of entity in the Bioregistry. (docs)
Release 22.12, December 2022
- Projects were added to the Professional Edition of Sample Manager, making this a common feature shared with other tiers.
Release 22.11, November 2022
- Improvements in the interface for managing Projects. (docs)
- New documentation:
Release 22.10, October 2022
- Improved interface for creating and managing Projects in Biologics. (docs)
Release 22.9, September 2022
- When exploring Media of interest, you can easily find and review any associated Notebooks from a panel on the Overview tab. (docs)
Release 22.8, August 2022
- Search for data across projects in Biologics. (docs)
Release 22.7, July 2022
- Biologics subfolders are now called 'Projects'; the ability to categorize notebooks now uses the term 'tags' instead of 'projects'. (docs | docs)
Release 22.6, June 2022
- New Compound Bioregistry type supports Simplified Molecular Input Line Entry System (SMILES) strings, their associated 2D structures, and calculated physical properties. (docs)
- Define and edit Bioregistry entity lineage. (docs)
- Bioregistry entities include a "Common Name" field. (docs)
Release 22.3, March 2022
- Mixture import improvement: choose between replacing or appending ingredients added in bulk. (docs)
Release 21.12, December 2021
Release 21.11, November 2021
- Apply a container specific prefix to all bioregistry entity and sample naming patterns. (docs)
Release 21.10, October 2021
- Customize the definitions of data classes (both bioregistry and media types) within the application. (docs)
Release 21.9, September 2021
- Customize the names of entities in the bioregistry (docs)
Release 21.7, July 2021
- Removal of the previous "Experiment" mechanism. Use workflow jobs instead.
Release 21.5, May 2021
- Nucleotide and Protein Sequence values can be hidden from users who have access to read other data in the system. (docs)
Release 21.4, April 2021
- Specialty Assays can now be defined and integrated, in addition to Standard Assays (docs)
- Creation of Raw Materials in the application uses a consistent interface with other sample type creation (docs)
Release 21.3, March 2021
- Biologics LIMS begins using the same user interface as Sample Manager.
- Release notes for this and other versions prior to this change can be found in the documentation archives.
Release 17.1, March 2017
Biologics: Navigate
- LabKey Biologics: Home Page
- Navigation
- Menu > Registry Sources - Browse all of the registry sources.
- Menu > Sample Types - A dashboard for tracking samples.
- Menu > Assays - Assay results for candidate molecules.
- Menu > Storage - Manage freezer contents and capacities.
- Menu > Workflow - Create and track custom workflow jobs and tasks.
- Menu > Media - A dashboard for ingredients, media, mixtures, and recipes. (Enterprise Edition Feature)
- Menu > Picklists - User-defined lists of samples.
- Menu > Notebooks - Electronic lab notebooks integrated directly with your data.
LabKey Biologics: Home Page
The main dashboard on the home page provides quick links into different aspects of the data.The top header bar is available throughout the application and includes:- The clickable logo, for returning to this dashboard at any time.
- A main Menu button, giving access to all of your data and activities.
- A set of common resources in the upper right:
- : Search options
- : A product selection menu for switching between Biologics and LabKey Server when needed
- : Notifications
- : Help Opens the documentation for LabKey Biologics LIMS.
- : Administration Settings
- A user menu, indicated by your avatar, or a default graphical one
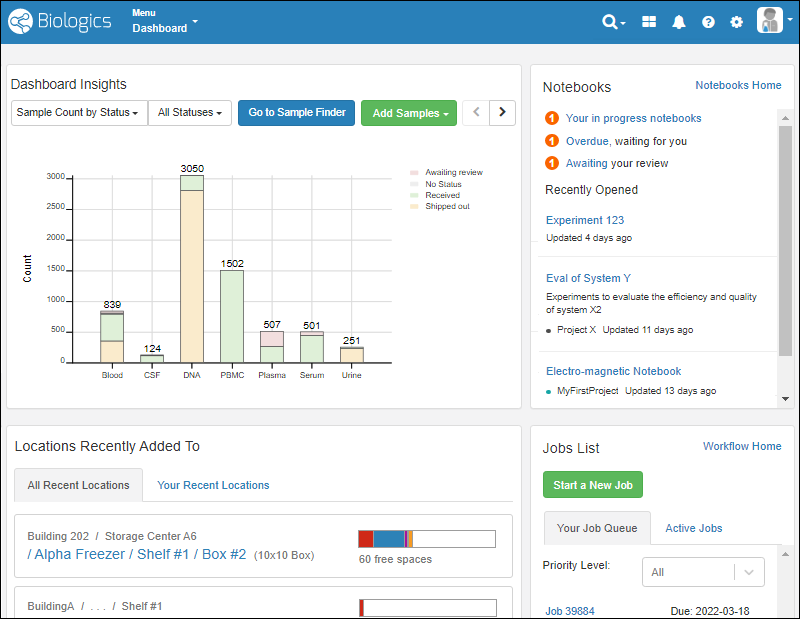 On the dashboard, you'll see panels displaying:
On the dashboard, you'll see panels displaying:
- Dashboard Insights: A selectable set of visualizations, defaulting to the Sample Count by Status.
- Sample Types can be selectively excluded from the graphs on this dashboard, as described here.
- Notebooks: See task status, recent notebooks, and link to your Electronic Lab Notebooks dashboard.
- Locations Recently Added To: Access to storage where samples have been added by you or others.
- Jobs List: The home for workflow jobs. If any tasks were assigned to you, they would be shown on Your Job Queue and can be filtered by priority.
Header Menus
The menus in the upper right offer these selections:- : Search options:
- Site-wide text search
- Find samples by barcode () or ID ()
- Sample Finder: Find samples by their properties or associated data
- : A product selection menu for switching between Biologics and LabKey Server when needed
- : Notifications
- Documentation:
- Help: Opens the documentation for LabKey Biologics LIMS.
- Release Notes: Biologics
- : Administration Settings:
- Application Settings: See below.
- Audit Logs
- Groups
- Permissions
- Folders
- Users
- User avatar menu
- Notification Settings
- Profile: Your user details, including the option to add an avatar image.
- Sign Out (or Sign In)
Application Settings
Select > Application Settings to manage:- Display Settings
- Notebook Settings
- BarTender Configuration
- ID/Name Settings
- Manage Sample Statuses
- Audit Logging
- Protected Data Settings
Navigation
The main Menu is available throughout the application and gives you quick access to all aspects of Biologics. It will include the name of the category you are in ("Dashboard" if not one of the specific categories).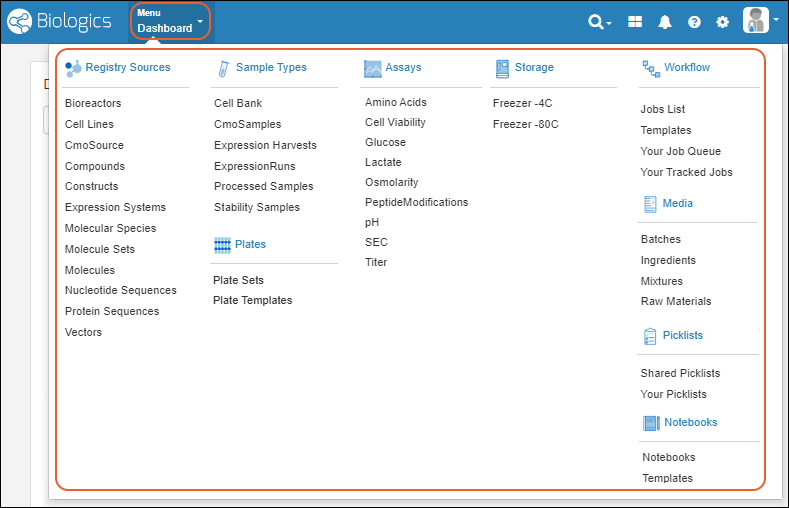 Click the menu categories for sub-dashboards, such as for Registry Sources, Notebooks, or Sample Types. Click individual menu items for that specific category.
Click the menu categories for sub-dashboards, such as for Registry Sources, Notebooks, or Sample Types. Click individual menu items for that specific category.
Biologics: Projects and Folders
- Options for Sharing Biologics Data
- Use Multiple Biologics Folders
- ID/Name Settings
- Shared Data Structures (Sample Types Registry Source Types, Assays, Storage Systems)
Options for Sharing Biologics Data
Biologics runs in the context of a LabKey project or folder container. Each container can have unique permissions assignments and data, supporting a secure and isolated workspace. If desired, many resources, including assay designs and sample type definitions, can also be shared across multiple containers. Further, within a LabKey project of type "Biologics", an administrator can add folders which allow data partitioning.The Biologics administrator should determine the configuration that best represents the organization.Example Scenarios:1. You might have two groups doing distinct work with no need to share data or compare results. These two groups could each use a separate Biologics container (LabKey project or folder) and keep all resources within it. Permissions are assigned uniquely within each container, with no built-in interaction between the data in each.2. If two groups needed to use the same assay designs or sample types, but would not otherwise share or compare the actual data, those definitions could be placed in the Shared project, with all collection and analysis in containers (LabKey projects or folders) for the two groups as in option 1. Note that in this scenario, samples created of those shared types would have the same naming pattern and if you used a container-specific prefix, it would not be applied to the samples of the shared type.3. If multiple groups need to share and compare data, and also keep clear which data came from which group originally, you could apply a container prefix in each group's folder so that all data would be consistently identified. In this scenario, each container would need its own sample type definitions so that the naming patterns would propagate the prefix into the created samples.4. For more integrated sharing, configure Biologics in a top level LabKey project, then add one or more subfolders, all also of folder type "Biologics". Bioregistry and Sample definitions can be shared among the Folders and permissions controlled independently. Learn about this option in the next section.5. Storage systems (freezers, etc.) can be shared among folders with different permissions. Users will only be able to see details for the stored samples to which they have been granted access. Learn more about shared storage in the Sample Manager documentation:Manage Multiple Biologics Folders
When using Biologics in a top-level LabKey Server container, administrators have the option to manage multiple Biologics Folders directly from within the admin interface. Note that this is not supported when the top level Biologics container is a folder or subfolder in LabKey Server.To manage Folders, select > Folders. You can customize types of data and specific storage systems available per Folder. Learn more in the Sample Manager documentation here: When in use, you'll see Folders listed on the left side of the main menu. Select the desired Folder, then click the item of interest on the menu or the links for its dashboard or settings. You will see the contents of the application 'scoped' to the selected Folder. Selecting the top-level container (aka the "home") will show all data the user can access across all Folders to which they have been granted "Read" permissions.Cross-Folder Actions
When multiple Biologics Folders are in use, the notes about cross-folder actions are the same as for Sample Manager, where Registry Sources are called Sources. Learn more in this topic:Restricted Visibility of Inaccessible Data
When a user has access to a subset of folders, there can be situations where this user will be restricted from seeing data from other folders, including identifiers such as Sample IDs that might contain restricted details. In detail pages, listing pages, and grids, entities a user does not have access to will show "unavailable" in the display instead of a name or rowid. Learn more in this topic:ID/Name Settings
Select > Application Settings and scroll down to the ID/Name Settings section to control several options related to naming of entities in the application.Force Usage of Naming Patterns for Consistency
To maintain consistent naming, particularly when using container-specific naming prefixes, you may want to restrict users from entering their own names for entities. Learn more here:Apply Naming Prefix
You can apply a container-specific naming prefix that will be added to naming patterns to assist integration of data from multiple locations while maintaining a clear association with the original source of that data.This prefix is typically short, 2-3 characters long, but will not be limited. Prefixes must be unique site-wide, and should be recognizable to your users. Before setting one, make sure you understand what will happen to the naming patterns and names of existing entities in your application.- The prefix will be applied to names created with naming patterns for all Sample Types and Registry Source Types in the container.
- New samples and entities created after the addition of the prefix will have names that include the prefix.
- Existing samples and entities created prior to the addition of the prefix will not be renamed and thus will not have the prefix (or might have a different previously-applied prefix).
- Sample aliquots are typically created and named including the name of the sample they are aliquoted from. This could mean that after the prefix is applied, new aliquots may or may not include the prefix, depending on whether the originating sample was created before or after the prefix was applied. Learn more about aliquot naming here: aliquotIDs.
- Select > Application Settings.
- Scroll down to the ID/Name Settings section.
- Enter the prefix to use. You will see a preview of what a name generated with the Naming Pattern with the prefix applied might look like using a representative example, Blood-${GenId}:
- Click Apply Prefix to apply it.
- This action will change the Naming Pattern for all new and existing Sample Types and Registry Source Types. No existing IDs/Names will be affected. Are you sure you want to apply the prefix?
- Click Yes, Save and Apply Prefix to continue.
Naming Pattern Elements/Tokens
The sampleCount and rootSampleCount tokens are used in Naming Patterns across the application.Learn more about these naming pattern tokens in this topic:Shared Data Structures (Sample Types, Registry Source Types, Assays, and Storage)
Sample Types, Registry Source Types, Assays, and Storage Systems are defined in the home folder and available in subfolders, provided an administrator has not 'hidden' them. Administrators can edit these shared data structures from the home any folder, but the changes will be made at the home level, applying to all folders.In addition, any Sample Types, Registry Source Types, and Assay definitions defined in the /Shared project will be available for use in LabKey Biologics. You will see them listed on the menu and dashboards alongside local definitions.From within the Biologics application, users editing (or deleting) a shared definition will see a banner indicating that changes will affect other folders. The same message is applied when editing a data structure from a folder within the Biologics application.Related Topics
Biologics: Bioregistry
- Create Registry Sources
- Register Nucleotide Sequences
- Register Protein Sequences
- Register Leaders, Linkers, and Tags
- Register Molecules
- Registry Reclassification
- Biologics: Terminology
- Protein Sequence Annotations
- CoreAb Sequence Classification
- Biologics: Chain and Structure Formats
- Molecules, Sets, and Molecular Species
- Vectors, Constructs, Cell Lines, and Expression Systems
- Compounds and SMILES Lookups
- Entity Lineage
- Customize the Bioregistry
- Bulk Registration of Entities
- Use the Registry API
- Biologics: Detail Pages and Entry Forms
Create Registry Sources
- Create Registry Sources in the User Interface
- Create/Import Registry Sources from File
- Naming Patterns
- Hide Name Entry/Edit Options
- Review Entity Details
Create Registry Sources in the User Interface
In this example, we show creation of a new cell line. Other kinds of sources will have different fields that compose them, and may also have additional tabs in the creation wizard. See specific documentation listed at the end of this topic.- From the main menu, click the type of registry source to create. Then use the Add > menu:
- Add Manually: Use the data entry wizard.
- Add Manually from Grid: Create one or more using a grid interface.
- Import from File
Add Manually
- Provide the details in the registration wizard:
- Name: Provide a short unique name, or leave this field blank to have one generated using the naming pattern for this Registry Source Type.
- Hover over the to see an example generated name.
- Description: Optional, but will be shown in the grids and can be a helpful way to illustrate the entity.
- Common Name: Every entity includes a field for the common name.
- Remaining fields: Required fields are marked with an asterisk.
- When the fields are completed, click Finish to create the new entity.
Add Manually from Grid
Using the grid interface to create Registry Sources as described in the Sample Manager documentation for Sources.Note that this option not supported for all Registry Source Types. You will not see this option on the menu for Nucleotide Sequences, Protein Sequences, Molecules, Molecular Species, etc.Create/Import Entities from File
For bulk registration of many entities, including registry sources, samples, assay result data, ingredients, and raw materials, you can make use of importing new data from file. Templates are available to assist you in reliably uploading your data in the expected format.Learn more in the Sample Manager documentation for importing Samples from file.Update or Merge Registry Sources from File
To update data for existing Registry Sources from file, or to import a spreadsheet merging updates with creation of new Registry Sources, use Edit > Update from File.Learn more in the Sample Manager documentation for updating Samples from file.Entity Naming Patterns
If you do not provide a name, the naming pattern for the Registry Source Type will be used to generate one. Hover over the to see the naming pattern in a tooltip, as well as an 'example name' using that pattern.| Registry Source Type | Default Naming Pattern |
|---|---|
| Cell Lines | CL-${genId} |
| Compounds | CMP-${genId} |
| Constructs | C-${genId} |
| Expression Systems | ES-${genId} |
| Molecular Species | MSp-${genId} |
| Molecule Sets | MS-${genId} |
| Molecules | M-${genId} |
| Nucleotide Sequences | NS-${genId} |
| Protein Sequences | PS-${genId} |
| Vectors | V-${genId} |
Hide Name Entry/Edit Options
An administrator can hide the Name field for insert, update, or both. When insert of names is hidden, they will be generated using the naming pattern for the registry source type. When update of names is hidden, names remain static after entity creation.This can be done:- On an application-wide basis by disabling user-supplied names entirely.
- For some but not all Registry Source Types, by using Query Metadata overrides.
Review Registry Source Details
Once created, you'll see a grid of all entities of the given type when you select it from the main menu.To see details for a specific Registry Source, click the name.- Overview: Panels for details, samples, related entities (for built in Registry Source Types), notebooks, and parent sources.
- Lineage: See the lineage in graph or grid format.
- Samples: Contains a grid of all samples 'descended' from this source.
- Assays: See all assay data available for samples 'descended' from this source.
- Jobs: All jobs involving samples 'descended' from this source.
- Sequence (Available for Protein and Nucleotide Sequences).
- Molecular Property Calculator (Available for Molecules)
Related Topics
Register Nucleotide Sequences
Nucleotide Sequence Validation
For nucleotide sequences, we allow DNA and RNA bases (ACTGU) as well as the IUPAC notation degenerate bases (WSMKRYBDHVNZ). On import, whitespace will be removed from a nucleotide sequence. If the sequence contains other letters or symbols, an error will be raised.For protein sequences, we only allow standard amino acids letters and zero or more trailing stop codon '*'. On import, whitespace will be removed from a protein sequence. If the sequence contains stop codons in the middle of the sequence or a other letters or symbols, an error will be raised.When translating a nucleotide codon triple to a protein sequence, where the codon contains one or more of the degenerate bases, the system attempts to find a single amino acid that could be mapped to by all of the possible nucleotide combinations for that codon. If a single amino acid is found, it will be used in the translated protein. If not, the codon will be translated as an 'X'.For example, the nucleotide sequence 'AAW' is ambiguous since it could map to either 'AAA' or 'AAT' (representing Lysine and Asparagine respectively), so 'AAW' will be translated as an 'X' However, 'AAR' maps to either 'AAA" or 'AAG' which are both are translated to Lysine, so it will be translated as a 'K'.Create a Nucleotide Sequence
To add a new nucleotide sequence to the registry:- Open the Nucleotide Sequence page, then select Add > Add manually.
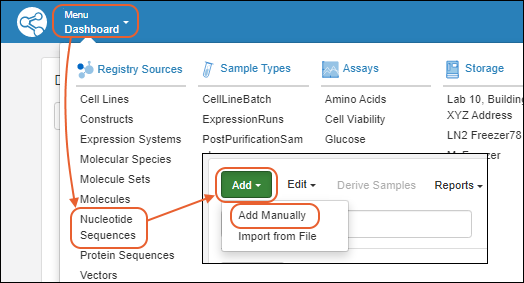 The creation wizard has two tabs:
The creation wizard has two tabs:
Details
On the Register a new Nucleotide Sequence page, in the Details panel, populate the fields: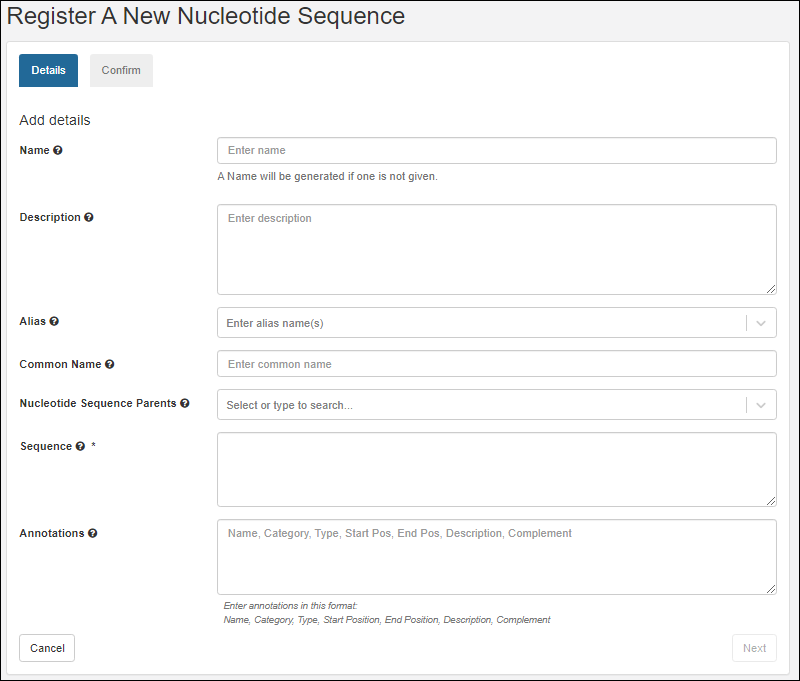
- Name: Provide a name, or one will be generated for you. Hover to see the naming pattern
- Description: (Optional) A text description of the sequence.
