Table of Contents |
guest 2026-03-19 |
Sample Manager
Release Notes: Sample Manager
Release Notes: LabKey LIMS
Release Notes: Biologics LIMS
Sample Manager & LIMS Validated
Get Started with Sample Manager
Sample Manager Dashboard
Folder Organization
Cross-Folder Actions
User Accounts, Groups, and Roles
My Account
Permission Roles
Manage Groups
Manage Notifications
Sample Manager - FAQ
Request Support
Contact Us
Explore Sample Manager
Ontology Concept Annotations
Help Links
Click to Explore
Samples
Create a Sample Type
Add Samples
Create Samples from Grid
Import Samples from File
Sample ID Naming
Aliquot Naming Patterns
ID and Name Settings
Sample Lineage / Parentage
Aliquots, Derivatives, and Sample Pooling
Sample Expiration
Sample Timeline
Manage Samples
Manage Sample Status
Edit Selected Samples
Sample Finder
Built-in Sample Reports
Sample Search
Print Labels with BarTender
BarTender Integration
Barcode Fields
Sample Picklists
Sources
Create Sources
Associate Samples with Sources
Manage Sources
Experiment / Assay Data
Describe Assay Data Structure
Import Assay Data
Manage Assay Designs
Manage Assay Data
Storage Management
Create Storage
Store Samples
Manage Storage
Manage Storage Unit Types
View Storage Details
Edit Storage Definition
Manage Storage Locations
Move Stored Samples
Manage Stored Samples
Storage Roles
Check Out and Check In
View Storage Activity
Migrate Storage Data into LabKey
Workflow
Start a New Job
Manage Job Queue
Complete Tasks
Edit Jobs and Tasks
Create a Job Template
Manage Jobs and Templates
Electronic Lab Notebooks (ELN)
Notebook Dashboard
Author a Notebook
Notebook References
Notebook ID Generation
Notebook Review
Notebook Amendments
Notebook Templates
Export Notebook
Configure Puppeteer
Troubleshoot ELN Images (Guide)
Troubleshooting ELN Images
ELN: Frequently Asked Questions
Data Resources
Use Sample Manager with LabKey Server
Use Sample Manager with Studies
Data Grid Basics
Data Import Guidelines
Custom Grid Views
Identifying Fields
Audit History
Search
Field Editor
Field Properties Reference
Attach Images and Other Files
URL Field Property
Date, Time, and Number Formats
String Expression Format Functions
LabKey SQL Syntax
LabKey LIMS
LIMS: Downloadable Templates
LIMS: Samples
Print Labels with BarTender
LIMS: Assay Data
LIMS: Charts
LIMS: Storage Management
LIMS: Workflow
Biologics LIMS
Introduction to LabKey Biologics
Release Notes: Biologics
Biologics: Navigate
Biologics: Projects and Folders
Biologics: Bioregistry
Create Registry Sources
Register Nucleotide Sequences
Register Protein Sequences
Register Leaders, Linkers, and Tags
Vectors, Constructs, Cell Lines, and Expression Systems
Registry Reclassification
Biologics: Terminology
Protein Sequence Annotations
CoreAb Sequence Classification
Biologics: Chain and Structure Formats
Molecules, Sets, and Molecular Species
Register Molecules
Molecular Physical Property Calculator
Compounds and SMILES Lookups
Entity Lineage
Customize the Bioregistry
Bulk Registration of Entities
Use the Registry API
Biologics: Plates
Biologics: Assay Data
Biologics: Specialty Assays
Biologics: Assay Integration
Biologics: Upload Assay Data
Biologics: Assay Batches and QC
Biologics: Media Registration
Managing Ingredients and Raw Materials
Registering Mixtures (Recipes)
Registering Batches
Biologics Administration
Biologics: Detail Pages and Entry Forms
Biologics: Protect Sequence Fields
Manage Notebook Tags
Biologics Admin: URL Properties
Explore LabKey Biologics with a Trial
LIMS Enterprise
Release Notes: LIMS Enterprise
LIMS Suite
Sample Manager
- See how Sample Manager can level-up your lab work.
