Antibody discovery, engineering, and characterization work involves a great deal of uncertainty about the materials at hand. There are important theoretical calculations necessary for analysis as well as variations of molecules that need to be explored. Scientists want to consider and run calculations several different ways based on variations/modifications they are working with for analysis and inclusion in a notebook.
In the case of a structure format not being recognized, e.g. some scFv diabody, it will not be properly classified and the calculations will be wrong. Providing the ability to reclassify and recalculate molecular physical properties is key for assisting with scenarios such as "What if this S-S bond formed or didn’t?"
Using the built-in
Molecular Physical Property Calculator, you can view the persisted calculations to make your input conditions and calculation type clearer and select alternative conditions and calculations for your entity.
We currently calculate average mass, pI (isoelectric point), and 𝜺 (extinction coefficient) from the sequence, molecular stoichiometry, number of free Cysteine and disulfide bonds. While including stoichiometry, free Cys vs S-S, these inputs include sequence scope/type.
View Physical Property Calculator
From the grid of
Molecules, select the Molecule of interest. Click the
Physical Property Calculator tab.
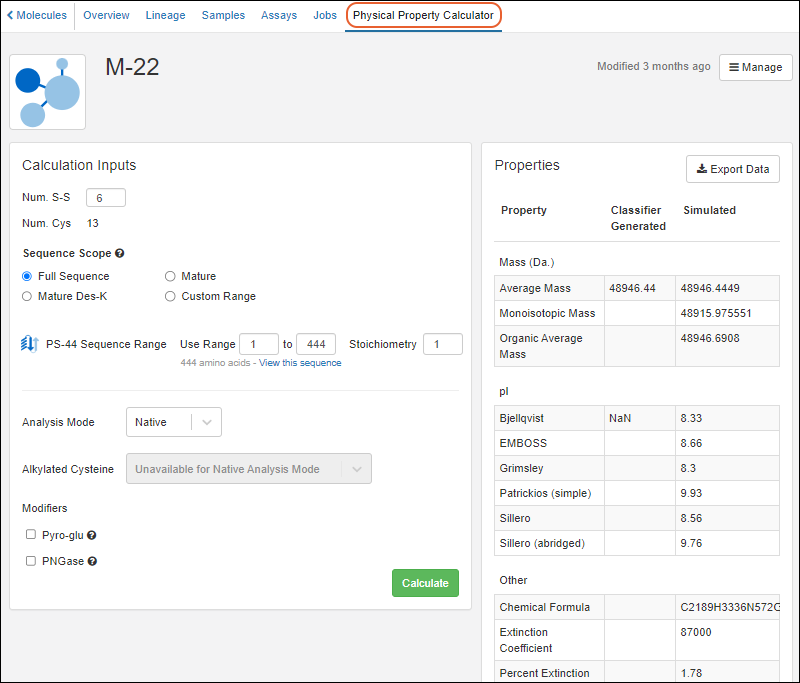
Calculation Inputs
In the
Calculation Inputs panel, you'll enter the values to use in the calculation:
- Num. S-S: Dynamically adjusted to be half of the "Num. Cys" value.
- Num Cys: A display only value derived from the sequence chosen.
- Sequence Scope: Select the desired scope. Sequence ranges will adjust based on your selection of any of the first three options. Use "Custom Range" for finer control of ranges.
- Full Protein Sequence: complete amino acid sequence of a protein, including all the amino acids that are synthesized based on the genetic information.
- Mature Protein Sequence: final, active form of the protein, after post-translational modifications and cleavages, or other changes necessary for the protein to perform its intended biological function.
- Mature Des-K Protein Sequence: mature protein sequence with the C-Terminal Lysine removed.
- Custom Range
- Sequence Ranges for each component sequence. These will adjust based on the range selected using radio buttons, or can be manually set using the "Custom Range" option.
- Use Range: Enter the start and end positions.
- Click View this sequence to open the entire annotated sequence in a new tab for reference.
- Stoichiometry
- Analysis Mode. Select from:
- Alkylated Cysteine (only available in "Reduced" mode). If available, select the desired value.
- Modifiers
- Pyro-glu: Check the box for "Cyclize Gln (Q) if at the N-terminus"
- PNGase: Check the box for "Asn (N) -> Asp (D) at N-link sites"
Click
Calculate to see the calculations based on your inputs.
Properties
When you click
Calculate, the right hand panel of
Properties will be populated. You'll see both what the
Classifier Generated value is (if any) and the
Simulated value using the inputs you entered.
Calculations are provided for:
- Mass
- Average Mass
- Monoisotopic Mass
- Organic Average Mass
- pI - Isoelectric Point calculated by different methods:
- Bjellqvist
- EMBOSS
- Grimsley
- Patrickios (simple)
- Sillero
- Sillero (abridged)
- Other
- Chemical Formula
- Extinction Coefficient - ε
- Percent Extinction Coefficient
- Sequence Length
- Amino Acid (AA) composition
Export Calculations
To export the resulting calculated properties, click
Export Data.
The exported Excel file includes both the calculated properties and the inputs you used to determine them. For example, the export of the above-pictured M-17 calculation would look like this:
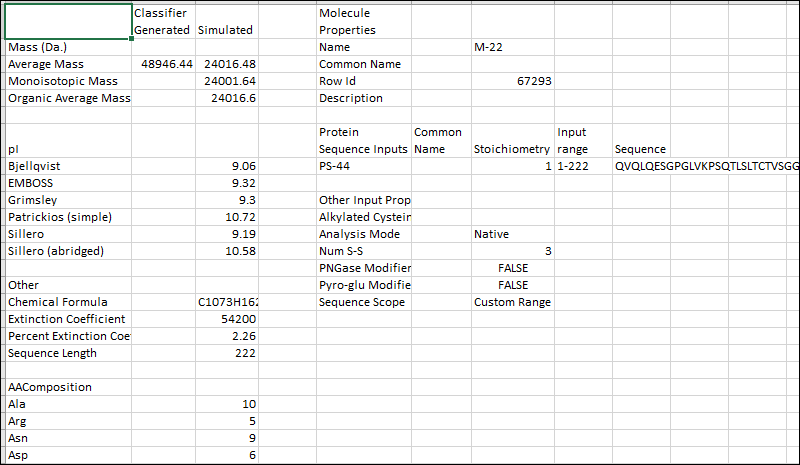
Related Topics