- Alias: (Optional) Alternative names for the sequence. Type a name, click enter when complete. Continue to add more as needed.
- Common Name: (Optional) The common name for this sequence, if any.
- Nucleotide Sequence Parents: (Optional) Parent components. A related sequence the new sequence is derived from, for example, related as a mutation. You can select more than one parent. Start typing to narrow the pulldown menu of options.
- Sequence: (Required) The nucleotide sequence
- Annotations: (Optional) A comma separated list of annotation information:
- Name - a freeform name
- Category - region or feature
- Type - for example, Leader, Variable, Tag, etc.
- Start and End Positions are 1-based offsets within the sequence.
- Description
- Complement
Confirm
Review the details on the Confirm tab.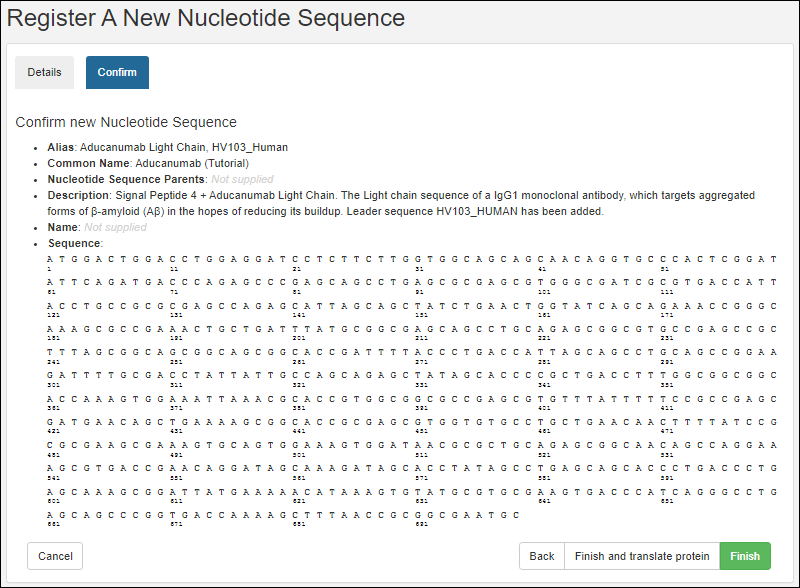 Options to complete registration:
Options to complete registration:
- Finish: Register this nucleotide sequence and exit.
- Finish and translate protein: Both register this nucleotide sequence and register the corresponding protein. This option will take you to the registry wizard for a new protein, prepopulating it with the protein sequence based on the nucleotide sequence you just defined.
Related Topics
Register Protein Sequences
- Via the nucleotide sequence wizard. When registering a nucleotide sequence, you have the option of continuing on to register the corresponding protein sequence.
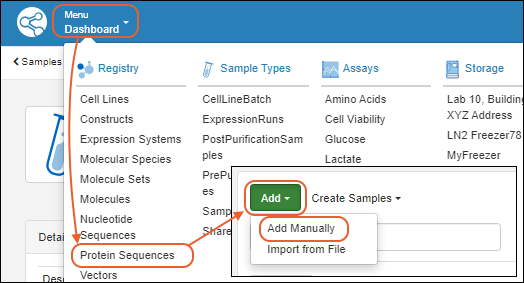
- Via the header bar. Select Registry > Protein Sequences.
- Select Add > Add manually.
Protein Sequence Wizard
The wizard for registering a new protein sequence proceeds through five tabs:Details
- Name: Provide a name, or one will be generated for you. Hover to see the naming pattern
- Description: (Optional) A text description of the sequence
- Alias: (Optional) List one or more aliases. Type a name, click enter when complete. Continue to add more as needed.
- Organisms: (Optional) Start typing the organism name to narrow the pulldown menu of options. Multiple values are accepted.
- Protein Sequence Parents: (Optional) List parent component(s) for this sequence. Start typing to narrow the pulldown menu of options.
- Seq Part: (Optional) Indicates this sequence can be used as part of a larger sequence. Accepted values are 'Leader', 'Linker', and 'Tag'. When set, chain format must be set to 'SeqPart'.
Sequence
On the sequence tab, you can translate a protein sequence from a nucleotide sequence as outlined below. If you prefer to manually enter a protein sequence from scratch click Manually add a sequence at the bottom.- Nucleotide Sequence: (Optional) The selection made here will populate the left-hand text box with the nucleotide sequence.
- Translation Frame: (Required). The nucleotide sequence is translated into the protein sequence (which will be shown in the right-hand text box) by parsing it into groups of three. The selection of translation frame determines whether the first second or third nucleotide in the series 'heads' the first group of three. Options: 1,2,3.
- Sequence Length: This value is based on the selected nucleotide sequence.
- Nucleotide Start: This value is based on the nucleotide sequence and the translation frame.
- Nucleotide End: This value is based on the nucleotide sequence and the translation frame.
- Translated Sequence Length: This value is based on the nucleotide sequence and the translation frame.
- Protein Start: Specific the start location of the protein to be added to the registry.
- Protein End: Specific the end location of the protein to be added to the registry.
Annotations
The annotations tab displays any matching annotations found in the annotation library. You can also add annotations manually at this point in the registration wizard.- Name: a freeform name
- Type: for example, Leader, Variable, Tag, etc. Start typing to narrow the menu options.
- Category: 'Feature' or 'Region'
- Description: (Optional)
- Start and End Positions: 1-based offsets within the sequence
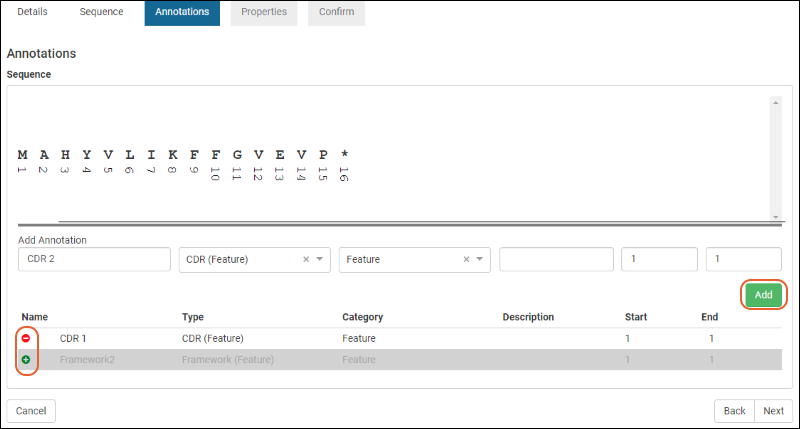 For complete details on using the annotation panel see Protein Sequence Annotations.Click Next to continue the wizard.
For complete details on using the annotation panel see Protein Sequence Annotations.Click Next to continue the wizard.
Properties
- Chain Format: select a chain format from the dropdown (start typing to filter the list of options). An administrator defines the set of options on the ChainFormats list. LabKey Biologics will attempt to classify the protein's chain format if possible.
- ε: the extinction coefficient
- Avg. Mass The average mass
- Num. S-S The number of disulfide bonds
- pI The isoelectric point
- Num Cys. The number of cysteine elements
Confirm
The Confirm panel provides a summary of the protein about to be added to the registry.Click Finish to add the protein to the registry.Editing Protein Sequence Fields
Once you have defined a protein sequence, you can locate it in a grid and click the name to reopen to see the details. Some fields are eligible for editing. Those that are "in use" by the system or other entities cannot be changed. All edits are logged.Related Topics
Register Leaders, Linkers, and Tags
- From the main menu, click Protein Sequences.
- Select Add > Add Manually.
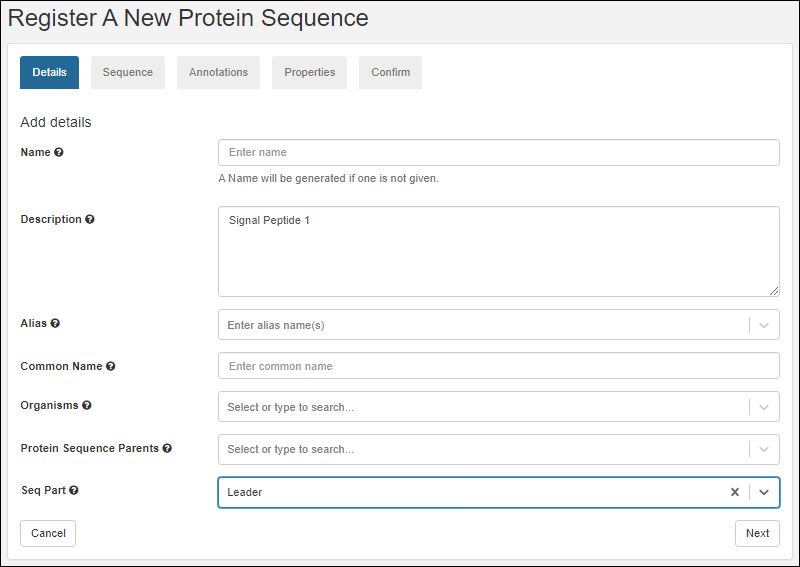
- This will start the wizard Register A New Protein Sequence.
- Leader
- Linker
- Tag
- Click Next.
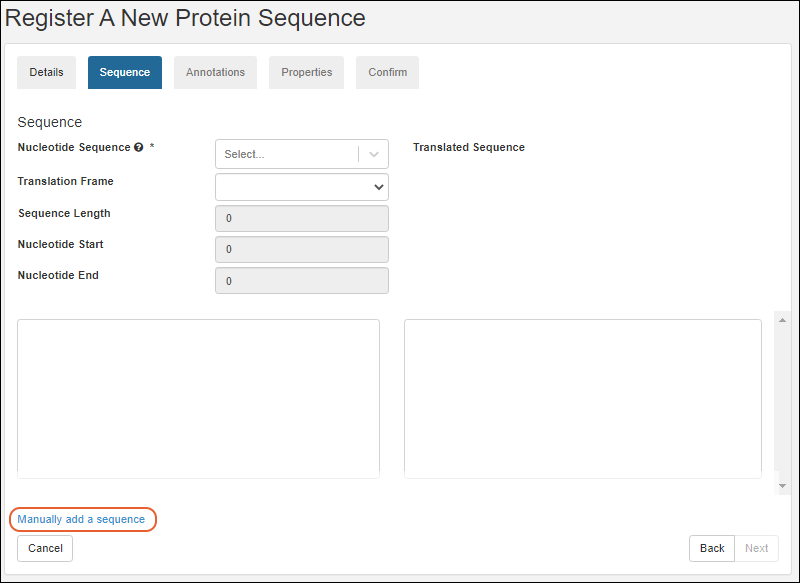
- On the Sequence tab, scroll down and click Manually add a sequence.
- Enter the sequence and click Next.
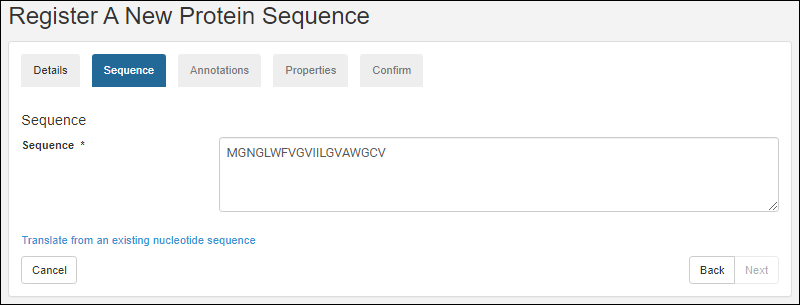
- Complete the rest of the wizard as described in the topic Register Protein Sequences.
Related Topics
Vectors, Constructs, Cell Lines, and Expression Systems
Definitions
- Vectors are typically plasmids that can inject genetic material into cells. Must have a specific nucleotide sequence.
- Constructs are Vectors which have been modified to include the genetic material intended for injection into the cell.
- Cell Lines are types of cells that can be grown in the lab.
- Expression Systems are cell lines that have been injected with a construct.
Add Manually within the User Interface
Select the desired entity from the main menu, then select Add > Add Manually.| Entity Type | Default Fields |
|---|---|
| Vector | Name Common Name Description Alias Sequence Selection Methods Vector Parents |
| Construct | Name Common Name Description Alias Vector Cloning Site Complete Sequence Insert Sequences Construct Parents |
| Cell Lines | Name Common Name Description Alias Expression System Stable Clonal Organisms Cell Line Parents |
| Expression Systems | Name Common Name Description Alias Host Cell Line Constructs Expression System Parents |
Add Manually from Grid
Select the desired entity from the main menu, then select Add > Add Manually from Grid. Learn about creating entities with this kind of grid in the topic:Import from File
Creation of many entities of a given type can also be done in bulk via file import using a template for assistance. Select Add > Import from File and upload the file. Learn more in this topic:Related Topics
Registry Reclassification
Reclassify Protein Sequence
From the Protein Sequences dashboard, select the protein sequence you wish to reclassify. Select Manage > Reclassify.Update Annotations
On the first page of the reclassification wizard, you will see the current Chain Format and Annotations for the sequence.- Review any updates to the Annotations that have been introduced since this protein sequence was last classified. These will be highlighted in green as shown below this list.
- Review the Chain Format assignment that may have been recalculated since the original classification. If needed, you can select another Chain Format.
- Add additional Annotations. Complete the required elements of the Add Annotation section, then click Add.
- Delete previous annotations using the in the "Delete" column. You'll see the deleted annotation struck out and have the option to re-add it by clicking the before continuing.
Adjust Molecules
After reviewing and adjusting the annotations, click Next to continue to the Molecules reclassification step.View Classification Status from Molecule
Starting from the overview page for a Molecule, you can see the Classification Status in the Details panel.Related Topics
Biologics: Terminology
- Definitions: "Registry Source" vs "Sample"
- Diagram of Registry Source Relationships
- Details of Each Registry Source Type
Definitions: "Registry Source" vs "Sample"
In LabKey Biologics LIMS, the terms "registry" and "samples" refer to different components or aspects of the system. The Registry is a database or collection of information about biologics entities, while Samples refer to the actual biological material or representations of it that are being managed within the system. The registry helps organize and provide context to the information about these entities, and samples are the tangible or digital representations linked to these entries in the registry.Registry Sources:- "Registry" typically refers to a comprehensive database or collection of information related to biologics entities. This can include details about biologics, such as antibodies, cell lines, or other biological molecules, that are being studied or managed within the system. The registry serves as a centralized repository for metadata and information about these biological entities, "Registry Sources", often including details like names, identifiers, descriptions, origin, properties, and associated data.
- "Samples" in LabKey Biologics usually refer to the physical or virtual representations of biological material that have been collected, stored, and are being managed within the system. Samples can include a variety of biological materials such as tissues, cell lines, fluids, or other substances. Each sample is typically associated with specific information, like collection date, source, location, processing history, and other relevant details. Samples are often linked to entries in the registry to provide a clear relationship between the biological entity and the actual physical or virtual sample associated with it.
Diagram of Registry Source Relationships
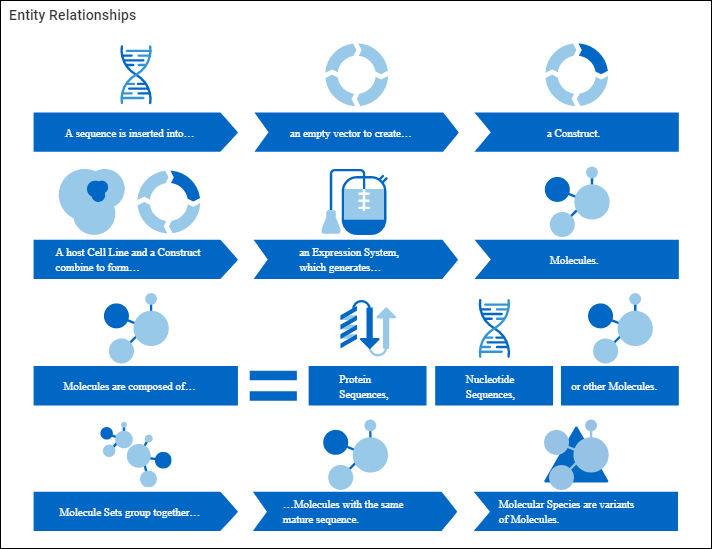
- A Sequence is inserted into an empty
- Vector to create a
- Construct.
- A host Cell Line and a Construct combine to form an
- Expression System, which generates
- Molecules.
- Molecules are composed of:
- Protein Sequences
- Nucleotide Sequences
- and/or other molecules.
- Molecule Sets group together molecules with the same mature sequence.
- Molecular Species are variants of molecules.
Molecule
Composed of a mixture of protein sequences, nucleotide sequences, chemistry elements, and other molecules. Generally, "molecule" refers to the target entity.| Example Molecules |
|---|
| Molecule W = 1 (protein sequence A) |
| Molecule X = 1 (protein sequence B) + 2 (protein sequence C) |
| Molecule Y = 1 (nucleotide sequence D) |
| Molecule Z = 2 (nucleotide sequence E) + 2 (protein sequence F) + 2 (molecule W) |
Molecule Set
The set of molecules that only differ from one another in terms of their leader sequences. See Vectors, Constructs, Cell Lines, and Expression Systems.Molecular Species
After protein expression and other processes, all the entities that are detected for a particular molecule (different cleavage sites, post-translational modifications, genomic drift). See Vectors, Constructs, Cell Lines, and Expression Systems.Protein Sequence
Single sequence comprised of amino acids (20 different amino acids).| Example Protein Sequences |
|---|
| protein sequence A = ACELKYHIKL CEEFGH |
| protein sequence B = HIKLMSTVWY EFGHILMNP |
Nucleotide Sequence
A single sequence comprised of nucleic acids, can be either DNA (A, T, G, C) or RNA (A, U, G, C).| Example Nucleotide Sequences |
|---|
| nucleotide sequence A (DNA) = AGCTGCGTGG GCACTCACGCT |
| nucleotide sequence B (RNA) = AGCUGUUGCA GCUUACAUCGU |
Vector
DNA molecule used as a vehicle to artificially carry foreign genetic material into another cell. The most common type of vector is a plasmid - a long double-stranded section of DNA that has been joined at the ends to circularize it. In molecular biology, generally, these plasmids have stretches of DNA somewhere in their makeup that allow for antibiotic resistance of some sort, providing a mechanism for selecting for cells that have been successfully transfected with the vector.Construct
A vector (generally a plasmid) that has had a stretch of DNA inserted into it. Generally, this DNA insertion encodes for a protein of interest that is to be expressed.Cell Line
A host cell line is transfected with a Construct, bringing the new DNA into the machinery of the cell. These transfected cells (called an Expression System) are then processed to select for cells that have been successfully transfected and grown up, allowing for the production of cells that have the construct and are manufacturing the protein of interest. At times, these transfections are transient and all of the cells are used in the process. Other times, a new stable cell line (still a Cell Line) is produced that can continue to be used in the future.Expression System
A combination of a Construct and a host cellRelated Topics
Protein Sequence Annotations
Annotation Detail Page
To view protein sequence annotations, go to Registry > Protein Sequences, and click a protein id in the Name column. Click the Sequence tab.- The upper panel represents the sequence graphically, with different annotations called out with colored bars.