- Getting started with Sample Manager
- LabKey Sample Manager - Guided Tours
LabKey Sample Manager is one of a suite of LabKey products designed to help you track the entire lifecycle of samples in your lab and all associated research data. The topics in this section will help you learn to use the Starter Edition of Sample Manager. All related products use the same general interface, adding additional functionality at each tier.
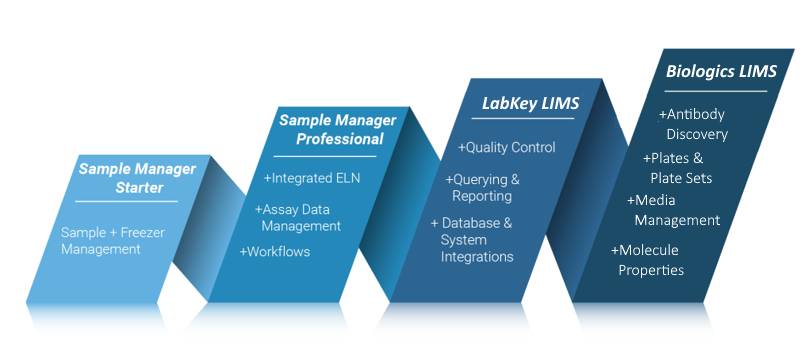
Sample Manager Starter Edition
The intuitive interface makes it easy to:- Manage and track your laboratory samples with customizable types and sources.
- Use storage management tools to track locations of samples in freezers and other storage. (Learn more in the documentation.)
- Retain an audit ready log of actions performed on every sample.
Sample Manager Professional Edition
With the Professional Edition of Sample Manager, you will have all of the Starter Edition features, plus be able to:- Create laboratory workflows and standardize sample processing.
- Upload associated assay and experiment result data and link it to your samples.
- Record your work in electronic lab notebooks integrated directly with your data. (Learn more in the documentation.)
- Partition your data by folder, sharing a storage environment and other resources.
LabKey LIMS
LabKey LIMS includes all features in the Professional Edition of Sample Manager, and adds more, including but not limited to:- Generate customizable reports with ease.
- Add automation with transformation scripts for assay data.
- Customize the downloadable templates available to users importing data
Biologics LIMS
Biologics adds even more functionality to what is available with LabKey LIMS. Learn more here:Topics
- Get Started with Sample Manager
- Sample Manager Dashboard
- Folder Organization
- User Accounts, Groups, and Roles
More Answers
- Release Notes: Sample Manager: What has changed in each version of Sample Manager.
- Sample Manager - FAQ
How Do I Know the Version of my Application
An administrator can see which version is running by selecting > Application Settings. The version is shown in the upper right of every tab in the Administration section.Using Sample Manager with Google Translate
Note that if you are using Google Translate and run into an error similar to the following, it may be due to an "expected" piece of text not appearing in the displayed language at the time the application expects it. Try refreshing the browser or temporarily disabling Google Translate to complete the action.Failed to execute 'removeChild' on 'Node': The node to be removed is not a child of this node.Release Notes: Sample Manager
Release 26.3, March 2026
- LIMS Enterprise Edition is now available.
- Better warnings for unknown fields - when importing cross-sample types, we provide better feedback for unknown fields.
- Improved experience when adding samples to storage - the "Search for Samples" grid now includes the Identifying Fields for a Sample.
- Text Choice options have been increased from 200 items to 500 items.
- Expanded audit coverage - the audit log now captures the creation and editing of grid views.
- Exact text searches now supported using double quotes.
Release 26.2, February 2026
- Column widths now adjust dynamically, allowing more columns to be visible at once with less horizontal scrolling. (docs)
- Configured URL links can now be opened in a new browser tab for easier comparison and multitasking. (docs)
- Entities you don't have access to in lineage views are now shown as restricted rather than being omitted, preserving full context without exposing details. (docs)
- Sample Status is available as a filter for "All Sample Types" in Sample Finder. (docs)
Release 26.1, January 2026
- Support for multiple unit types provides improved inventory and material management. (docs)
- Move workflow jobs to different folders to better reflect changes in projects or organization. (docs)
- Client APIs can query and update samples using the RowId value; using the LSID value is no longer required.
