When a protein sequence is added to the registry, the system searches an internal library for matching annotations. Matching annotations are automatically associated with the protein sequence and displayed in a viewer. Protein annotations can also be added manually, edited, and removed for a registered sequence (protein or nucleotide).
Learn more about the methodology used in this internal library in this topic:
When you register a new protein sequence, this methodology is used to apply annotations representing these classifications. It will not override information provided in a GenBank file. After import and initial classification using this methodology, you can then refine or add more annotations as needed.
Annotation Detail Page
To view protein sequence annotations, go to
Registry > Protein Sequences, and click a protein id in the Name column. Click the
Sequence tab.
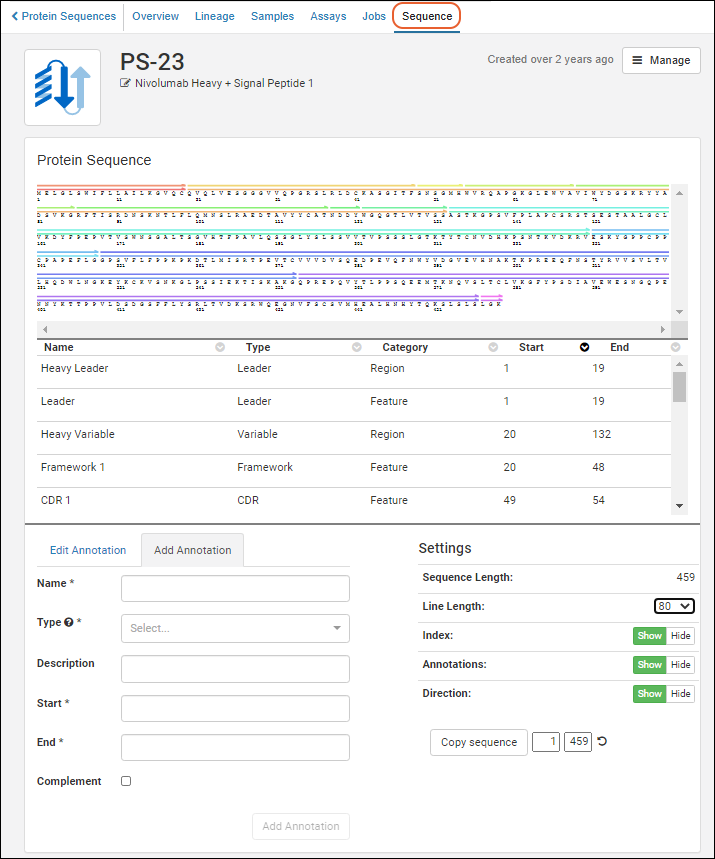
A somewhat simplified version of the annotations viewer is also used within the
protein sequence registration wizard.
Graphical Representation
Clicking an annotation’s color bar highlights its description in the grid below, For example, if you click the green bar for sequence position 24-34, that annotation’s details are highlighted in the grid below, and vice versa.
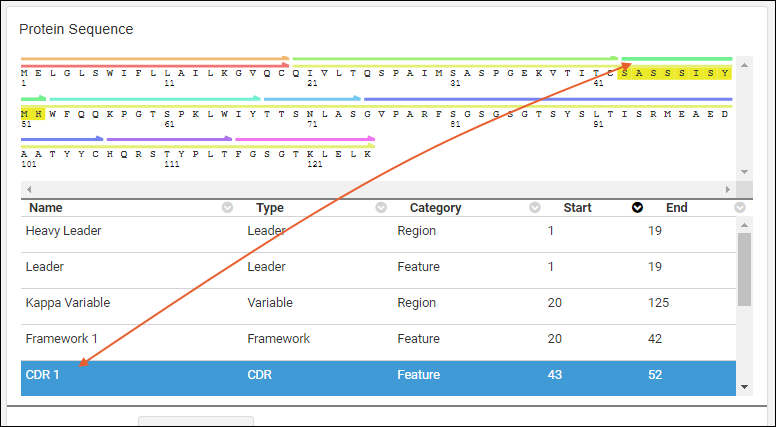
Details Grid
The annotations details grid can be sorted by any of the available columns. The grid will always default to sorting the "Start" column in ascending order
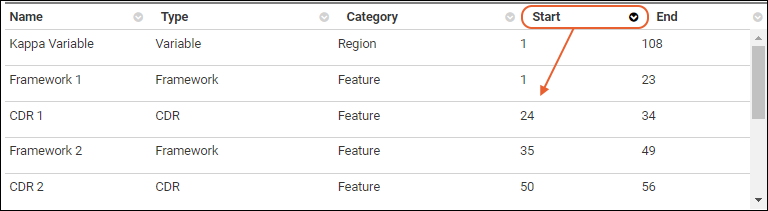
Annotation Editor
The annotation editor has two tabs:
Edit Annotation and
Add Annotation. Adding or editing an annotation will refresh the grid and viewer to include your changes.
- Note the asterisks marking required fields on the Add Annotation tab. The button at the bottom of the form will remain grayed out until all fields have been filled out.
- The Type dropdown is pre-populated with the items in the list "AnnotationTypes".