- The panel below provides annotation details in a grid view.
- The bottom left panel provides an editor with two tabs for adding and editing annotations.
- The bottom right panel offers controls of the display area
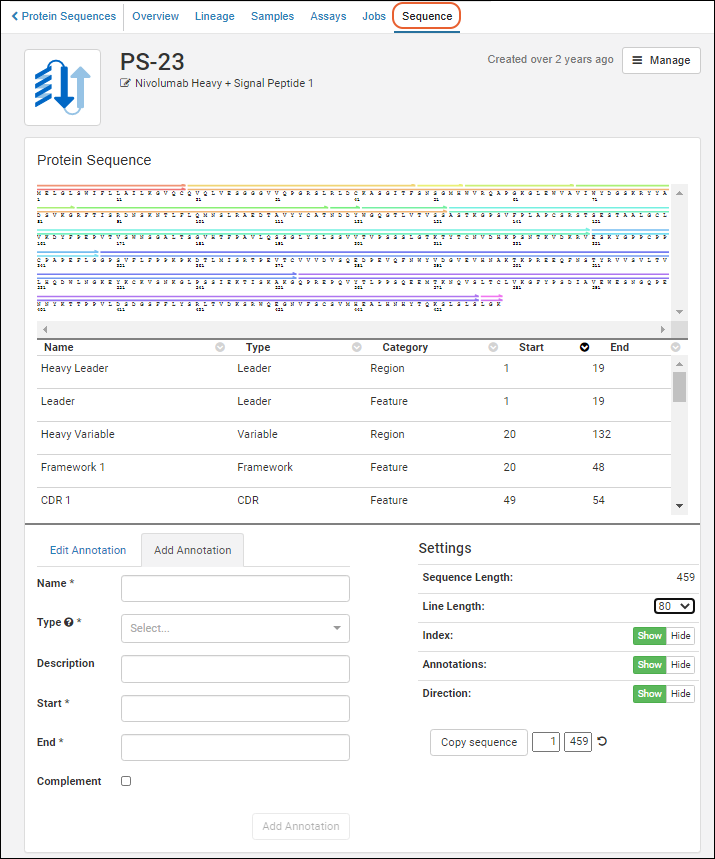 A somewhat simplified version of the annotations viewer is also used within the protein sequence registration wizard.
A somewhat simplified version of the annotations viewer is also used within the protein sequence registration wizard.
Graphical Representation
Clicking an annotation’s color bar highlights its description in the grid below, For example, if you click the green bar for sequence position 24-34, that annotation’s details are highlighted in the grid below, and vice versa.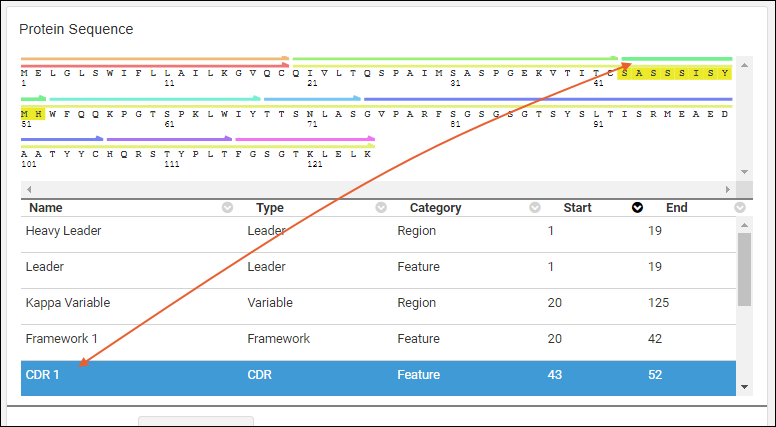
Details Grid
The annotations details grid can be sorted by any of the available columns. The grid will always default to sorting the "Start" column in ascending order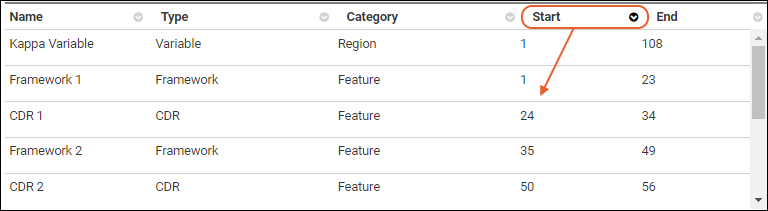
Annotation Editor
The annotation editor has two tabs: Edit Annotation and Add Annotation. Adding or editing an annotation will refresh the grid and viewer to include your changes.- Note the asterisks marking required fields on the Add Annotation tab. The button at the bottom of the form will remain grayed out until all fields have been filled out.
- The Type dropdown is pre-populated with the items in the list "AnnotationTypes".
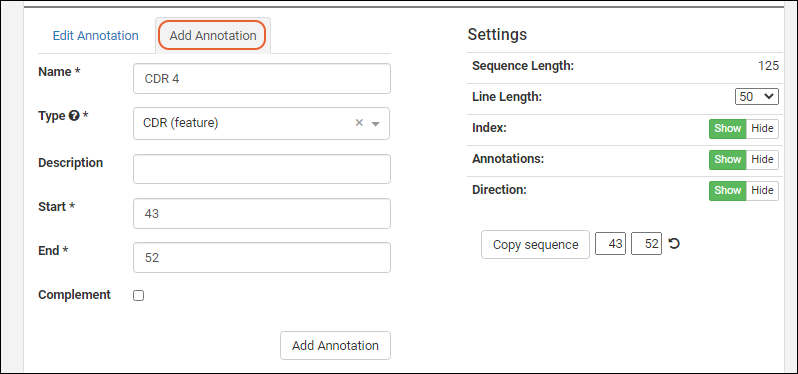 In order to edit an annotation, select one from the grid or viewer and click the Edit Annotation tab.
In order to edit an annotation, select one from the grid or viewer and click the Edit Annotation tab.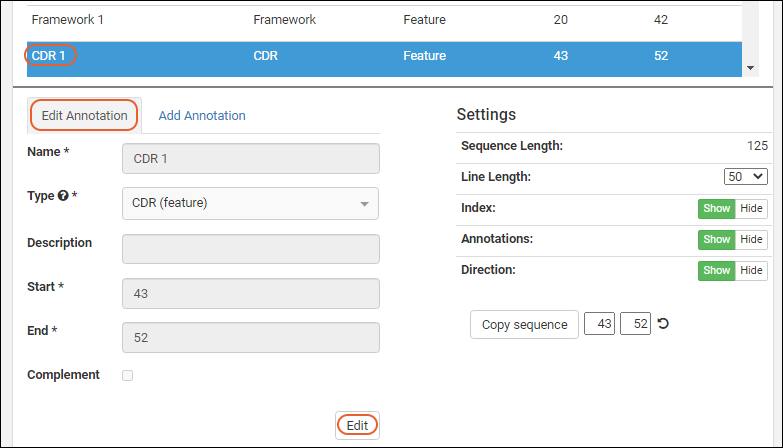 Click the Edit button to enable the field inputs and action buttons:
Click the Edit button to enable the field inputs and action buttons:
- Remove Annotation: Removes the annotation and deletes the details row from the grid.
- Cancel: Cancels the edit.
- Save: Save any changes to the selected annotation.
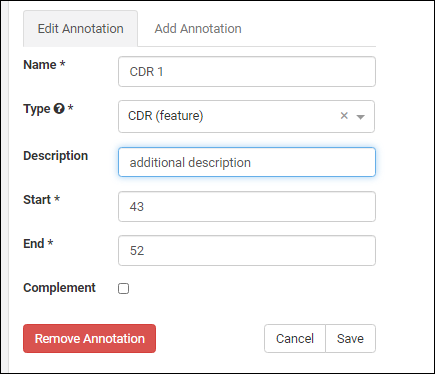
Add AnnotationType
To add a new option to the Annotation Type dropdown, such as "Protease Cleavage Site", switch to the LabKey Server interface and edit the "AnnotationTypes" list. Insert a new row, specifying the name, abbreviation and category you want to use.Settings/Controls
The controls in the bottom right panel let you customize the display region.- Sequence Length: The length of the entire sequence.
- Line Length: Select to control the display scale. Options: 50, 60, 80, 100.
- Index: Show or hide or show the position index numbers.
- Annotations: Show or hide the annotation color bars.
- Direction: Show or hide the direction indicators on the bars.
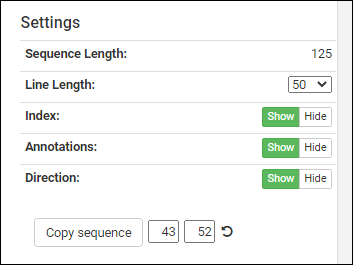 Below these controls, you can see a sequence selection widget next to the Copy sequence button. The position boxes for start and end update to the currently selected annotation. These can be manually changed to any index values in the sequence, or can be reset to the full sequence by pressing the refresh icon.Clicking Copy sequence will copy the indicated segment to your clipboard for pasting (useful when registering a new protein sequence into the Registry).
Below these controls, you can see a sequence selection widget next to the Copy sequence button. The position boxes for start and end update to the currently selected annotation. These can be manually changed to any index values in the sequence, or can be reset to the full sequence by pressing the refresh icon.Clicking Copy sequence will copy the indicated segment to your clipboard for pasting (useful when registering a new protein sequence into the Registry).
Related Topics
CoreAb Sequence Classification
- CoreAb Sequence Classification Introduction
- Classification Process
- Reference Germline Data Extraction Process
CoreAb Sequence Classification: White Paper Introduction
This methodology was published in March 2020, and is also available here:- https://just-evotecbiologics.com/white-papers-application-notes/coreab-sequence-classification/
- Author: J. Alex Taylor
- Publication: March 2020
Keywords: antibody; antibody numbering; structural numbering; antibody engineering; antibody analysis
Classification Process
The CoreAb Java library (developed at Just - Evotec Biologics) contains algorithms for the classification and alignment of antibodies and antibody-like sequences. A high-level summary of the classification process is presented in Figure 1. The first step in the classification process is the detection of antibody variable and constant regions specified in the detection settings. The default regions for detection are kappa variable, lambda variable, heavy variable, light constant, heavy constant Ig (CH1), heavy constant Fc-N (CH2), and heavy constant Fc-C (CH3). A Position-Specific Sequence Matrix (PSSM) has been pre-built for each type and is used as a low threshold first pass filter for region detection using the Smith-Waterman algorithm to find local alignments. Each local alignment is then refined by a more careful alignment comparison to the germline gene segments from species specified in the detection settings. If germline data for the query’s species of origin does not exist or is incomplete in the resource files contained in CoreAb, other, more complete, germline gene data sets from other species can be used to identify homologous regions. The germline sequences are stored as ASN-aligned so that the resulting region alignments are also ASN-aligned.To generate an alignment for variable regions, the PSSM-matched sub-sequence is aligned to both germline V-segments and J-segments and these results are combined to synthesize an alignment for the entire variable region. For heavy variable regions, the germine D-segments are aligned to the residues between the V-segment match and the J-segment match. As a final step in the variable region alignment refinement process, CDR regions are center-gapped to match the AHo/ASN numbering system. Fig. 1 High-level antibody classification pseudocode1. Identify antibody variable and constant regions (domains)
a. Loop over the region types that were specified in settings
i. Use a PSSM for the region type to find local alignments in the query
ii. Loop over each local alignment from the query
1. Loop over the germline sets that were specified in settings
a. Generate a refined region alignment for the PSSM alignment
i. Only keep alignments that meet the minimum number of identities and
region percent identity specified in settings
ii. Assign ASN numbering
iii. If variable region, refine the alignment and adjust CDR gapping
iv. If constant region and alignment is < 10 aa, toss it unless it is at
the start of the region
2. Identify potential leader region matches (can use SeqParts)
3. Resolve overlapping regions giving priority to the higher scoring region
4. Assign gaps between identified regions (can use SeqParts)
5. Cleanup constant regions
6. Assign chain and structure format (based on the arrangement of regions)
<AbFormatCache version='2'>
<ChainFormats>
...
<ChainFormat id='55' name='DVD-IgG1 Heavy Chain' abbrev='DVD-IgG HC' description='DVD-IgG1 Heavy Chain'>
{Ldr} ; HV ; Lnk ; HV ; IgG1:HCnst-Ig ; IgG1:Hinge ; IgG1:Fc-N ; IgG1:Fc-C ; IgG1:HCnst-Po
</ChainFormat>
<ChainFormat id='56' name='IgG1 Fc-Fusion' abbrev='IgG1 Fc-Fusion' description='IgG1 Fc-Fusion'>
{Ldr} ; Unk ; {Lnk}; IgG1:Hinge ; IgG1:Fc-N ; IgG1:Fc-C ; IgG1:HCnst-Po
</ChainFormat>
<ChainFormat id='57' name='IgG1 Heavy Chain F(ab')2' abbrev='IgG1 HC F(ab')2' description='IgG1 Heavy Chain F(ab')2'>
{Ldr} ; HV ; IgG1:HCnst-Ig ; IgG1:Hinge<C113,C116>
</ChainFormat>
...
</ChainFormats>
<StructureFormats>
...
<StructureFormat id='28' name='DVD-IgG1' abbrev='DVD-IgG1' description='Dual Variable Domain (DVD) Bispecific IgG1 Antibody'>
<ChainCombination>
<ChainFormat id='53' stoichiometry='2' />
<ChainFormat id='55' stoichiometry='2' />
</ChainCombination>
<ChainCombination>
<ChainFormat id='54' stoichiometry='2' />
<ChainFormat id='55' stoichiometry='2' />
</ChainCombination>
</StructureFormat>
<StructureFormat id='29' name='IgG1 Fc-Fusion' abbrev='IgG1 Fc-Fusion' description='IgG1 Fc-Fusion'>
<ChainCombination>
<ChainFormat id='56' stoichiometry='2' />
</ChainCombination>
</StructureFormat>
<StructureFormat id='30' name='IgG1 F(ab')2' abbrev='IgG1 F(ab')2' description='IgG1 F(ab')2 Fragment'>
<ChainCombination>
<ChainFormat id='5' stoichiometry='2' />
<ChainFormat id='57' stoichiometry='2' />
</ChainCombination>
<ChainCombination>
<ChainFormat id='8' stoichiometry='2' />
<ChainFormat id='57' stoichiometry='2' />
</ChainCombination>
</StructureFormat>
...
</StructureFormats>
</AbFormatsCache>
Reference Germline Data Extraction Process
The extraction and compilation of antibody germline gene data can be a difficult and time consuming process. In cases where gene annotation is provided by the NCBI, a CoreAb tool is used to extract and align the gene information. Incomplete or unannotated genomes require a more de novo approach. CoreAb also contains a tool that can scan for potential V-segments, J-segments, and D-segments using PSSMs designed to locate the Recombination Signal Sequence (RSS) sequences used to join the variable region segments. Manual curation is still required to filter and adjust the results but this automation can alleviate most of the tedious work. When possible, names for extracted genes are set to those from IMGT since that is the source of official naming. Figure 3 displays a section of an XML-formatted germline data resource file. Default XML-formatted germline data is included in CoreAb and loaded at runtime. Additional or alternate germline data can be provided by the user. Full or partial antibody gene data is currently included in CoreAb for the following organisms: Bos taurus, Camelus bactrianus, Camelus dromedarius, Canis familiaris, Cavia porcellus, Gallus gallus, Homo sapiens, Macaca mulatta, Mus musculus, Oryctolagus cuniculus, Ovis aries, Protopterus dolloi, Rattus norvegicus, Struthio camelus, and Vicugna pacos. Fig. 3 Snippet of the Bos taurus HV.xml germline data file of heavy variable genes extracted from genomic sequences.<?xml version='1.0' encoding='UTF-8' standalone='yes' ?>
<GermlineGeneSetRsrc igGeneGroup='IGHV' taxonId='9913'>
...
<GermlineGene id='IGHV1-20*01'>
<Note>Num RSS mismatches: 0</Note>
<GenomicLoc build='ARS-UCD1.2' chromosome='21' contig='NW_020190105.1' contigLength='69862954'
strand='Forward'>join(278220..278274,278353..278658)</GenomicLoc>
<DB_Xref db='Just-TempId'>IGHV1-4S*01</DB_Xref>
<DB_Xref db='IMGT/GENE-DB'>IGHV1-20*01</DB_Xref>
<Intron donorSite='a gtgtc' acceptorSite='acag g' seqLength='78'>
<GenomicLoc build='ARS-UCD1.2' chromosome='21' contig='NW_020190105.1' contigLength='69862954'
strand='Forward'>278275..278352</GenomicLoc>
<DNA>
gtgtctctgggtcagacatgggcacgtggggaagctgcctctgagcccacgggtcaccgtgcttctctctctccacag
</DNA>
</Intron>
<AlignedRegions>
<AlignedRegion region='HLdr'>
<Protein>
MNPLWEPPLVLSSPQSGVRVLS
</Protein>
<DNA>
atgaacccactgtgggaacctcctcttgtgctctcaagcccccagagcggagtgagggtcctgtcc
</DNA>
</AlignedRegion>
<AlignedRegion region='HV'>
<Protein>
QVQLRES-GPSLVKPSQTLSLTCTVSG-FSLSS-----YAVGWVRQAPGKALEWLGGISS----GGSTYYNPALKSRLSITKDNSKSQVSLSVSSVTPEDTATYYCAK-----------------------------------------
</Protein>
<DNA RSS_3prime='cacagtg aggggaaatcagtgtgagcccag acaaaaacc' splitCodon3prime='ga'>
caggtgcagctgcgggagtcgggccccagcctggtgaagccctcacagaccctctccctcacctgcacggtctctggattctcactgagcagctatgctgtaggctgggtccgccaggctccagggaaggcgctggagtggctcggtggtataagcagtggtggaagcacatactataacccagccctgaaatcccggctcagcatcaccaaggacaactccaagagccaagtctctctgtcagtgagcagcgtgacacctgaggacacggccacatactactgtgcgaagga
</DNA>
</AlignedRegion>
</AlignedRegions>
</GermlineGene>
...