- Sample names (SampleId) can be updated via a file, when RowId is provided. (docs)
Release 25.12, December 2025
- Amount and Units Fields - Improvements have been made to ensure that the Amounts & Units fields function as paired fields. SM-VAL-25.12.A (docs)
- Negative Amount Values Disallowed - Sample Manager now enforces that the Amounts field cannot have a negative value. SM-VAL-25.12.B (docs)
- Identifying Fields - Identifying fields are now shown in more assay import scenarios. SM-VAL-25.12.C (docs)
Release 25.11, November 2025
- Audit log captures additional information on the method or webpage location used to insert, update, and delete records. SM-VAL-25.11.A (docs)
- When an ELN notebook is recalled by an administrator, the author will now receive an email notification, improving visibility and timely follow-up. SM-VAL-25.11.B
- The Customize Grid View and Filter pop-up dialogs now list fields alphabetically, making it faster and more intuitive to find and select fields. SM-VAL-25.11.C
- Two factor authentication is available for configuration by LabKey. Consult your Account Manager for changes. SM-VAL-25.11.D
Release 25.10, October 2025
- Amounts and Units Changes - Amount and Unit fields are now enforced as a pair—both must be completed together or left empty. SM-VAL-25.10.A (docs)
- Required Fields in Workflow Jobs - Required fields in workflow jobs are now enforced during job creation, instead of during job completion. SM-VAL-25.10.B (docs)
- Identifying Fields - Administrators can now set up to 6 identifying fields. SM-VAL-25.10.C (docs)
Release 25.9, September 2025
- Improved Audit Logging Behavior - The LabKey Client APIs will now respect the audit level configured by the system to improve adherence to compliance and ease development. When both the system and API parameters specify an auditing level, the higher, more detailed level is applied. (docs) SM-VAL-25.9.A
- Several improvements were made to overall system reliability and performance.
Release 25.8, August 2025
- Improved ELN Editing: ELN editors now get faster feedback when pasting images into an ELN: files pasted into an ELN now fail immediately if they can't be loaded. SM-VAL-25.7-B (docs)
- The CheckedOut date/time stamp is now an available column in sample grids. SM-VAL-25.7-D
- You can now view all audit events for a transaction in one place. SM-VAL-25.8.A (docs)
- The audit log now records original file names when duplicates are automatically renamed. SM-VAL-25.8-B (docs)
Release 25.7, July 2025
- Improvements were made to address overall system reliability and performance.
- Continued investment in automated testing and internal quality checks to support ongoing feature development.
Release 25.7.8, September 2025
- Selection order is retained when editing in a grid. SM-VAL-25.7-A
- Improved ELN Editing: ELN editors now get faster feedback when pasting images into an ELN: files pasted into an ELN now fail immediately if they can't be loaded. SM-VAL-25.7-B (docs)
- Improved import feedback: When attachment fields are supplied with data in a file import or file update for Sources, users will be provided an error message that Attachment data cannot be provided via a file. SM-VAL-25.7-C
- The CheckedOut date/time stamp is now an available column in sample grids. SM-VAL-25.7-D (docs)
- We have addressed an issue with moving assay runs. Moving assay runs that have multiple file fields now associate correctly. SM-VAL-25.7-E
- We have addressed an issue with cross-sample-type import or cross-folder sample import, where the CheckedOut column was being ignored. SM-VAL-25.7-F
- We have addressed an issue with cross-sample-type import and cross-folder sample import, where Yes/No text fields were being inadvertently converted to Boolean values. SM-VAL-25.7-G
- We have addressed an issue where samples being removed from storage could not be assigned a Locked sample status type. SM-VAL-25.7-H
Release 25.6, June 2025
- Lineage details can be used in aliquot naming patterns. SM-VAL-25.6-A (docs)
- Users can enter a reason when they make changes to a Sample Type, Source Type, or Assay Design. SM-VAL-25.6-B (docs)
- Fields of type "Sample" can be set to validate that values already exist in the system. SM-VAL-25.6-C (docs)
- Several improvements were made to address overall system reliability and performance.