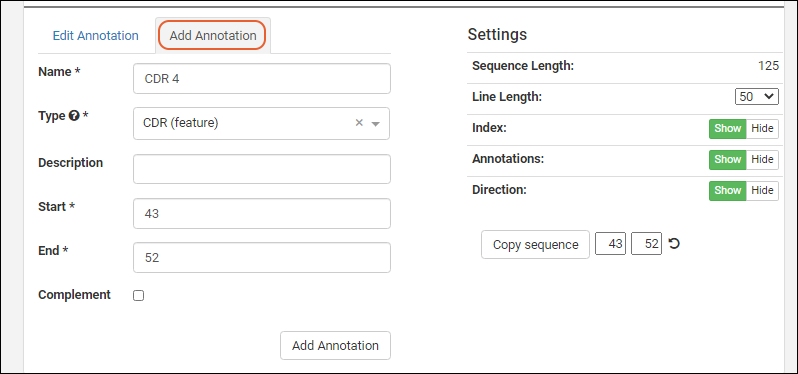
In order to edit an annotation, select one from the grid or viewer and click the
Edit Annotation tab.
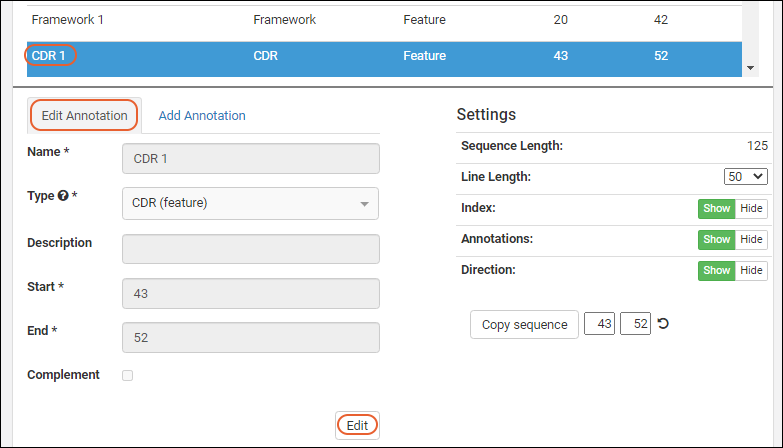
Click the
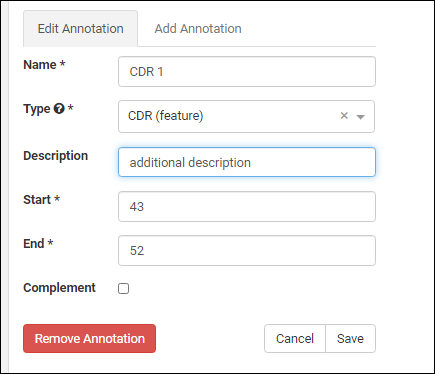
Edit button to enable the field inputs and action buttons:
- Remove Annotation: Removes the annotation and deletes the details row from the grid.
- Cancel: Cancels the edit.
- Save: Save any changes to the selected annotation.
Add AnnotationType
To add a new option to the Annotation Type dropdown, such as "Protease Cleavage Site", switch to the LabKey Server interface and edit the "AnnotationTypes" list. Insert a new row, specifying the name, abbreviation and category you want to use.
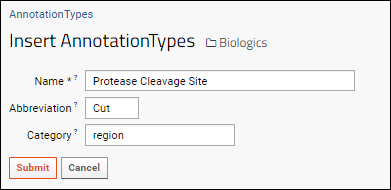
This option will now be available on the dropdown when adding or editing annotations.
Settings/Controls
The controls in the bottom right panel let you customize the display region.
- Sequence Length: The length of the entire sequence.
- Line Length: Select to control the display scale. Options: 50, 60, 80, 100.
- Index: Show or hide or show the position index numbers.
- Annotations: Show or hide the annotation color bars.
- Direction: Show or hide the direction indicators on the bars.
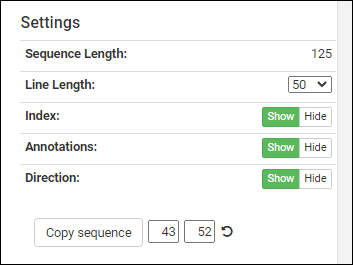
Below these controls, you can see a sequence selection widget next to the
Copy sequence button. The position boxes for start and end update to the currently selected annotation. These can be manually changed to any index values in the sequence, or can be reset to the full sequence by pressing the
refresh icon.
Clicking
Copy sequence will copy the indicated segment to your clipboard for pasting (useful when
registering a new protein sequence into the Registry).
Related Topics
 A somewhat simplified version of the annotations viewer is also used within the protein sequence registration wizard.
A somewhat simplified version of the annotations viewer is also used within the protein sequence registration wizard.


 In order to edit an annotation, select one from the grid or viewer and click the Edit Annotation tab.
In order to edit an annotation, select one from the grid or viewer and click the Edit Annotation tab. Click the Edit button to enable the field inputs and action buttons:
Click the Edit button to enable the field inputs and action buttons:

 Below these controls, you can see a sequence selection widget next to the Copy sequence button. The position boxes for start and end update to the currently selected annotation. These can be manually changed to any index values in the sequence, or can be reset to the full sequence by pressing the refresh icon.Clicking Copy sequence will copy the indicated segment to your clipboard for pasting (useful when registering a new protein sequence into the Registry).
Below these controls, you can see a sequence selection widget next to the Copy sequence button. The position boxes for start and end update to the currently selected annotation. These can be manually changed to any index values in the sequence, or can be reset to the full sequence by pressing the refresh icon.Clicking Copy sequence will copy the indicated segment to your clipboard for pasting (useful when registering a new protein sequence into the Registry).