</GermlineGeneSetRsrc>
Related Topics
Biologics: Chain and Structure Formats
Overview
After the classification engine has determined the pattern of regions within the sequence, the chain and structure format information that was provided is used to first match the regions to a chain pattern and then to match the assigned chain formats to a unique structure format.If antibody regions are present in a ProteinSeq, but it does not match a chain format, it will be assigned a chain format of "Unrecognized Antibody Chain" and the Molecule is assigned a structure format of "Unrecognized Antibody Format".If no antibody regions are present, the ProteinSeq is assigned a chain format of "Non-Antibody Chain" and the Molecule structure format is set to "Non-Antibody".In order for changes to the chain and structure formats to take effect, an administrator needs to clear caches from the Administration Console.Chain Formats
Chain formats are stored in the ChainFormats List.A chain format specification is specified at the region level using region abbreviations. Recognized regions are listed in the following table.| Region | Abbreviation |
|---|---|
| Leader | Ldr |
| Light Variable | KV/LmdV |
| Light Constant | KCnst/LmdCnst |
| Kappa Leader | KLdr |
| Kappa Variable | KV |
| Kappa Constant Ig Domain | KCnst-Ig |
| Post Kappa Constant Ig Present in some species as a short C-terminal tail | KCnst-Po |
| Lambda Leader | LmdLdr |
| Lambda Variable | LmdV |
| Lambda Constant Ig Domain | LmdCnst-Ig |
| Post Lambda Constant Ig Present in some species as a short C-terminal tail | LmdCnst-Po |
| Heavy Leader | HLdr |
| Heavy Variable | HV |
| Heavy Constant Ig Domain | HCnst-Ig |
| Heavy Constant Hinge | Hinge |
| Heavy Constant Fc N-terminal Domain | Fc-N |
| Heavy Constant Fc C-terminal Domain | Fc-C |
| Post Heavy Constant Ig | HCnst-Po |
| Linker | Lnk |
| Tag | Tag |
| Protease Cleavage Site | Cut |
| Unrecognized | Unk |
| TCR-alpha Leader | TRALdr |
| TCR-alpha Variable | TRAV |
| TCR-alpha Constant Ig Domain | TRACnst-Ig |
| TCR-alpha Constant Connector | TRACnst-Connector |
| TCR-alpha Constant Transmembrane Domain | TRACnst-TM |
| TCR-beta Leader | TRBLdr |
| TCR-beta Variable | TRBV |
| TCR-beta Constant Ig Domain | TRBCnst-Ig |
| TCR-beta Constant Connector | TRBCnst-Connector |
| TCR-beta Constant Transmembrane Domain | TRBCnst-TM |
| TCR-delta Leader | TRDLdr |
| TCR-delta Variable | TRDV |
Chain Format Syntax
1. Regions are separated by a semicolon. Optional regions are surrounded by braces {}.Example 1: A kappa light chain is specified as:{Ldr} ; KV ; KCnst-Ig ; {KCnst-Po}{Ldr | Unk<M>} ; (HV ; Lnk ; KV/LmdV | KV/LmdV ; Lnk ; HV){Ldr} ; HV ; IgG2:HCnst-Ig ; IgG2:Hinge ; IgG2:Fc-N ; IgG2:Fc-C ; IgG2:HCnst-Po{Ldr} ; HV ; IgG1:HCnst-Ig ; IgG1:Hinge<!113-123>{Ldr} ; HV ; IgG1:HCnst-Ig ; IgG1:Hinge ; IgG1:Fc-N ; IgG1:Fc-C<C15,W30> ; IgG1:HCnst-Po{Ldr} ; HV[3] ; IgG1:HCnst-Ig ; Unk<EPKSCD> ; Lnk ; HV[2] ; Unk<A> ; KCnst/LmdCnst ;
IgG1:Hinge<!1-107> ; IgG1:Fc-N ; IgG1:Fc-C<C15,W30> ; IgG1:HCnst-Po{Ldr} ; HV⊃VHH ; IgG1:Hinge ; IgG1:Fc-N ; IgG1:Fc-C ; IgG1:HCnst-PoStructure Formats
Structure formats are stored in the StructureFormats List where the only important information is the name and abbreviation. The more important table is ChainStructureJunction which describes the chain combinations that map to a structure format.For example, there are two chain combinations for an IgG1, 2 copies of a Kappa Light Chain + 2 copies of an IgG1 Heavy Chain or 2 copies of a Lambda Light Chain + 2 copies of an IgG1 Heavy Chain. In the ChainStructureJunction this is represented like this:| Structure Format | Chain Format | Combination | Stoichiometry | Num Distinct* | Fv Num Overrides** |
|---|---|---|---|---|---|
| IgG1 | Kappa Light Chain | 1 | 2 | 1 | |
| IgG1 | IgG1 Heavy Chain | 1 | 2 | 1 | |
| IgG1 | Lambda Light Chain | 2 | 2 | 1 | |
| IgG1 | IgG1 Heavy Chain | 2 | 2 | 1 |
Related Topics
Molecules, Sets, and Molecular Species
Definitions
Molecules are composed of various components and have a set of physical properties that describe them. When a molecule is added to the registry, the following additional entities are calculated and added, depending on the nature of the molecule. These additional entities can include:- Molecular Species
- Molecule Sets
- Other Protein Sequences
Molecule Components
A molecule can be created (= registered) based on one or more of the following:- a protein sequence
- a nucleotide sequence
- other molecules
Rules for Entity Calculation/Creation
When the molecule contains only protein sequences, then:- A mature molecular species is created, consisting of the leaderless segments of the protein sequences, provided a leader portion is identifiable. The leader segment has to be:
- Have annotation start with residue #1
- The annotation Type is "Leader"
- The annotation Category is "Region"
- (If no leader portion is identifiable, then the species will be identical to the Molecule which has just been created.)
- Additionally, a mature desK molecular species is created (provided that there are terminal lysines on heavy chains).
- New protein sequences are created corresponding to any species created, either mature, mature des-K, or both. The uniqueness constraints imposed by the registry are in effect, so already registered proteins will be re-used, not duplicated.
- A molecule set is created, provided that the mature molecular species is new, i.e., is not the same (components and sequences) as any other mature molecular species of another molecule. If there is an already existing mature species in the registry, then the new molecule is associated with that set.
- Physical properties are not calculated.
- Molecular species are not created.
- A molecule set is created, which has only this molecule within it.
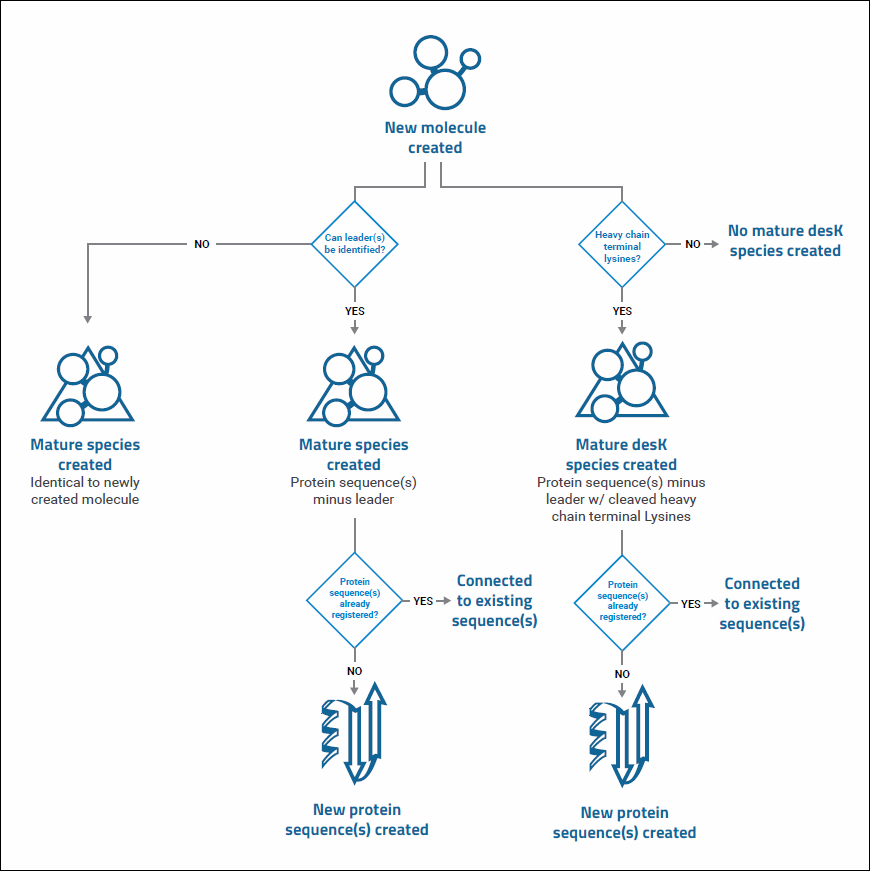
Aliases and Descriptions
When creating a Molecule, the auto-generated Molecule Set (if there is a new one) will have the same alias as the Molecule. If the new Molecule is tied to an already existing Molecule Set, the alias is appended with alias information from the new Molecule.When creating a molecule, the auto-generated molecular species (both mature and mature desK) should have the same alias as the molecule. Similarly, when creating a molecular species from a molecule (manually), the alias field will pre-populate with the alias from the molecule.For molecular species that are auto-generated, if it is creating new protein sequences (one or more) as the components of that molecular species, they have a Description:- Mature of “PS-15”
- Mature, desK of “PS-16”
- Mature of “PS-17”
- Mature, desK of “PS-18”
Related Topics
Register Molecules
Create a Molecule
To add a new molecule to the registry:- Open the Molecules creation wizard.
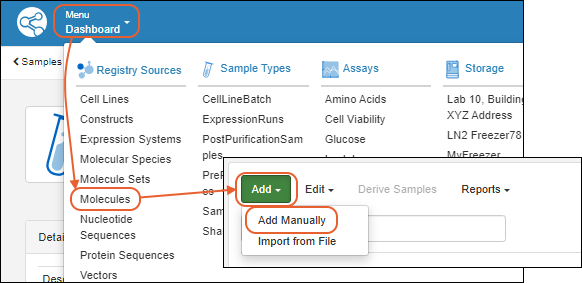
Add Details
On the first tab of the wizard, enter the following:- Name: Provide a name, or one will be generated for you. Hover to see the naming pattern
- Description: (Optional) A text description of the molecule.
- Alias: (Optional) Alternative names for the molecule.
- Common Name: (Optional) The common name of the molecule, if any.
- Molecule Parents: (Optional) Parent molecules for the new molecule.
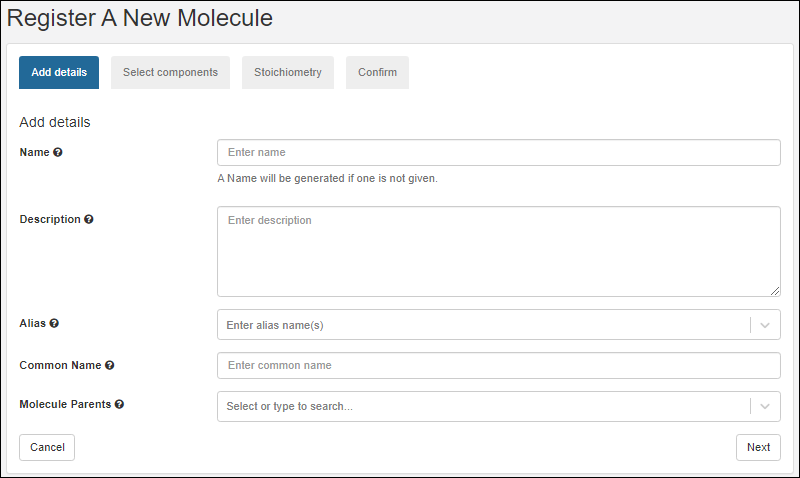 Click Next to continue.
Click Next to continue.
Select Components
- On the Select components tab, search and select existing components of the new molecule.
- After selecting the appropriate radio button, search for the component of interest.
- Type ahead to narrow the list.
- You will see a details preview panel to assist you.
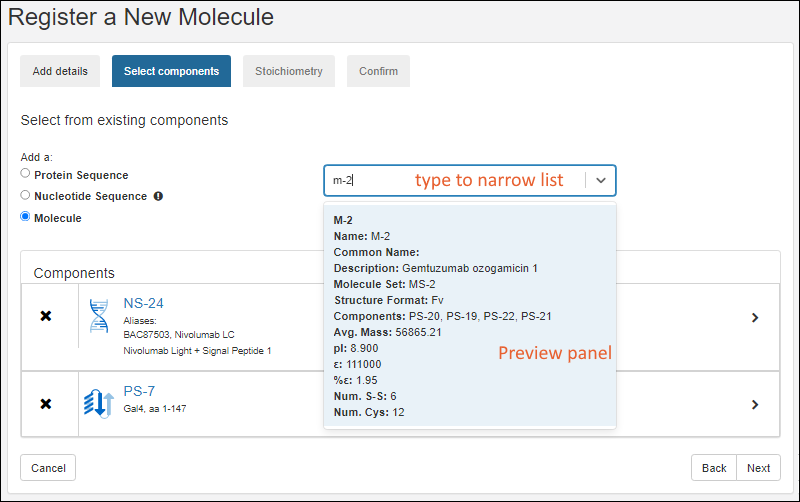
- Once you have added a component, it will be shown as a panel with entity icon. Click the to expand details.
Stoichiometry
LabKey Biologics will attempt to classify the structure format of the molecule's protein components, if possible. The structure format is based on the component protein chain formats.On the Stoichiometry pane enter:- Stoichiometry from each component
- Structure Format: Select a format from the pulldown list. The list is populated from the StructureFormat table.
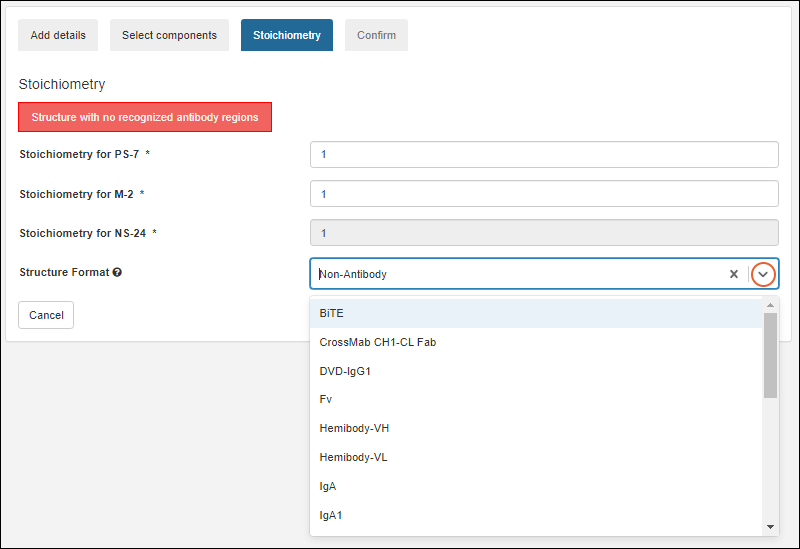 Click Next to continue.
Click Next to continue.
Confirm
On the final tab, confirm the selections and click Finish to add the molecule to the registry.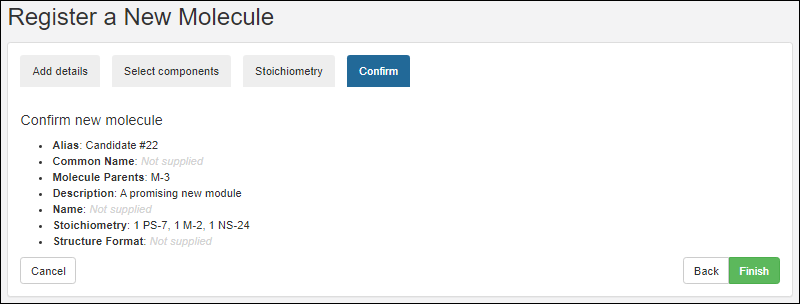 The new molecule will be added to the grid.
The new molecule will be added to the grid.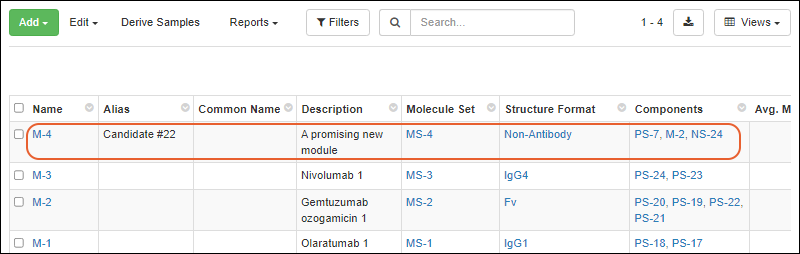
Related Topics
Molecular Physical Property Calculator
View Physical Property Calculator
From the grid of Molecules, select the Molecule of interest. Click the Physical Property Calculator tab.Calculation Inputs
In the Calculation Inputs panel, you'll enter the values to use in the calculation:- Num. S-S: Dynamically adjusted to be half of the "Num. Cys" value.
- Num Cys: A display only value derived from the sequence chosen.
- Sequence Scope: Select the desired scope. Sequence ranges will adjust based on your selection of any of the first three options. Use "Custom Range" for finer control of ranges.
- Full Protein Sequence: complete amino acid sequence of a protein, including all the amino acids that are synthesized based on the genetic information.
- Mature Protein Sequence: final, active form of the protein, after post-translational modifications and cleavages, or other changes necessary for the protein to perform its intended biological function.
- Mature Des-K Protein Sequence: mature protein sequence with the C-Terminal Lysine removed.
- Custom Range
- Sequence Ranges for each component sequence. These will adjust based on the range selected using radio buttons, or can be manually set using the "Custom Range" option.
- Use Range: Enter the start and end positions.
- Click View this sequence to open the entire annotated sequence in a new tab for reference.
- Stoichiometry
- Analysis Mode. Select from:
- Native
- Reduced
- Alkylated Cysteine (only available in "Reduced" mode). If available, select the desired value.
- Modifiers
- Pyro-glu: Check the box for "Cyclize Gln (Q) if at the N-terminus"
- PNGase: Check the box for "Asn (N) -> Asp (D) at N-link sites"
Properties
When you click Calculate, the right hand panel of Properties will be populated. You'll see both what the Classifier Generated value is (if any) and the Simulated value using the inputs you entered.Calculations are provided for:- Mass
- Average Mass
- Monoisotopic Mass
- Organic Average Mass
- pI - Isoelectric Point calculated by different methods:
- Bjellqvist
- EMBOSS
- Grimsley
- Patrickios (simple)
- Sillero
- Sillero (abridged)
- Other
- Chemical Formula
- Extinction Coefficient - ε
- Percent Extinction Coefficient
- Sequence Length
- Amino Acid (AA) composition
Export Calculations
To export the resulting calculated properties, click Export Data.The exported Excel file includes both the calculated properties and the inputs you used to determine them. For example, the export of the above-pictured M-17 calculation would look like this:Related Topics
Compounds and SMILES Lookups
SMILES Lookup Field
The Compound registry source type uses a custom SMILES field type only available in the Biologics module for this specific registry source. This datatype enables users to provide a SMILES string, e.g. "C1=CC=C(C=C1)C=O", that will return a 2D structure image, molecular weight and other computed properties.The SMILES string is used to search the Java library CDK ("Chemistry Development Kit") , a set of open source modular Java libraries for Cheminformatics. This library is used to generate the Structure2D image and calculate masses.Create/Import Compounds
New Compounds can be created manually or via file import. It's easier to get started understanding the lookup process by creating a single compound in the user interface.Create Compound: Carbon Dioxide
As an example, you could create a new compound, supplying the SMILES string "O=C=O" and the Common Name "carbon dioxide".Import File of SMILES Strings
When a file of SMILES strings is Created/imported, each is used to query for the respective 2D structure, molecular weight and set of computed properties. During the Create/Import operation if Name/ID isn’t specified, the SMILES string is used for Name.Related Topics
- Create Registry Sources
- Biologics: Bioregistry
- CDK (Chemistry Development Kit) : Open Source modular Java libraries for Cheminformatics
Entity Lineage
View Lineage
Any parents and children of an Registry Source or Sample can be viewed by clicking the Lineage tab on the details page. The main lineage dashboard shows a graphical panel on the left and details on the right.Two lineage views are available: (1) a graphical tree and (2) a grid representation.
The main lineage dashboard shows a graphical panel on the left and details on the right.Two lineage views are available: (1) a graphical tree and (2) a grid representation.