Release 25.5, May 2025
- Several improvements were made to address overall system reliability and performance.
- Continued investment in automated testing and internal quality checks to support ongoing feature development.
Release 25.4, April 2025
- Several improvements were made to address overall system reliability and performance.
- Continued investment in automated testing and internal quality checks to support ongoing feature development.
Release 25.3, March 2025
Maintenance Release 25.3.2, April 2025
- Bulk edit grids now show amounts as entered, regardless of selected units.
- Field names longer than 40 characters are now supported, though not recommended.
Release 25.3.0, March 2025
- You can now use numeric positions for boxes, plates, and tube racks (instead of xy coordinates) when that will better match your lab. SM-VAL-25.3-A (docs)
- Improved support for using special characters in column names and data. SM-VAL-25.3-B (docs)
- Updates to our content security policy (CSP) to enforce strong settings that will block serious cybersecurity threats. Administrators can add allowed external resources if needed. SM-VAL-25.3-C (docs)
Release 25.2, February 2025
- The main dashboard has been simplified. SM-VAL-25.2-A (docs)
- The last storage location is remembered on a per sample type basis, making it easier for users who work with different materials to return to the right locations. SM-VAL-25.2-B (docs)
- Multiple Sample or SourceIDs can now be edited in the editable grid. SM-VAL-25.2-C (docs)
- If desired, you can prevent this by disallowing user-defined names. (docs)
- Adding samples to workflow jobs has been streamlined. SM-VAL-25.2-D (docs)
- Workflow jobs, tasks, and notification subscriptions can be assigned by group as well as by individual user. SM-VAL-25.2-E (docs)
Release 25.1, January 2025
- Creating or deriving samples of multiple types at once can now be done with a streamlined interface. SM-VAL-25.1-A (docs)
- The process of adding lineage details when creating and deriving samples and sources is more consistent. SM-VAL-25.1-B (docs)
- Special characters are not allowed in the names of structures like Sample Types, Source Types, and Assay Designs. SM-VAL-25.1-C (docs)
- When importing new samples and sources from a file, first navigate to the folder where you want to create them. SM-VAL-25.1-D (docs)
Release 24.12, December 2024
- The main dashboard shows "Locations Recently Added To" for easy access to recent storage locations. SM-VAL-24.12-A (docs)
- Improvements in creating and deriving samples and sources let you choose the type and number to create up front. SM-VAL-24.12-B (docs)
- Use identifying fields when editing samples in grids and viewing tooltips during sample creation and derivation. SM-VAL-24.12-C (docs)
- Importing assay data from a workflow job will now only choose relevant sample IDs. SM-VAL-24.12-D (docs)
Release 24.11, November 2024
- Customize which Identifying Fields are shown when users choose samples or sources from dropdowns. SM-VAL-24.11-B (docs)
- Date, time, and datetime fields are now limited to a set of common display patterns, making it easier for users to choose the desired format. SM-VAL-24.11-C (docs)
- A simplified interface for editing lineage and storage information replaces the previous additional tabs on editable grids. SM-VAL-24.11-D (docs)
- Include "Calculation" fields in your sample, source, and assay definitions that can perform calculations on any combination of system and user-defined fields. SM-VAL-24.11-A (docs)
- Folders that are no longer in use can be archived to hide them from view. SM-VAL-24.11-E (docs)
- An administrator may amend any notebook. SM-VAL-24.11-F (docs)
Release 24.10, October 2024
- Lineage relationships can now be marked as required when creating or updating samples or sources. SM-VAL-24.10-A (docs)
- The term "Remove" is now used instead of "Discard" when a sample is taken out of a storage system. SM-VAL-24.10-B (docs)
- The term "Folder" is now used instead of "Project" to describe a subcontainer or partition of data within the application. All data scoping, user access, and configuration options are unchanged. SM-VAL-24.10-C (docs)
Release 24.9, September 2024
- More easily create storage during sample import by adding labels on terminal storage units. SM-VAL-24.9-A (docs)
- Make naming samples easier with the ability to name your samples based on source or sample ancestors regardless of hierarchy. SM-VAL-24.9-B (docs)
- When adding samples to storage, you can use the Sample Creation Order, i.e. the "reverse" of the default grid order. SM-VAL-24.9-C (docs | docs)
- Resolved an issue with editing samples with mixed parent sample or source IDs. In certain scenarios, where projects were in use, samples being edited in the grid with sample or source parents whose sample IDs were a mixture of numeric and non numeric, lineage was unintentionally removed. SM-VAL-24.7.5-J
Release 24.8, August 2024
- More easily get your sample templates to BarTender by exporting the "BarTender Template". SM-VAL-24.8-A (docs)
- Editable grids now validate entered values as you go, rather than waiting for save to check all entries. SM-VAL-24.8-B (docs)
- Resolved an issue with moving boxes and deleting storage hierarchy. In certain scenarios, boxes containing samples were unintentionally deleted when their previous location was simultaneously deleted while they were being moved. SM-VAL-24.7.3-H
Release 24.7, July 2024
Maintenance Release 24.7.5, September 2024
- Resolved an issue with editing samples with mixed parent sample or source IDs. In certain scenarios, where projects were in use, samples being edited in the grid with sample or source parents whose sample IDs were a mixture of numeric and non numeric, lineage was unintentionally removed. SM-VAL-24.7.5-J
Maintenance Release 24.7.3, August 2024
- Resolved an issue with moving boxes and deleting storage hierarchy. In certain scenarios, boxes containing samples were unintentionally deleted when their previous location was simultaneously deleted while they were being moved. SM-VAL-24.7.3-H
Release 24.7.0, July 2024
- The storage dashboard now lists recently used freezers first, and loads the first ten by default, making navigating large storage systems easier for users. SM-VAL-24.7-A (docs)
- New roles, Sample Type Designer and Source (Data Class) Designer, improve the ability to customize the actions available to individual users. SM-VAL-24.7-B (docs)
- Fields with a description, shown in a tooltip, now show an icon to alert users that there is more information available. SM-VAL-24.7-C (docs)
- Lineage across multiple Sample Types can be viewed using the "Ancestor" node on the "All Samples" tab of the grid. SM-VAL-24.7-D (docs)
- Editable grids support the locking of column headers and sample identifier details making it easier to tell which cell is being edited. SM-VAL-24.7-F (docs)
- Exporting data from a grid while data is being edited is no longer supported. SM-VAL-24.7-E (docs)
- Workflow jobs cannot be deleted if they are referenced from a notebook. SM-VAL-24.7-G (docs)
Release 24.6, June 2024
- More easily identify if a file wasn't uploaded with an "Unavailable File" indicator. SM-VAL-24.6-A (docs)
- The process of adding samples to storage has been streamlined to make it clearer that the preview step is only showing current contents. SM-VAL-24.6-B (docs)
- Customize the certification language used during ELN signing to better align with your institution's requirements. SM-VAL-24.6-C (docs)
- Workflow templates can now be copied to make it easier to generate similar templates. SM-VAL-24.6-D (docs)
Release 24.5, May 2024
- Assign custom colors to sample statuses to more easily distinguish them at a glance. SM-VAL-24.5-A (docs)
- Easily download and print a box view of your samples to share or assist in using offline locations like some freezer "farms". SM-VAL-24.5-B (docs)
- Use any user-defined Sample properties in the Sample Finder. SM-VAL-24.5-C (docs)
- More easily understand Notebook review history and changes including recalls, returns for changes, and more in the Review Timeline panel. SM-VAL-24.5-D (docs)
- Administrators can require a reason when a notebook is recalled. SM-VAL-24.5-E (docs)
- Add samples to any project without first having to navigate there. SM-VAL-24.5-F (docs)
- Edit samples across multiple projects, provided you have the appropriate permissions. SM-VAL-24.5-G (docs)
Release 24.4, April 2024
- Maintenance release 24.4.1 addresses an issue with uploading files during bulk editing of Sample data. SM-VAL-24.4.1-K
- Users can better comply with regulations by entering a Reason for Update when editing sample, source or assay data. SM-VAL-24.4-A (docs)
- More quickly find the location you need when adding samples to storage (or moving them) by searching for storage units by name or label. SM-VAL-24.4-B (docs)
- Choose either an automatic (specifying box starting position and Sample ID order) or manual fill when adding samples to storage or moving them. SM-VAL-24.4-C (docs)
- More easily find samples in Sample Finder by searching with user-defined fields in sample parent and source properties. SM-VAL-24.4-D (docs)