Lineage Graph
The graphical tree shows parents above and children below the current entity. Click any node in the tree to see details on the right. Hover for links to the overview and lineage of the selected node, also known as the seed.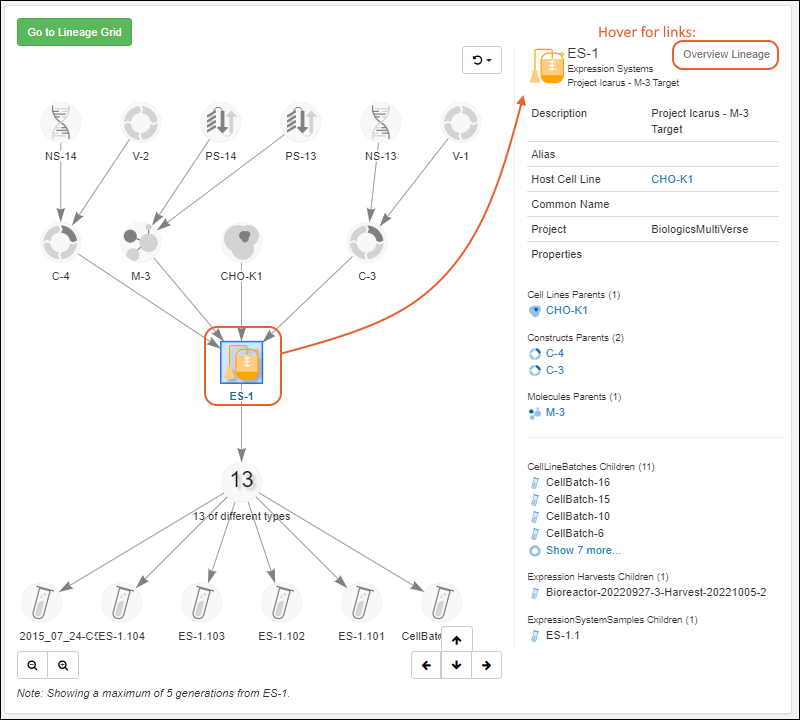
- Zoom out and in with the and buttons in the lower left.
- Step up/down/left and right within the graph using the arrow buttons.
- Up to 5 generations will be displayed. Walk up and down the tree to see further into the lineage.
- Click anywhere in the graph and drag to reposition it in the panel.
- Refresh the image using the button. There are two options:
- Reset view and select seed
- Reset view
Graph Filters for Samples
When viewing lineage for Samples, you will have a filter button. Use checkboxes to control visibility in the graph of derivatives, source parents, and aliquots.Lineage Grid
Click Go to Lineage Grid to see a grid representation of lineage. The grid view is especially helpful for viewing lengthy lineages/derivation histories. Control visibility of parents and children in the grid view using Show Parents and Show Children respectively.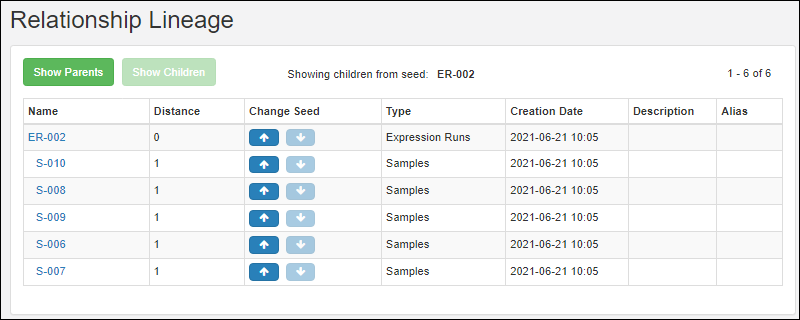 Items in the Name column are clickable and take you to the details page for that entity.Arrow links in the Change Seed column reset the grid to the entity in the Name column, showing you the parents or children for that entity.
Items in the Name column are clickable and take you to the details page for that entity.Arrow links in the Change Seed column reset the grid to the entity in the Name column, showing you the parents or children for that entity.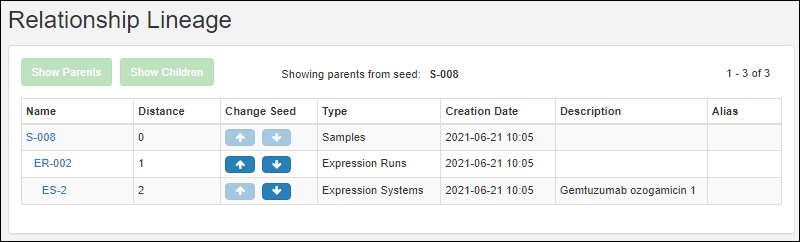
Troubleshooting
Note that if any entity names that just strings of digits, you may run into issues if those "number-names" overlap with row numbers of other entities of that type. In such a situation, when there is ambiguity between name and row ID, the system will presume that the user intends to use the value as the name.Related Topics
Customize the Bioregistry
- Definitions
- Edit Existing Registry Source Types
- Create New Registry Source Type
- Lineage for Custom Registry Source Types
- Delete Custom Registry Source Type
Definitions
The term Registry Source Type applies to all of the following entities in the Biologics application. These are two of the categories of Data Class available in LabKey Server. Within the Biologics application you cannot see or change the category, though if you access these structures outside the application you will be able to see the category.- Registry Source Types: (Category=registry)
- CellLine
- Construct
- ExpressionSystem
- MolecularSpecies
- Molecule
- MoleculeSet
- NucSequence
- ProtSequence
- Vector
- Media Types: (Category=media)
- Ingredients
- Mixtures
Edit Existing Registry Source Types
The default set of Registry Source and Media Types in Biologics are designed to meet a common set of needs for properties and fields for the Bioregistry and Media sections.If you want to customize these designs, you can edit them within the Biologics application following the same interface as used for Sample Manager "Sources".Open the desired type from the main menu. All entries under Registry Sources and Media are editable. Select Manage > Edit [Registry Source Type] Design.Create New Registry Source Type
From the main menu, click Registry Sources. Click Create Source Type.- For example, to use "DC-" followed by the date portion of when the entity is added, plus an incrementing number you could use:
DC-${now:date}-${genId}
Lineage for Custom Registry Source Types
The lineage relationships among the built-in Registry Source Types are pre-defined. Learn more in this topic: Biologics: TerminologyWhen you create your own Custom Registry Source Types, you can include Parent Aliases in the definition, making it possible to define your own lineage relationships.To add a new lineage relationship, either define it when you create the new Registry Source Type or edit it later.Delete Custom Registry Type
You cannot delete the built in Registry Source Types, but if you add a new custom one, you will have the ability to delete it. Before deleting the type, you may want to delete all members of the type. If any members are depended upon by other entities, deletion will be prevented, allowing you to edit to change the connections before deleting the type.- Select the custom type to delete from the Registry section of the main menu.
- Select Manage > Delete [Registry Source Type].
Related Topics
Bulk Registration of Entities
Supported File Formats
The file formats supported are listed in the file import UI under the drop target area.Tabular file formats are supported for all entity types:- Excel: .xls, .xlsx
- Text: .csv, .tsv
- GenBank: .genbank, .gb, .gbk
Assemble Bulk Data
When assembling your entity data into a tabular format, keep in mind that each Registry Source Type has a different set of required column headings.- Acquire a template Excel file for a given entity by clicking Template from the Registry Source Type overview page.
- Open the Excel file, and add data as appropriate under the column headers. See examples below.
Indicate Lineage Relationships
Lineage relationships (parentage) of entities can be included in bulk registration by adding "DataInputs/<DataClassType>" columns and providing parent IDs.For example, to include a Vector as a 'parent' for a bulk registered set of Expression Systems, after obtaining the template for Expression Systems, add a new column named "DataInputs/Vector" and provide the parent vector name for each row along with other fields defining the new Expression Systems.Bulk Upload Registry Source Data
After you have assembled your information into a table, you can upload it to the registry:- Go the Registry Source Type you wish to import.
- Select Add > Import from File.
- On the import page, you can download a template if you don't have one already, then populate it with your data.
- Confirm that the Source Type you want is selected, then drag and drop your file into the target area and click Import.
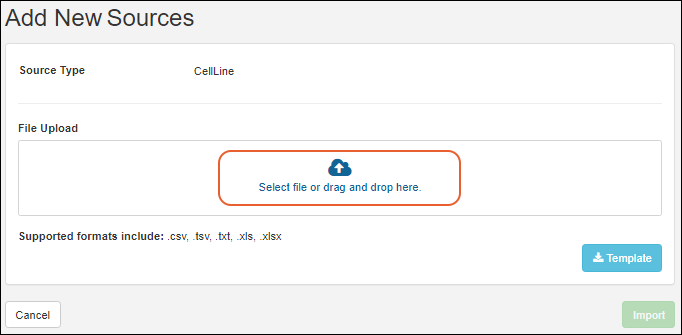
Bulk Data Example Files
Example Nucleotide Sequence File
Notes:- Annotations: Add annotation data using a JSON snippet, format is shown below.
| name | alias | description | flag | protSequences | sequence | annotations |
|---|---|---|---|---|---|---|
| NS-23 | Signal Peptide 1 | An important sequence. | FALSE | [{name: "PS-23"}] | CCCCTCCTTG GAGGCGCGCA ATCATACAAC CGGGCACATG ATGCGTACGC CCGTCCAGTA CGCCCACCTC CGCGGGCCCG GTCCGAGAGC TGGAAGGGCA | [ |
| name | alias | description | flag | protSequences | sequence | annotations |
|---|---|---|---|---|---|---|
| NS-100 | Signal Peptide 1 | some description | FALSE | [{name: "PS-100", nucleotideStart=1, nucleotideEnd=30, translationFrame=2}] | ATGGAGTTGGGACTGAGCTGGATTTTCCTTTTGGCTATTTTAAAAGGTGTCCAGTGT |
Example Protein Sequence File
Notes:- Organisms: A comma separated list of applicable organisms. The list, even if it has only one member, must be framed by square brackets. Examples: [human] OR [human, rat, mouse]
- ?: The column header for the extinction coefficient (ε).
- %?: The column header for the % extinction coefficient (%ε).
| Name | Alias | Description | Nuc Sequences | Chain Format | Avg. Mass | pI | ? | %? | Num. S-S | Num. Cys | Organisms | Sequence |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| PStest-150 | Test sequence for import | 1 | 13999.64 | 8.030 | 35500 | 2.54 | 1 | 2 | [mouse, rat] | EVQLVESGEL IVISLIVESS PSSLSGGLVQ GGGSLRLSCA ASGELIVISL IVESSPSSLS YSFTGHWMNW VRQAPGKGLE WVGIMIHPSD SETRYNQKFK DELIVISLIV ESSPSSLSIR FTISVDKSKN TLYLQMNSLR AEDTAVYYCA RIGIYFYGTT YFDYIWGQGT |
Mixtures and Batches
The text 'unknown' can entered for certain fields. For Mixtures, the Amount field; for Mixture Batches, the Amount and the RawMaterial fields.Mixture Bulk Upload| Type | Ingredient/Mixture | Amount Unit Type |
|---|---|---|
| Ingredient | I-2 | unknown |
| Ingredient | Amount Used | Raw Material Used |
|---|---|---|
| Sodium phosphate dibasic anhydrous | 5 | RawMat-1234 |
| Sodium Chloride | unknown | unknown |
| Potassium chloride | unknown | unknown |
Related Topics
Use the Registry API
Identity Service
When registering a sequence or molecule, use the identity of the sequence to determine uniqueness. External tools can use the following API to get the identity of a sequence prior to registration. To get the identity of a sequence use either the identity/get.api or identity/ensure.api. The ensure.api will create a new identity if the sequence hasn't been added yet.To get or ensure the identity of a single nucleotide sequence:LABKEY.Ajax.request({
url: LABKEY.ActionURL.buildURL("identity", "get.api"),
jsonData: {
items: [{
type: "nucleotide", data: "GATTACA"
}]
}
});LABKEY.Ajax.request({
url: LABKEY.ActionURL.buildURL("identity", "get.api"),
jsonData: {
items: [{
type: "nucleotide", data: "GATTACA"
},{
type: "protein", data: "ELVISLIVES"
}]
}
});LABKEY.Ajax.request({
url: LABKEY.ActionURL.buildURL("identity", "get.api"),
jsonData: {
items: [{
type: "molecule", items: [{
type: "protein", data: "MAL", count: 2
},{
type: "protein", data: "SYE", count: 3
]}
}]
}
});Manual Data Entry
You can enter molecules, sequences, cell lines, etc., using registration wizards within the Biologics application. For an example use of the wizard, see Register Nucleotide SequencesCut-and-Paste or Import from a File
The LabKey import data page can also be used to register new entities. For example, to register new nucleotide sequences:- Go to the list of all entity types (the DataClasses web part).
- Click NucSequence.
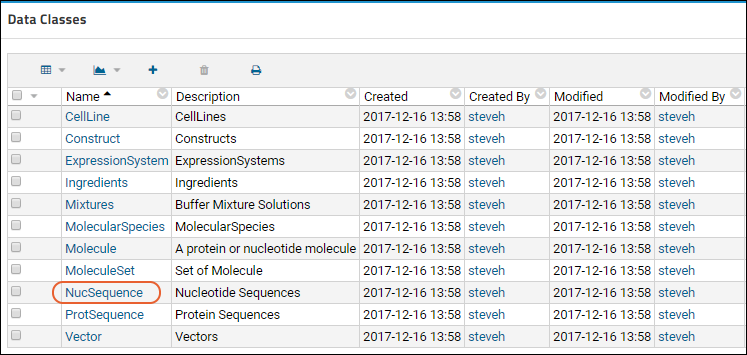
- In the grid, select > Import Bulk Data.
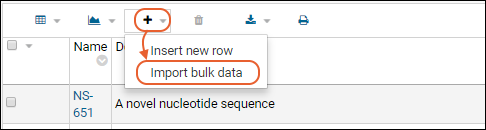
- Click Download Template to obtain the full data format.
- Upload an Excel file or paste in text (select tsv or csv) matching the downloaded template. It might be similar to:
| Description | protSeqId | translationFrame | translationStart | translationEnd | sequence |
|---|---|---|---|---|---|
| Anti_IGF-1 | PS-7 | 0 | 0 | 0 | caggtg... |
- Go to the list of all entity types
- Click ProtSequence
- In the grid, select > Import Bulk Data.
- Click Download Template to obtain a template showing the correct format.
- Paste in a TSV file or upload an Excel file in the correct format. It might be similar to:
| Name | Description | chainFormatId | organisms | sequence |
|---|---|---|---|---|
| PS-1 | PS1038-1 | 1 | ["human","mouse"] | MALWMRL... |
- Go to the list of all entity types
- Click Molecule
- In the grid, select > Import Bulk Data.
- Paste in data of the format:
| Description | structureFormatId | seqType | components |
|---|---|---|---|
| description | 1 | 1 | [{type: "nucleotide", name: "NS-1", stoichiometry: 3}] |
| description | 1 | 3 | [{type: "protein", name: "PS-1", stoichiometry: 2}, {type: "protein", ident: "ips:1234"}] |
| description | 1 | 3 | [{type: "chemical", name: "CH-1"}, {type: "molecule", name: "M-1"}] |
Register via Query API
The client APIs also can be used to register new entities.Register Nucleotide Sequence
From your browser's dev tools console, enter the following to register new nucleotide sequences:LABKEY.Query.insertRows({
schemaName: "exp.data",
queryName: "NucSequence",
rows: [{
description: "from the client api",
sequence: "gattaca"
},{
description: "another",
sequence: "cattaga"
}]
});Register Protein Sequence
To register new protein sequences:LABKEY.Query.insertRows({
schemaName: "exp.data",
queryName: "ProtSequence",
rows: [{
name: "PS-100",
description: "from the client api",
chainFormatId: 1,
sequence: "ML"
}]
});Register Molecule
To register new molecules:LABKEY.Query.insertRows({
schemaName: "exp.data",
queryName: "Molecule",
rows: [{
description: "from the client api",
structureFormatId: 1,
components: [{
type: "nucleotide", name: "NS-202"
}]
},{
description: "another",
structureFormatId: 1,
components: [{
type: "protein", name: "PS-1"
},{
type: "protein", name: "PS-100", count: 5
}]
}]
});Lineage, Derivation, and Samples
Parent/child relationships within an entity type are modeled using derivation. For example, the Lineage page for this nucleotide sequence (NS-3) shows that two other sequences (NS-33 and NS-34) have been derived from it.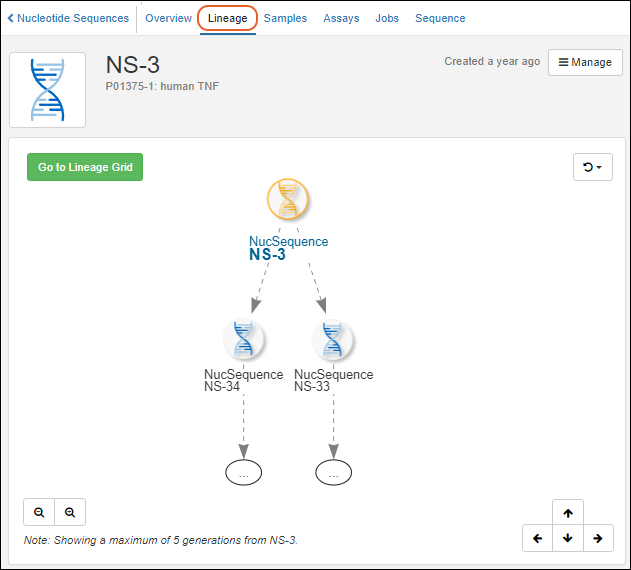 To create new children, you can use the "experiment/derive.api" API, but it is still subject to change. The dataInputs is an array of parents each with an optional role. The targetDataClass is the LSID of the entity type of the derived datas. The dataOutputs is an array of children each with an optional role and a set of values.
To create new children, you can use the "experiment/derive.api" API, but it is still subject to change. The dataInputs is an array of parents each with an optional role. The targetDataClass is the LSID of the entity type of the derived datas. The dataOutputs is an array of children each with an optional role and a set of values.LABKEY.Ajax.request({
url: LABKEY.ActionURL.buildURL("experiment", "derive.api"),
jsonData: {
dataInputs: [{
rowId: 1083
}],
targetDataClass: "urn:lsid:labkey.com:DataClass.Folder-5:NucSequence",
dataOutputs: [{
role: "derived",
values: {
description: "derived!",
sequence: "CAT"
}
}]
}
})LABKEY.Ajax.request({
url: LABKEY.ActionURL.buildURL("experiment", "derive.api"),
jsonData: {
dataInputs: [{
rowId: 1083
}],
targetSampleType: "urn:lsid:labkey.com:SampleType.Folder-5:Samples",
materialOutputs: [{
role: "sample",
values: {
name: "new sample!",
measurement: 42
}
}]
}
})Register Parents/Inputs
To indicate parents/inputs when registering an entity, use the columns "DataInputs/<DataClassName>" or "MaterialInputs/<SampleTypeName>". The value in the column is a comma separated list of values in a single string. This works for both DataClass and SampleType:LABKEY.Query.insertRows({
schemaName: "exp.data",
queryName: "MyDataClass",
rows: [{
name: "blush",
"DataInputs/MyDataClass": "red wine, white wine"
}]
});LABKEY.Query.insertRows({
schemaName: "samples",
queryName: "SamplesAPI",
rows: [{
name: "red wine"
},{
name: "white wine"
},{
name: "blush",
"MaterialInputs/SamplesAPI": "red wine, white wine"
}]
});Related Topics
Biologics: Plates
Plates Dashboard
Click Plates (under Sample Types) on the main menu. You can also go directly to Plate Sets or Plate Templates from here.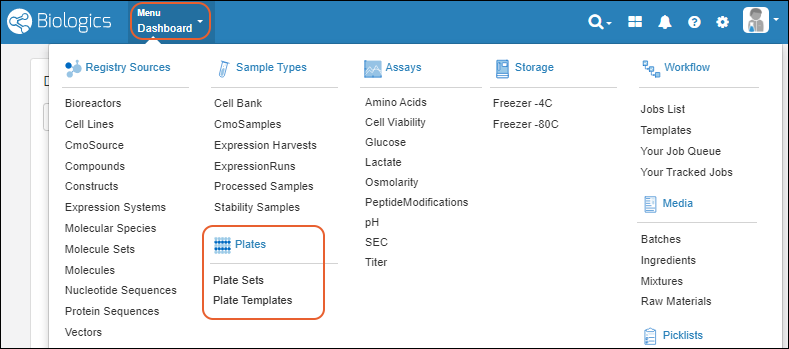
Plate Sets
- Create Plate Set
- Manage > Archive
- Manage > Plate Templates
Plate Templates
Plate templates let you set out some common features and well roles for given campaigns.Related Topics
Biologics: Assay Data
Documentation
Learn more about using the general assay framework in the Sample Manager documentation here:Additional Features with LabKey LIMS
Additional Features with Biologics LIMS
Biologics: Specialty Assays
Create a New Assay Design
To create a new assay design in Biologics LIMS, follow this process. Make sure to start from the container where you want to define the assay.- From the main menu, click Assays, then click Create Assay Design.