- Lineage graphs have been updated to reflect "generations" in horizontal alignment, rather than always showing the "terminal" level aligned at the bottom. SM-VAL-24.4-E (docs)
- Hovering over a column label will show the underlying column name. SM-VAL-24.4-F (docs)
- Up to 20 levels of lineage can be displayed using the grid view customizer. SM-VAL-24.4-G (docs)
- Administrators of the Professional Edition can set the application to require users to provide reasons for updates as well as other actions like deletions. SM-VAL-24.4-H (docs)
- Moving entities between projects is easier now that you can select from multiple projects simultaneously for moves to a new one. SM-VAL-24.4-J (docs)
Sample Manager User Conference - March 28, 2024
 Watch the session recordings to hear what's new in Sample Manager, best practices for using key features, and other users sharing how Sample Manager is used in their labs.
Watch the session recordings to hear what's new in Sample Manager, best practices for using key features, and other users sharing how Sample Manager is used in their labs.
Release 24.3, March 2024
Maintenance Release 24.3.5, May 2024
- Resolved an issue with importing across Sample Types in certain time zones. Date, datetime, and time fields were sometimes inconsistently translated.
- Resolved an issue with uploading files during bulk editing of Sample data.
Version 24.3.0, March 2024
- Storing or moving samples to a single box now allows you to choose the order in which to add them as well as the starting position within the box. (docs)
- When discarding a sample, by default the status will change to "Consumed". Users can adjust this as needed. (docs)
- While browsing into and out of storage locations, the last path into the hierarchy will be retained so that the user returns to where they were previously. (docs)
- The storage location string can be copied; pasting outside the application will include the slash separators, making it easier to populate a spreadsheet for import. (docs)
- The icons and are now used for expanding and collapsing sections, replacing and . (docs)
- Assay run fields can now use the datatypes "Date" or "Time". (docs)
- The ":withCounter" naming pattern modifier is now case-insensitive. (docs)
- Workflow templates can be edited before any jobs are created from them, including updating the tasks and attachments associated with them. (docs)
Release 24.2, February 2024
- In Sample Finder, use the "Equals All Of" filter to search for Samples that share up to 10 common Sample Parents or Sources. (docs)
- Include fields of type "Date" or "Time" in Samples, Sources, and Assay Results. (docs)
- Users can now sort and filter on the Storage Location columns in Sample grids. (docs)
- Sample types can be selectively hidden from the Insights panel on the dashboard, helping you focus on the samples that matter most. (docs)
- The Sample details page now clarifies that the "Source" information shown there is only one generation of Source Parents. For full lineage details, see the "Lineage" tab. (docs)
- Up to 10 levels of lineage can be displayed using the grid view customizer. (docs)
- Menu and dashboard language is more consistent about shared team resources vs. your own items. (docs)
- The application can be configured to require reasons (previously called "comments") for actions like deletion and storage changes. Otherwise providing reasons is optional. (docs)
- Developers can generate an API key for accessing client APIs from scripts and other tools. (docs)
Release 24.1, January 2024
- A banner message within the application links you directly to these release notes to help you learn what's new in the latest version. (docs)
- Search for samples by storage location. Box names and labels are now indexed to make it easier to find storage in larger systems. (docs)
- Only an administrator can delete a storage system that contains samples. Non-admin users with permission to delete storage must first remove the samples from that storage in order to be able to delete it. (docs)
- Electronic Lab Notebook enhancements:
- All notebook signing events require authentication using both username and password. Also available in version 23.11.4. (docs | docs)
- From the notebook details panel, see how many times a given item is referenced and easily open its details or find it in the notebook. (docs)
- Workflow jobs can be referenced from Electronic Lab Notebooks. (docs)
- Editing during ELN is fixed after an issue on Chromium-based browsers (eg. Chrome, Edge) caused users to experience jumpiness during notebook authoring. Also available in version 23.11.7 and 24.1.2.