Select the Assay Type
- Select the Assay Type. For most cases, leave the Standard Assay tab selected.
- Learn more about using Specialty Assays in this topic: defineAssaySchema.
- Click Choose Standard Assay.
- Creating a standard assay in Biologics is the same as described in the Sample Manager documentation here:
- Creating specialty assays follows a similar process. Specific properties of the specialty assays can be found in this section:
Related Topics
Biologics: Assay Integration
Connect Assays with Samples
To associate assay data with corresponding samples, include a field of type Sample in the run or result fields. You can name the field as you like and decide whether to allow it to be mapped only to a specific sample type, or to "All Samples".Learn more in the Sample Manager documentation.Connect Assays with Other Entities
To add Entity fields to an assay design, add a field of type Lookup that points to the desired Entity/DataClass. For example, the screenshot below shows how to link to Molecules.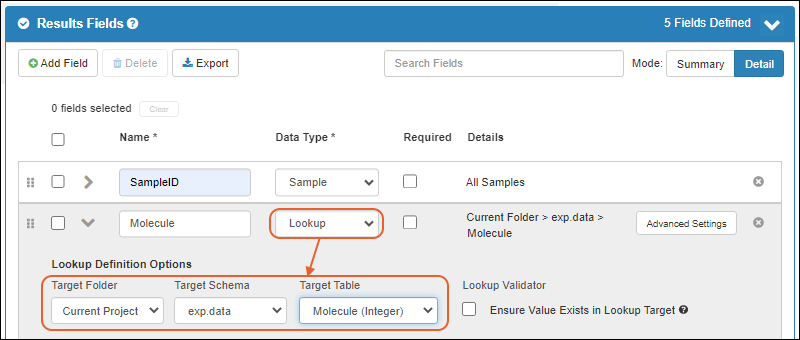
Create Integrated Grid Views
Once the assay design includes a lookup to a Sample or another Entity type (such as a Molecule), you can add other related fields to the assay design using Views > Customize Grid View.Learn about customizing and using grid views in the Sample Manager documentation.Related Topics
Biologics: Upload Assay Data
- Import Assay Data
- Import Assay Data Associated with Selected Samples
- Re-import a Run
- Delete Assay Runs
Import Assay Data
From the main menu, under Assays, click the name of the assay design you wish to import into.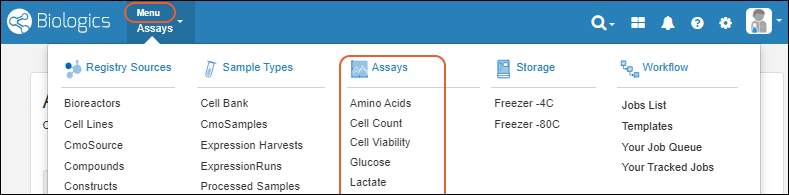 Follow the same general process as for importing assay data into Sample Manager described here. Some additional notes apply:
Follow the same general process as for importing assay data into Sample Manager described here. Some additional notes apply:
- Batch Details: If the assay design includes batch fields, you will be prompted to enter values for them.
- When importing Results, if your assay design includes File fields, you can simultaneously upload the referenced files in a second panel of the file upload tab.
Import Assay Data Associated with a Sample
An alternative way to import assay data lets you directly associate assay data with some sample(s) in the registry.From the Samples grid, select one or more samples, and then select Assay (on narrower browsers this will be a section of the More > menu). Expand the Import Assay Data section, then click the [Assay Design Name]. Scroll to select, or type into the filter box to narrow the list.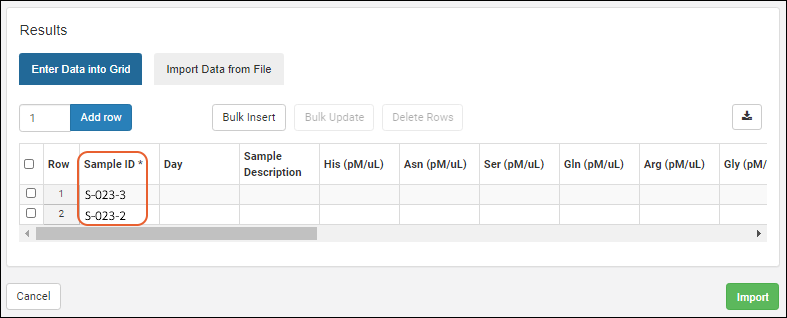 Enter the necessary values and click Import.
Enter the necessary values and click Import.
Re-Import Run
In cases where you need to revise assay data after uploading to the server, you can replace the data by "re-importing" a run.To re-import a run of assay data:- Go the details page for the target run. (For example, go to the runs table for an assay, and click one of the runs.)
- Click the Manage menu and select Re-import Run.
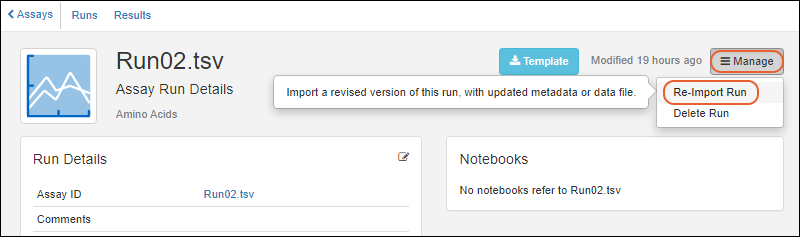
- For instrument-specific assay designs (such as NAb and Luminex), you will be directed to the specific assay’s import page in LabKey Server. Follow the procedure for re-import as dictated by the assay type: Reimport NAb Assay Run or Reimport Luminex Run.
- For general purpose assay designs, you will stay within the Biologics application:
- Revise batch and run details as needed.
- Enter new Results data. The first few lines of the existing results are shown in a preview. Choose Import Data from File or Enter Data Into Grid as when creating a new run.
- Click Re-Import.
- The data replacement event is recorded by the "Replaced By" field.
Delete Assay Runs
To delete one or more assay runs, you can:- Start from the run details and select Manage > Delete Run.
- From the grid of Runs for the assay, select the run(s) to delete, then choose Edit > Delete.
Related Topics
Biologics: Assay Batches and QC
Assay Batch Fields
Click the name of an Assay Design to see it's summary page, with several tabs, showing various scopes of the data. In addition to Runs and Results, assays in Biologics LIMS may also have fields (and a tab) for Batches of runs.Assay Quality Control
Visibility of assay data in Biologics can be controlled by setting quality control states. While an admin can edit assay designs to use QC states within the Biologics application, the actual states themselves cannot be configured in the application.Set Up QC States (Administrator)
To configure states, an admin must:- Open the Assay Design where you want to use QC. Select Manage > Edit Design.
- Confirm that the QC States checkbox is checked. If not, check it and click Finish Updating [Assay Design Name].
- Use > LabKey Server > [current Biologics folder name] to switch interfaces.
- Configure the available QC states in the system, using this topic: assayQCconfig.
- Select > Manage Assays.
- Click the name of your assay design.
- Select QC State > Manage states.
- Define the states you want to use.
- Once complete, use > LabKey Biologics > [current Biologics folder name] to return to Biologics.
Use QC States (Users)
Once QC states have been configured in the system, users (with the appropriate roles) can assign those states to assay run data. Users can update QC states in the following cases:- When a user has both Reader and QC Analyst roles
- When a user has either Folder-, Project-, or Site Administrator roles
- Navigate to the assay run view page. (Cell Viability runs are shown below.)
- Select one or more runs.
- Select Edit > Update QC States.
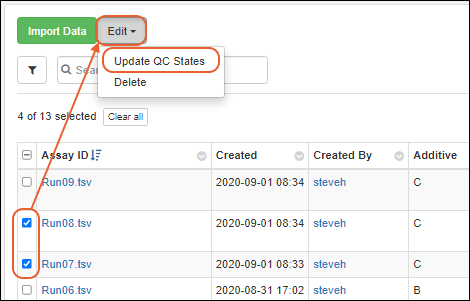
- In the popup dialog, use the dropdown menu to select the QC state. This state will be applied to all of the runs selected.
- Optionally add a comment.
- Click Save changes.
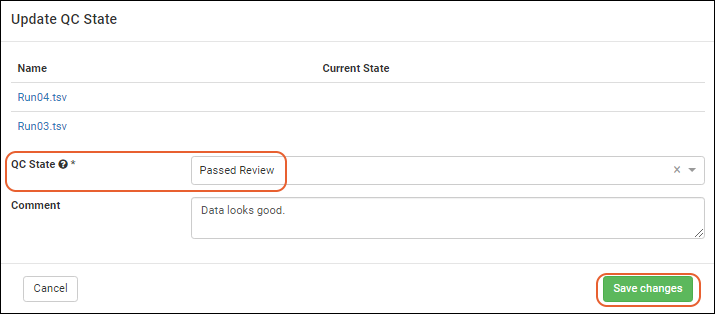
- The QC states will be reflected in the runs data grid. Admins and QC Analysts will see all of the runs, as shown below.
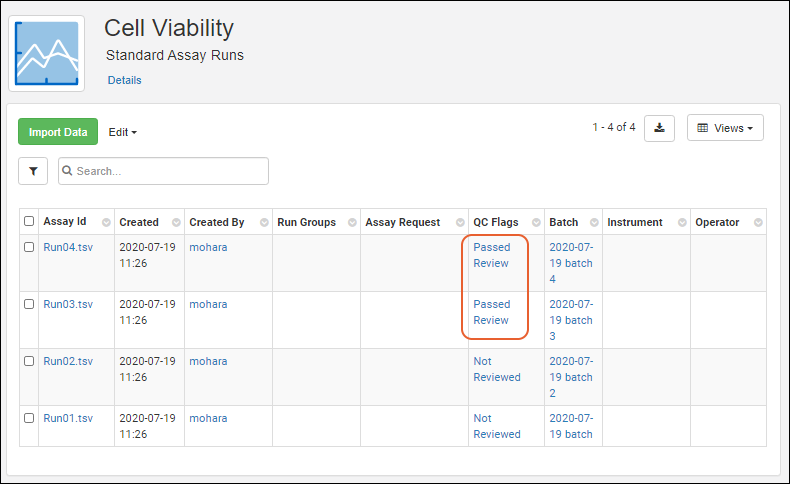
- It should be noted that if some of the QC states are not marked as "Public Data", they will not be included in the grid when viewed by non-admin users. For example, if "Not Reviewed" were marked as not public, the reader would see only the runs in public states:
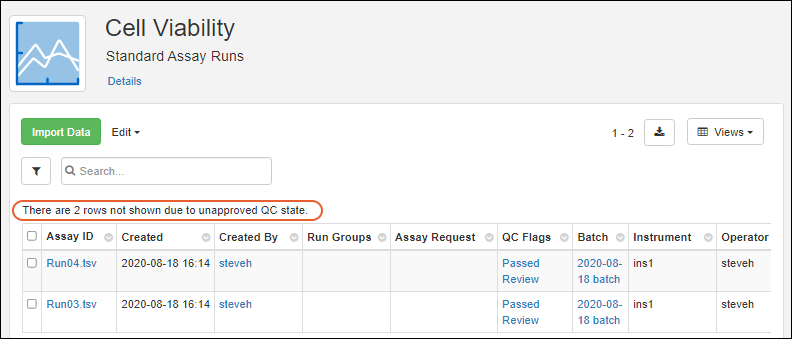
Related Topics
Biologics: Media Registration
Definitions
- Ingredients are virtual entities in LabKey Biologics, and capture the fixed natural properties of a substance. For example, the Ingredient "Sodium Chloride" includes its molecular weight, melting and boiling points, general description, etc. You register an Ingredient like this only once. Ingredients are managed with an interface similar to Registry Sources.
- Raw Materials are the particular physical instantiations of an Ingredient as real "samples" or "bottles". You register multiple bottles of the Raw Material Sodium Chloride, each with different amounts, sources, lot numbers, locations, vessels, etc. Raw Materials are managed using the Sample Type interface.
- Mixtures are recipes that combine Ingredients using specific preparation steps. Mixtures are virtual entities in LabKey Biologics. Each Mixture is registered only once, but are realized/instantiated multiple times by Batches. Mixtures are managed with an interface similar to Registry Sources.
- Batches are realizations of a Mixture recipe. They are physically real formulations produced by following the recipe encoded by some Mixture. Multiple Batches of the same Mixture can be added to the registry, each with its own volume, weight, vessel, location, etc. Batches are managed using the Sample Type interface.
Deletion Prevention
Media entities of any type cannot be deleted if they are referenced in an Electronic Lab Notebook. On the details page for the entity, you will see a list of notebooks referencing it.Topics
Managing Ingredients and Raw Materials
Definitions
In the LabKey Biologics data model, "Ingredients" are definitions, "Raw Materials" are physical things.- Ingredients are virtual entities that describe the properties of a substance. For example, the Ingredient "Sodium Chloride" includes its molecular weight, melting and boiling points, general description, etc. You register an Ingredient like this only once. Defining and creating ingredients uses the Sources UI.
- Raw Materials are physical instances of an Ingredient, tangible things that have a location in storage, with specified amounts, sources, lot numbers, locations, vessels, etc. Defining and creating raw materials uses the Samples UI.
- Mixtures are definitions. Recipe definitions, instructions for combining Ingredients. Mixtures are defined using a wizard but otherwise similar in structure and menus to Sources.
- Batches are physical things, what you get when you combine Raw Materials according to some Mixture recipe. Batches are defined using a wizard but otherwise similar in structure and menus to Samples.
Ingredients
Clicking Ingredients brings you to a grid of available (previously created) ingredients. Click Template to obtain an Excel template with the expected columns. Use the Add > menu to create new ingredients, with options similar to those for Sources.Work with Ingredients
Other actions from the grid of all ingredients are:- Edit: Select one or more ingredient rows, then choose whether to:
- Edit in Grid
- Edit in Bulk
- Delete: Note that ingredients cannot be deleted if they have derived sample or batch dependencies, or are referenced in notebooks.
- Derive Samples:
- Select one or more parent ingredients, click Derive Samples, then choose which type of sample to create.
- Reports:
- Find Derivatives in Sample Finder: Select one or more ingredients and choose this report option to open the Sample Finder filtered to show samples with the selected ingredient components. From there you can further refine the set of samples.
Ingredient Details
Click the name of an ingredient to see the details page. On the overview you can see values set for properties of the ingredient, and if any ELNs reference this ingredient, you will see links to them under Notebooks.Using the Create menu, you can add new Mixtures or Raw Materials with this Ingredient as a parent.Raw Materials
The raw materials used in mixtures are listed on the Raw Materials tab.Aliquot Raw Materials
As for samples, you can create aliquots of Raw Materials, or use certain Raw Materials to create derived samples or pooled outputs. Select the desired parent material(s) and choose Derive > Aliquot Selected or Derive or Pool as desired.Learn more in the Sample Manager documentation here: When users are selecting raw materials from a dropdown, you can use identifying fields to customize what information is shown.Deletion Prevention
Media entities of any type cannot be deleted if they are referenced in an Electronic Lab Notebook. On the details page for the entity, you will see a list of notebooks referencing it.Related Topics
Registering Mixtures (Recipes)
- Registering Mixtures
- Register a Mixture: Wizard Steps
- Register a Mixture using 'Bulk' Ingredients Table
- Registering Mixtures with Unknown Ingredients, Amounts, or Concentrations
- Delete a Mixture
- Use the Recipe API to Update Mixtures
Definitions
- Mixtures are recipes that combine Ingredients using specific preparation steps. Mixtures are virtual entities in LabKey Biologics. Each Mixture is registered only once, but are realized/instantiated multiple times by Batches.
- Batches are realizations of a Mixture recipe. They are physically real formulations produced by following the recipe encoded by some Mixture. Multiple Batches of the same Mixture can be added to the registry, each with its own volume, weight, vessel, location, etc.
Registering Mixtures
The virtual recipes are listed in the Media > Mixtures grid. Find a given Mixture by filtering, sorting, or searching the grid.- Create a new mixture "from scratch"
- Create a new mixture starting from an existing mixture or ingredient
- Registering Mixtures with Unknown Ingredients, Amounts, or Concentrations.
- Registering Batches with Unknowns.
Start a New Mixture "From Scratch"
From the main menu, select Media, then Mixtures and then click Add above the grid of available mixtures.
Start from an Existing Ingredient or Mixture
Any of the registered ingredients or mixtures can be clicked for a detailed view. The details view includes general information. Mixture detail pages show the mixture’s included ingredients and preparation steps.Select Manage > Create Mixture. The Ingredients tab will be prepopulated.
Register a Mixture: Wizard Steps
The new mixture wizard adds a new mixture to the registry, and performs a check to ensure that the mixture name is unique in the registry. Duplicate mixture names are not allowed in the registry.Wizard Step 1: Details
The Details step asks for basic information about the mixture, including a type such as powder or solution. Once all required fields have been filled in, the Next button will become clickable.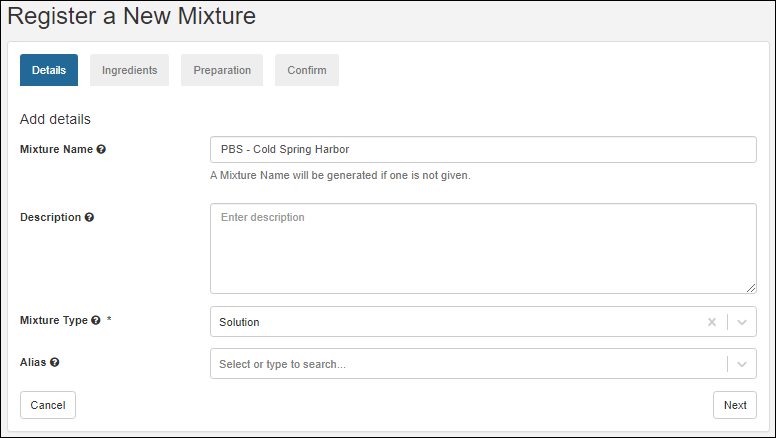 Upon clicking Next, the mixture name is checked against the registry to see if it is already in use. A warning displayed if a duplicate name is found.
Upon clicking Next, the mixture name is checked against the registry to see if it is already in use. A warning displayed if a duplicate name is found.
Wizard Step 2: Ingredients
The Ingredients step allows you to add single ingredients or existing mixtures to a new mixture recipe, as well as the required amounts and the amount unit. If you started the wizard from an ingredient or mixture it will be prepopulated here.Specify what Recipe Measure Type the mixture is using: Mass or Volume. This selection dictates what unit types are available for the ingredients below.- If Mass is selected, unit types will be: g/kg, g/g, mL/kg, mol/kg, mol/g, S/S
- If Volume is selected, unit types will be: g/kL, g/L, L/L, mL/L, mol/kL, mol/L, μL/L, μL/mL
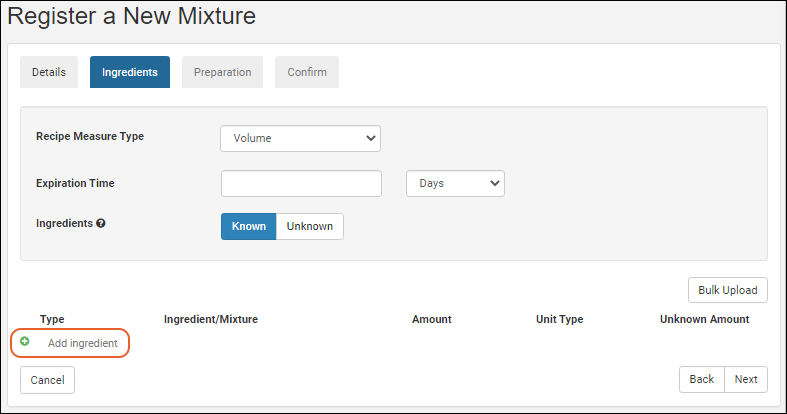 Under Type select either Ingredient or Mixture. The Ingredient/Mixture text boxes are 'type ahead' searches, that is, as you type, the registry will offer a filtered dropdown of options that contain your typed string. To control how many fields are shown with each option, you can provide a lookup view.
Under Type select either Ingredient or Mixture. The Ingredient/Mixture text boxes are 'type ahead' searches, that is, as you type, the registry will offer a filtered dropdown of options that contain your typed string. To control how many fields are shown with each option, you can provide a lookup view.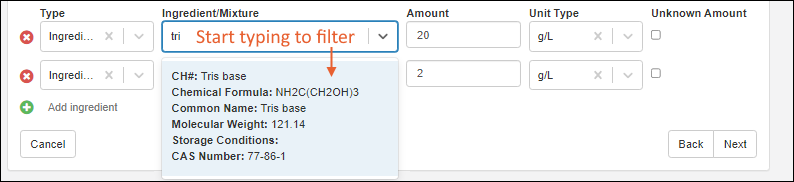 When at least one ingredient/mixture and the associated amount/amountUnit fields have been filled in, the Next button becomes enabled. If a selected Ingredient or Mixture is no longer desired, the red X button on the left can be clicked to remove it.
When at least one ingredient/mixture and the associated amount/amountUnit fields have been filled in, the Next button becomes enabled. If a selected Ingredient or Mixture is no longer desired, the red X button on the left can be clicked to remove it.
Wizard Step 3: Preparation
The preparation step lets you add one or more preparation instructions. Click Add Step to add one or more text boxes. When all instructions have been entered, click the Next button.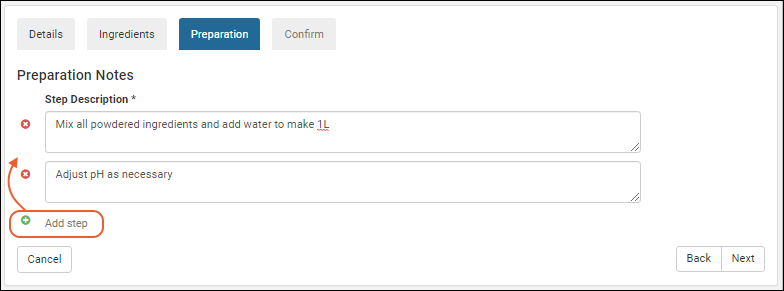
Wizard Step 4: Confirmation
The confirmation step summarizes all of the information entered so far.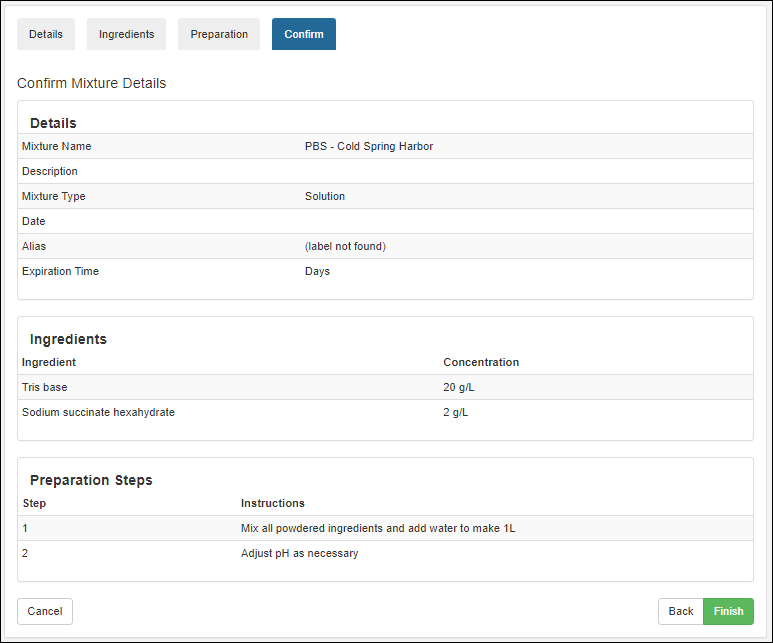 Click Finish to submit; the mixtures grid is shown with the new addition.
Click Finish to submit; the mixtures grid is shown with the new addition.
Register a Mixture using 'Bulk' Ingredients Table
This method of registering a mixture lets you enter the ingredients in a tabular format by copying-and-pasting from an Excel or TSV file.- Go to the Mixtures grid and click Add.
- On the Details tab, enter the Mixture Name and Mixture Type (and description and aliases if needed) then click Next.
- On the Ingredients tab, click Bulk Upload.
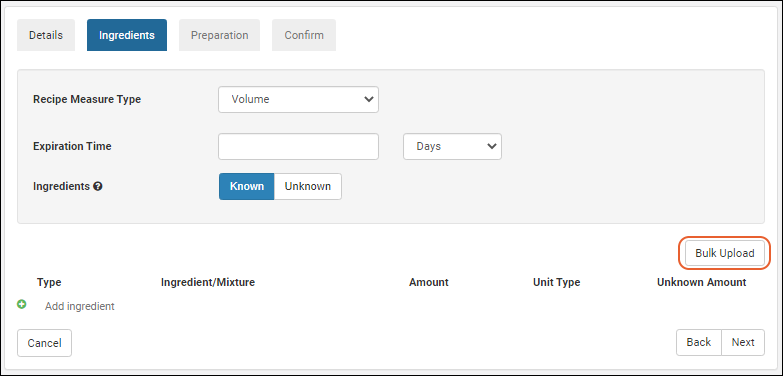
- A popup window appears showing the table headers which you can copy-and-paste into an empty Excel file.
- Fill out the table, adding separate lines for each ingredient in the mixture. Only the "Ingredient/Mixture" column is required. If a cell is left blank, you can complete the details of the mixture using the user interface.
- Type: (Optional) Specify either "Ingredient" or "Mixture".
- Ingredient/Mixture: (Required) The name of the ingredient or mixture which must be a pre-existing item already in the registry. Values from the Name or Scientific Name columns are supported.
- Amount: (Optional) A number that indicates the weight or volume of the ingredient.
- Unit Type: (Optional) Possible values are the same as when entering ingredients in the UI.
- Select whether to:
- Append new ingredients or
- Replace ingredients already listed.
- Copy-and-paste the table back into the popup window and click Add Ingredients.
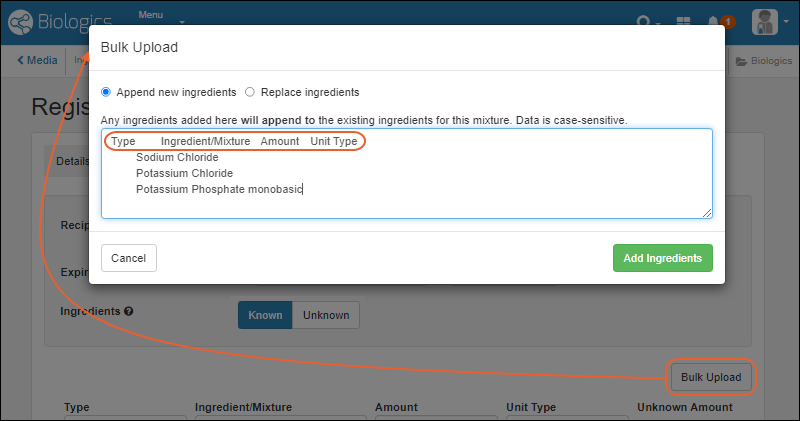
- Fill in any remaining fields, if necessary, and click Next.
- Complete the rest of the mixture wizard.
Registering Mixtures with Unknown Ingredients, Amounts, or Concentrations
In cases where you do not know the exact ingredients, amounts, or concentrations of a material, you can still add it to the registry. These scenarios are common when receiving a material from an outside vendor who does not disclose the exact formulation of the product.In such cases, the mixture registration process is largely the same, except for the Ingredients step, where you can toggle all ingredients and amounts as unknown, or toggle individual amounts as unknown.Clicking Unknown disables entry of any further details about the recipe: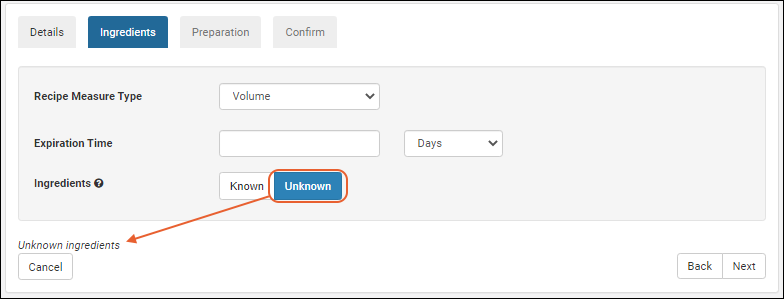 Selecting Known but checking an Unknown Amount box disables that specific ingredient's amount and unit type inputs (other ingredients may still have known amounts).
Selecting Known but checking an Unknown Amount box disables that specific ingredient's amount and unit type inputs (other ingredients may still have known amounts).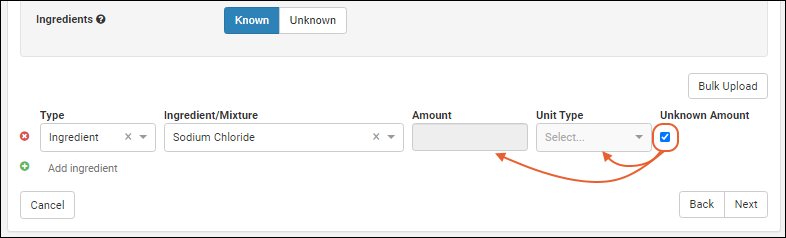 Note that you can register materials and mixtures without specifying ingredients or amounts, even when you do not explicitly check one of the 'unknown' boxes. Warning messages are provided to confirm that your registration is intentional.
Note that you can register materials and mixtures without specifying ingredients or amounts, even when you do not explicitly check one of the 'unknown' boxes. Warning messages are provided to confirm that your registration is intentional.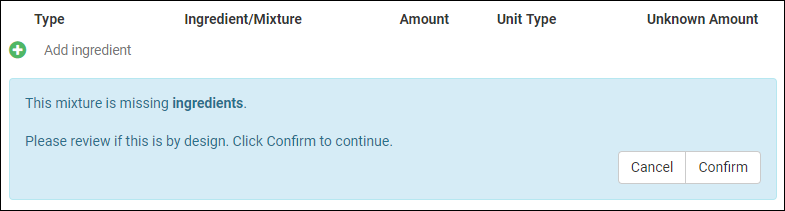 Once registered, mixtures with unknown ingredients and amounts behave just as any other mixture, with a slight change to their detail view to illustrate unknown amounts or ingredients.
Once registered, mixtures with unknown ingredients and amounts behave just as any other mixture, with a slight change to their detail view to illustrate unknown amounts or ingredients.
Bulk Registration of Mixtures That Include Unknowns
In bulk registration of Mixtures, some fields support the text "unknown" on import. For details, see Bulk Registration of Entities.Delete a Mixture
Each Mixture is composed of an entry in the Mixture DataClass table and a Protocol that defines the ingredients used and the steps. To delete the mixture completely, delete the Protocol and then you can delete the Mixture DataClass.- Switch to the LabKey Server interface via > LabKey Server > [Folder Name]
- Select > Go to Module > Experiment.
- Scroll down to the list of Protocols.
- Select the Protocol(s) to delete, and click to delete.
- You should now be able to delete the Mixture DataClass row.
Use the Recipe API to Update Mixtures
Mixtures can be updated after they are created by using the Recipe API. You can make these updates using PUT requests to recipe-recipe.api:- Mutation of ingredients and their associated metadata.
- Description can be changed on the underlying protocol.
- Recipe produces metadata.
- Recipe name
- Changing the recipe steps. Use the pre-existing recipe-steps.api endpoint if you're looking to edit these.
- Call recipe-read.api and receive the full recipe/mixture definition.
- Mutate the received object in the way you'd like (change ingredients, amounts, etc).
- Remove things you do not want to mutate (e.g. "produces", etc) or similarly copy to a new object only the properties you want to mutate.
- Call recipe-recipe.api and PUT the mutated object on the payload as the recipe.
Related Topics
- Managing Ingredients and Raw Materials: Register new ingredients and raw materials
- Registering Batches
Registering Batches
- Batch Design
- Batches
- Registering Batches with Unknowns
- Bulk Registration of Batches That Include Unknowns
- Aliquot Batches
- Add Detail to "Raw Materials Used" Dropdown
- Use the Recipe API to Update Batches
Batch Design
Batches represent a specific quantity of a given mixture. The mixture is a virtual entity, i.e. a recipe, used to create a real, physical batch of material. The design of the Batch, meaning the fields, properties, and naming pattern, is set by default to provide commonly used fields and defaults, but can be edited using Manage > Edit Batch Design.Batches
To create a new batch, click Add above the batches grid, or select Manage > Create Mixture Batch from any mixture detail page to include that mixture in your batch.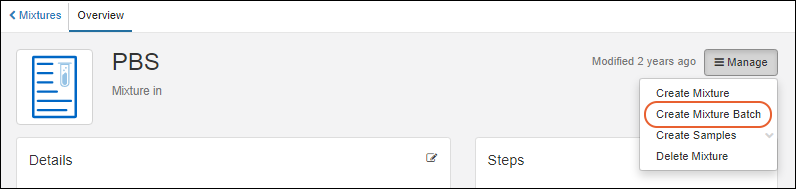 The steps in the batch creation wizard are similar to those in the mixture wizard.
The steps in the batch creation wizard are similar to those in the mixture wizard.
Details
On the Details tab, complete all required fields and any option fields desired. If the Batch Name is not provided, it will be automatically assigned a unique name using the naming pattern for Batches. Hover over the for any field to see more information.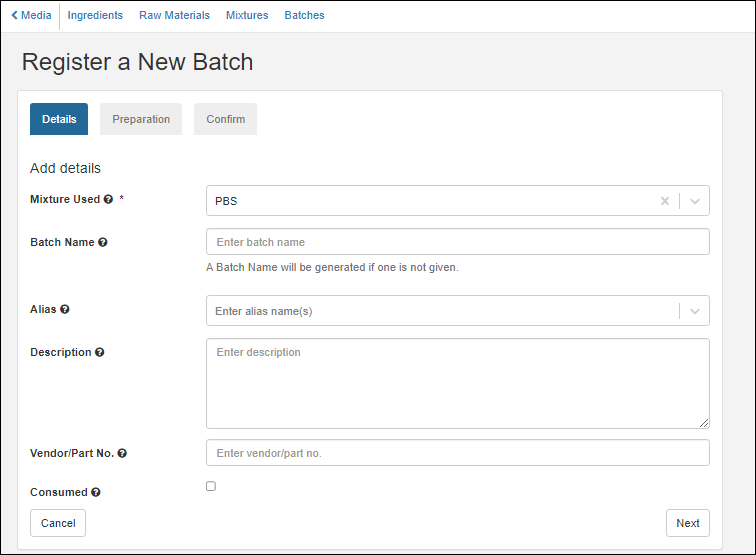 Click Next.
Click Next.
Preparation
On the Ingredients tab, each ingredient row is disabled until you add a Desired Batch Yield and Unit. Once selected, the rest of the page will become enabled and populate information.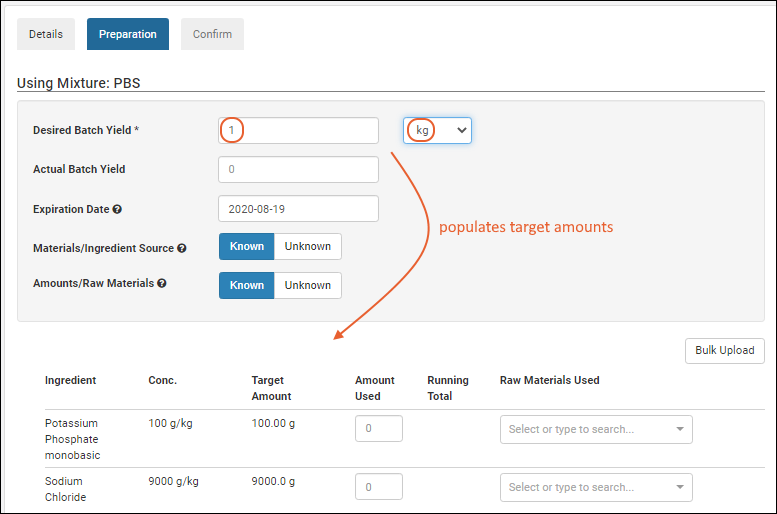 Enter the exact amounts of which raw materials were used to create the mixture recipe. (If ingredients and/or amounts are unknown, see options below.) For each raw material, specify a source lot by its id number. An administrator can enhance the details show to users in this dropdown or other dropdowns by using Identifying Fields.Each ingredient in the recipe may have one or more source lots of raw material This is useful when you exhaust one lot and need to use a second lot to complete the batch. For example, using 40g of Potassium Phosphate empties a lot, if the next lot contains 45, you would need an additional 15g from another lot to reach the target amount of 100g shown in the following. Add additional lots by clicking Add raw material for the specific ingredient.
Enter the exact amounts of which raw materials were used to create the mixture recipe. (If ingredients and/or amounts are unknown, see options below.) For each raw material, specify a source lot by its id number. An administrator can enhance the details show to users in this dropdown or other dropdowns by using Identifying Fields.Each ingredient in the recipe may have one or more source lots of raw material This is useful when you exhaust one lot and need to use a second lot to complete the batch. For example, using 40g of Potassium Phosphate empties a lot, if the next lot contains 45, you would need an additional 15g from another lot to reach the target amount of 100g shown in the following. Add additional lots by clicking Add raw material for the specific ingredient. Note that if one or more raw materials are indicated, the option to set an amount or raw material as unknown is no longer available.
You can also add ingredients (or other mixtures) as you register a batch. Click Add ingredient at the end of the ingredient list.
Note that if one or more raw materials are indicated, the option to set an amount or raw material as unknown is no longer available.
You can also add ingredients (or other mixtures) as you register a batch. Click Add ingredient at the end of the ingredient list.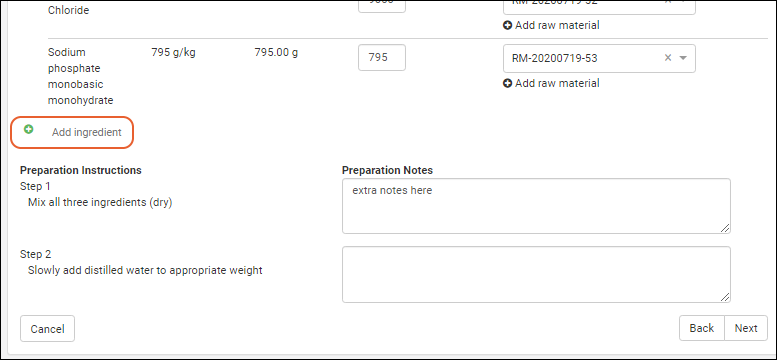 For each preparation step in the mixture recipe, the user can enter any preparation notes necessary in creating this particular batch. Notes are optional, but can provide helpful guidance if something unusual occurred with the batch.Once all required fields are filled, the Next button will become clickable.
For each preparation step in the mixture recipe, the user can enter any preparation notes necessary in creating this particular batch. Notes are optional, but can provide helpful guidance if something unusual occurred with the batch.Once all required fields are filled, the Next button will become clickable.
Confirmation
The confirmation step allows the user to view the information they have entered. If necessary, they can click back to return to previous tabs to make any updates. Preparation notes are collapsed, but can be expanded by clicking the right-arrow icon.Click Finish to register this batch and see the row as entered in the grid.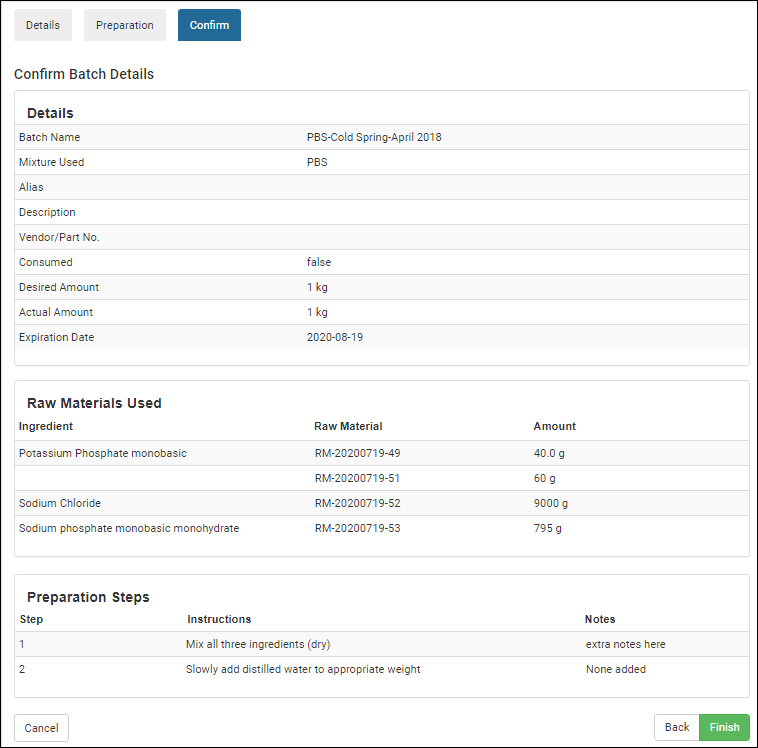 If multiple raw materials were used, the batch details panel will display material lot numbers and the amounts used.
If multiple raw materials were used, the batch details panel will display material lot numbers and the amounts used.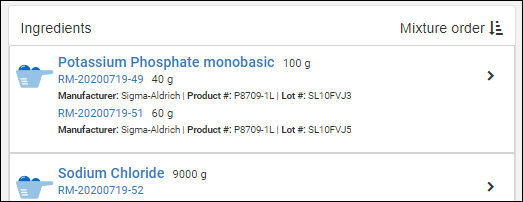 If additional ingredients were added to the batch, these modifications from the mixture recipe will be noted in a panel:
If additional ingredients were added to the batch, these modifications from the mixture recipe will be noted in a panel: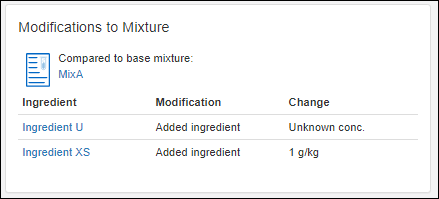
Amounts and Units
The amount, and units for that amount, are determined from the Batch Yield details provided on the Preparation tab. Three fields are stored and visible:- Recipe Amount (RecipeAmount) : The amount, i.e. "Desired Batch Yield". This value is stored as a property on the run that created the batch. Note that this field is not editable as it is not a property of the batch.
- Recipe Amount Units (Units and also RawUnits): The units value for the Recipe Amount field.
- Recipe Actual Amount (StoredAmount and also RawAmount): The "Actual Batch Yield".
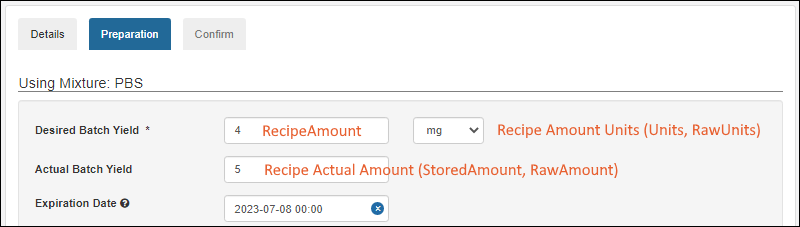 Note that if you have existing data prior to the addition of the Amount and Units default fields in version 23.4, the combined amount-with-units field will be parsed into the two fields. If we cannot parse that text field (such as in a case where the units value is not supported) the two fields will be left blank and need to be manually populated.
Note that if you have existing data prior to the addition of the Amount and Units default fields in version 23.4, the combined amount-with-units field will be parsed into the two fields. If we cannot parse that text field (such as in a case where the units value is not supported) the two fields will be left blank and need to be manually populated.
Registering Batches with Unknowns
When registering batches, the Ingredients step includes the option to mark:- all materials as unknown
- individual raw materials as unknown
- individual amounts as unknown
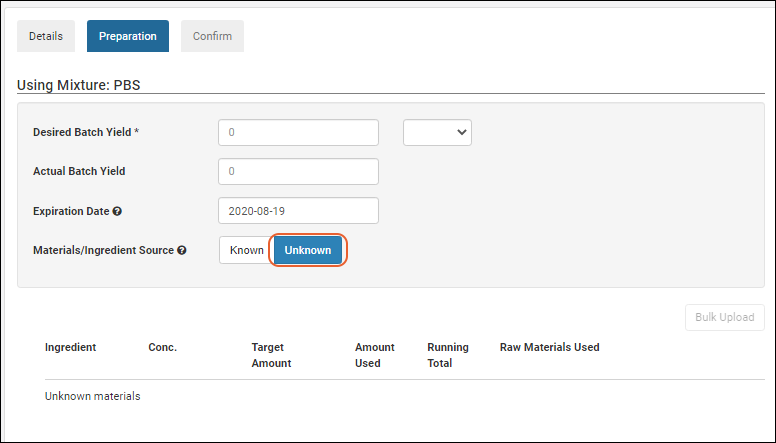 Confirmation warnings are provided if the user provides incomplete values and an 'unknown' box is not ticked.
Confirmation warnings are provided if the user provides incomplete values and an 'unknown' box is not ticked. Once entered into the registry, the unknown factors are reflected in the user interface.
Once entered into the registry, the unknown factors are reflected in the user interface.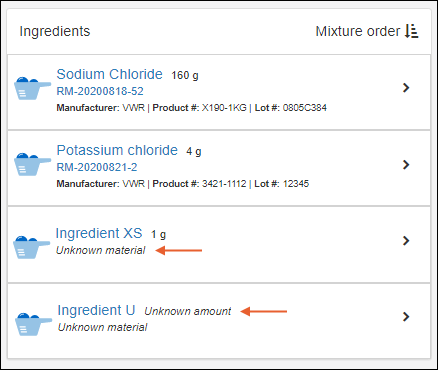
Bulk Registration of Batches That Include Unknowns
In bulk registration of Batches, some fields support the text "unknown" on import. For details, see Bulk Registration of Entities.Aliquot Batches
Instead of having to register sub-portions of mixture batches in advance, you can create large batches, then later create aliquots from them. You can specify the Aliquot Naming Pattern by editing the Mixture Batch Design. The default name of an aliquot is the name of the parent batch, followed by a dash and a counter for that parent. Learn more here: Select one or more Batches from the grid and choose Derive > Aliquot Selected.Add Detail to "Raw Materials Used" Dropdown
The "Name" of a Raw Material is it's generated ID; these are not particularly helpful to users selecting the raw materials for a batch. By customizing raw materials with Identifying Fields, administrators can provide their users with the necessary details when they select from the dropdown.For example, if you define "Product Number" and "Lot Number" as identifying fields, they will appear for users with the "Name" of the raw material.Use the Recipe API to Update Batches
Batches can be updated after they are created by using the Recipe API. You can make these updates using PUT requests to recipe-batch.api:- Mutation of materials and their associated metadata.
- Mutation of amount, amountUnits, and actualAmount of produces.
- Comments
- Changing which sample is produced.
- Changing the source recipe (mixture).
- Changing the batch steps. Use the pre-existing recipe-notes.api endpoint if you're looking to edit these.
- Call recipe-getBatch.api and receive the full mixture batch definition.
- Mutate the received object in the way you'd like (change materials, amounts, etc.).
- Remove things you do not want to mutate (e.g. "produces", etc.) or similarly copy to a new object only the properties you want to mutate.
- Call recipe-batch.api and PUT the mutated object on the payload as the batch.
Related Topics
Biologics Administration
Biologics Admin Topics
- Biologics: Detail Pages and Entry Forms
- Biologics: Protect Sequence Fields
- Manage Notebook Tags
- Biologics Admin: URL Properties
Related Topics
Biologics: Detail Pages and Entry Forms
Custom Details Pages
An administrator can create a special custom grid view to control the fields shown on an entity details page. If you create this special view using the Biologics interface, it will only show for your own user account. To make this change to the details view for all users, select > LabKey Server > [Your Biologics Project Name] to switch to the LabKey Server UI. Here you can save the "BIOLOGICSDETAILS" view and share it with all users.Customize the view for the entity type, then save it:- Uncheck "Make default view for all users"/"Default grid view for this page".
- Use the name "BIOLOGICSDETAILS".
- Check the box to "Make this grid view available to all users" (available only in the LabKey Server interface).
- Optionally check the box to "Make this grid view available in child folders."
Entry Forms
To modify entry and update forms, modify the fields in the field designer. For example, to add a field "NIH Registry Number" to the Cell Line entity:- From the main menu, click Cell Lines.
- Select Manage > Edit Cell Lines Design.
- Click the Fields section to open it.
- Expand the desired field details and enter a Label if you want to show something other than the field name to users.
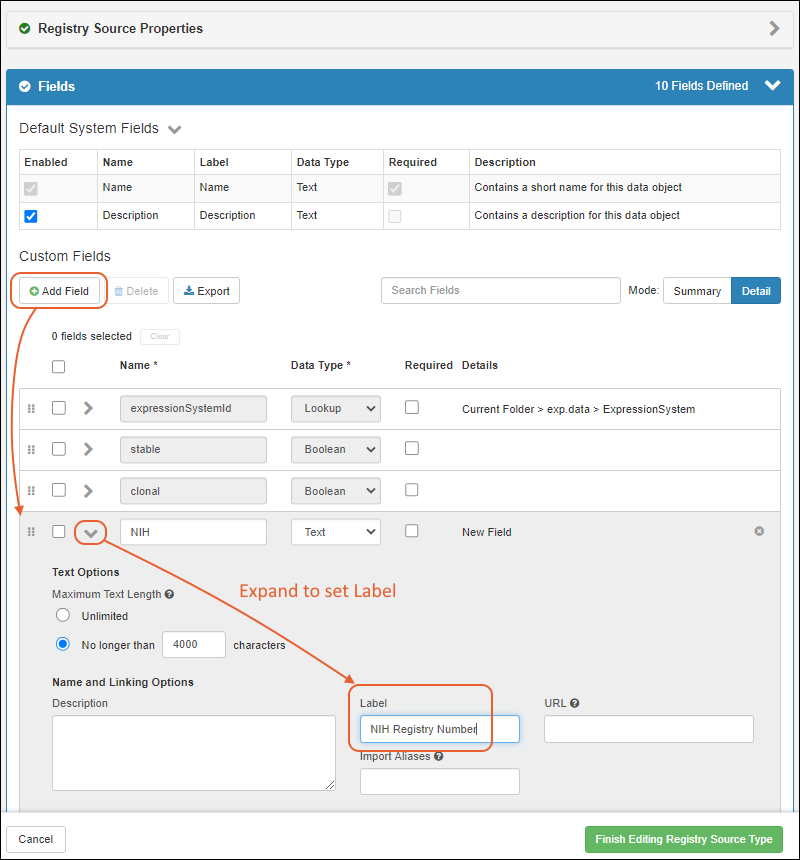
- Click Finish Editing Source Type.
- The field will appear in the entry form for Cell Lines when you Create a new one.
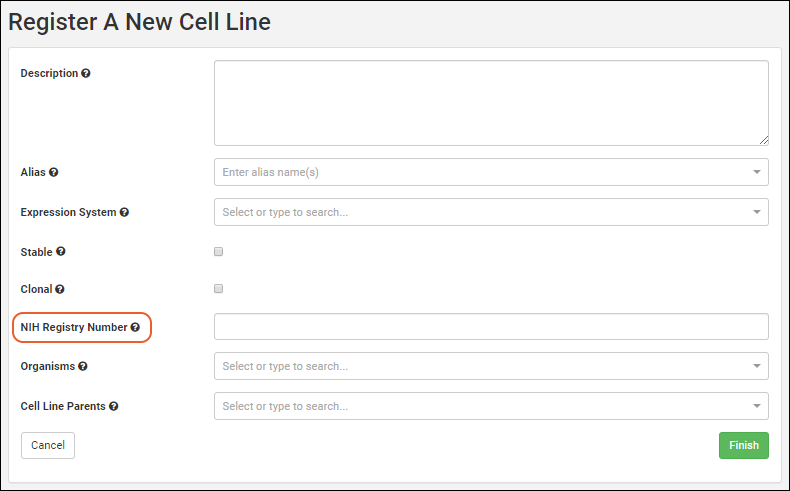 Follow similar procedures to delete or modify entry form fields for other entities.
Follow similar procedures to delete or modify entry form fields for other entities.
Customize Details Shown for Lookups
Lookup fields connect your data by letting a user select from a dropdown which joins in data from another table. By default, a lookup column will only show a single display field from the table, generally the first text field in the target table.Details for Registry Sources and Sample Types
By defining Identifying Fields for Registry Sources and Sample Types, including Media types, you can expose additional detail. Learn more here: As an example of how this can be used to make it easier for users to select the right raw materials for a batch, review this topic: Add Detail to "Raw Materials Used" Dropdown for Registering BatchesDetails for Lists
You can also customize the lookup view in the Sample Manager, LabKey LIMS, and Biologics LIMS for a list or other table by editing the metadata directly. For example, for dropdown targeting a list of Supplier names, it might also be helpful to users to show the state where that supplier is located. In this example, the "Suppliers" list would include the following XML metadata to set shownInLookupView to true for the desired field(s):<tables xmlns="http://labkey.org/data/xml">
<table tableName="Suppliers" tableDbType="NOT_IN_DB">
<columns>
<column columnName="State">
<shownInLookupView>true</shownInLookupView>
</column>
</columns>
</table>
</tables>
Related Topics
Biologics: Protect Sequence Fields
- Restrict Visibility of Nucleotide and Protein Sequences
- PHI/IP Protection Levels and Permissions
- Omit or Blank Protected Columns
- Conditional Viewing of Sequences
- Mark Other Fields as Protected
Restrict Visibility of Nucleotide and Protein Sequences
An administrator can enable Protected Data Settings within the Biologics application. Once enabled, a user (even an administrator) must have the "Restricted PHI Reader" role in the folder to view the protein sequences and nucleotide sequences themselves.- In the Biologics application, select > Application Settings.
- Scroll down to Protected Data Settings, then check the box to Require 'Restricted PHI' permission to view nucleotide and protein sequences.
- If you don't see this box, you do not have sufficient permissions to make this change.
PHI/IP Protection Levels and Permissions
There are four available 'levels' of protection, detailed in this topic: phiLevels- Not PHI
- Limited PHI
- Full PHI
- Restricted PHI: the highest level of information to be protected. Sequences are protected at this level.
- Use the menu to select LabKey Server and your Biologics folder.
- Select > Folder > Permissions.
- Assign the Restricted PHI Reader role to any user who should be able to read Sequences.
- Note that this role is required to see Sequences, even for users with admin access to the Biologics application.
- Learn more about setting permissions here: configuringPerms
- Save and Finish when permissions settings are complete.
Omit or Blank Protected Data
You can choose whether to show an empty column (the default) or omit the column entirely.- Select > LabKey Server > Your Biologics Folder.
- Select > Folder > Management.
- Click the Compliance tab.
- Confirm that Require PHI roles to access PHI columns is checked.
- Use the radio button to select either:
- Blank the PHI column. (Default) The column is still shown in grid views and is available for SQL queries, but it will be empty.
- Omit the PHI column. The column is completely unavailable to the user.
Conditional Viewing of Sequences
Once configured as described above, any user without 'Restricted PHI' permission will see that there is a Sequence column, but the heading will be shaded and no sequences shown. Hovering reveals the message "PHI protected data removed"Mark Other Fields as Protected
Other fields in Registry Source Types and Sample Types may also be protected using the PHI mechanism, when enabled. A user with lower than "Restricted PHI Reader" access will not see protected data. In the LabKey Server interface, they will see a banner to this effect.The mechanism of setting and user experience is slightly different for Registry Source Types and Samples.Protect Registry Source Type Fields
When a field in an entity Registry Source Type other than the Sequence fields is set to a higher level of PHI than the user is authorized to access, it will be hidden in the same way as the Sequence fields.To mark fields in Registry Source Type, such as registry entities including nucleotide and protein sequences, use the LabKey Server interface to edit these "Data Classes".- Use the menu to select LabKey Server and your Biologics folder.
- Click the name of the Data Class to edit, such as "NucSequence"
- Click Edit.
- In the Fields section, click to expand the desired field.
- Click Advanced Settings.
- Use the PHI Level dropdown to select the desired level.
- Click Apply.
- Click Save.
- Return to the Biologics application.
Protect Sample Fields
When a field in a Sample Type is protected at a higher level of PHI than the user can access, they will not see the empty column as they would for an entity field. To mark fields in Sample Types as PHI/IP, use the Biologics interface.- Click the name of your Sample Type on the main menu.
- Select Manage > Edit Sample Type Design.
- In the Fields section, click to expand the desired field.
- Click Advanced Settings.
- Use the PHI Level dropdown to select the desired level.
- Click Apply.
- Click Save.
Related Topics
Manage Notebook Tags
Manage Tags
From the main menu, click Notebooks, then select Manage > Tags to open the dashboard.Delete Unused Tag
To delete tags, an administrator can select one or more rows and click Delete. Tags in use for notebooks cannot be deleted.Add New Tag
To add a new tag, an administrator clicks Create Tag, provides a name and optional description, sets a tag color, then clicks Create Tag again in the popup.Require Tags
By default, new notebooks do not require a tag. An administrator can require a tag for every notebook by selecting > Application Settings and scrolling to the Notebook Settings section. Check the box to require a tag.Select Tag for Notebook
Notebook authors will see the colors and tags available when they create or edit notebooks:Related Topics
Biologics Admin: URL Properties
Last Page You Viewed
When you are in the LabKey Server interface, you can hand edit the URL to return to the last page you were viewing before switching to LabKey.Substitute the server and path for your biologics-app.view and append the lastPage property:https://example.lkpoc.labkey.com/Biologics%20Example/biologics-app.view?#/.lastPage
Reports Dashboard
To view the Reports dashboard, use "/reports":https://example.lkpoc.labkey.com/Biologics%20Example/biologics-app.view?#/reports
URL Redirects
The following redirects can be done directly to the URL for navigating a user programmatically. Note that these examples assume various rowID to assay mappings that may be different in your implementation.| URL ending like this | Can be represented by | Resolves to... |
|---|---|---|
| Any Biologics URL | #/.lastPage | The last page you were viewing |
| /assay-assayBegin.view?rowId=45 | #/assays/45 | #/assays/general/titer |
| /assay-assayResults.view?rowId=765 | #/assays/765 | #/assays/general/amino%20acids |
| /experiment-showDataClass.view?name=CellLine | #/rd/dataclass/cellline | #/registry/cellline |
| /experiment-showDataClass.view?name=Mixture | #/rd/dataclass/mixture | #/media/mixtures |
| /experiment-showData.view?rowId=3239 | #/rd/expdata/3239 | #/registry/molecularspecies/3239 |
| /experiment-showMaterials.view?rowId=3087 | #/rd/samples/3087 | #/samples/expressionsystemsamples/3087 |
| /list-grid.view?listId=18&pk=2 | #/q/lists/18/2 | #/q/lists/mixturetypes/2 |
| /reports | #/reports | Reports Dashboard |
| /assays | #/assays | Assay Dashboard |
| /query-begin.view | #/q | Schema/Query Browser |
| /list-begin.view | #/q/lists | Lists |