Release 23.12, December 2023
- Samples can be moved to multiple storage locations at once. (docs)
- New storage units can be created when samples are added to storage. (docs)
- The table of contents for a notebook now includes headings and day markers from within document entries. (docs)
- Workflow templates can now have editable assay tasks allowing common workflow procedures to have flexibility of the actual assay data needed. (docs)
- When samples or aliquots are updated via the grid, the units will be correctly maintained at their set values. This addresses an issue in earlier releases and is also fixed in the 23.11.2 maintenance release.
Release 23.11, November 2023
- Moving samples between storage locations now uses the same intuitive interface as adding samples to new locations. (docs)
- Sample grids now include a direct menu option to "Move Samples in Storage". (docs)
- Standard assay designs can be renamed. (docs)
- Users can share saved custom grid views with other users. (docs)
- Authorized users no longer need to navigate to the home project to add, edit, and delete data structures including Sample Types, Registry Source Types, Assay Designs, and Storage. Changes can be made from within a subproject, but still apply to the structures in the home project. (docs)
- The interface has changed so that the 'cancel' and 'save' buttons are always visible to the user in the browser. (docs)
Release 23.10, October 2023
- Adding Samples to Storage is easier with preselection of a previously used location, and the ability to select which direction to add new samples: top to bottom or left to right, the default. (docs)
- When Samples or Sources are created manually, any import aliases that exist will be included as parent fields by default, making it easier to set up expected lineage relationships. (docs)
- Grid settings are now persistent when you leave and return to a grid, including which filters, sorts, and paging settings you were using previously. (docs)
- Header menus have been reconfigured to make it easier to find administration, settings, and help functions throughout the application. (docs)
- Panels of details for Sample Types, Sources, Storage, etc. are now collapsed and available via hover, putting the focus on the data grid itself. (docs)
Release 23.9, September 2023
- View all samples of all types from a new dashboard button. (docs)
- When adding samples to storage, users will see more information about the target storage location including a layout preview for boxes or plates. (docs)
- Longer storage location paths will be 'summarized' for clearer display in some parts of the application. (docs)
- Naming patterns can now incorporate a sampleCount or rootSampleCount element. (docs)
Release 23.8, August 2023
- Sources can have lineage relationships, enabling the representation of more use cases. (docs)
- Two new calculated columns provide the "Available" aliquot count and amount, based on the setting of the sample's status. (docs)
- The amount of a sample can be updated easily during discard from storage. (docs)
- Customize the display of date/time values on an application wide basis. (docs)
- The aliquot naming pattern will be shown in the UI when creating or editing a sample type. (docs)
- The allowable box size for storage units has been increased to accommodate common slide boxes with up to 50 rows. (docs)
- Options for saving a custom grid view are clearer. (docs)
Release 23.7, July 2023
- Define locations for storage systems within the app. (docs)
- Create new freezer hierarchy during sample import. (docs)
- Import & update samples across multiple sample types from a single file. (docs)
- Administrators have the ability to do the actions of the Storage Editor role. (docs)
- Improved options for bulk populating editable grids, including better "drag-fill" behavior and multi-cell cut and paste. (docs | docs)
- Enhanced security by removing access or identifiers to data in different projects in lineage, sample timeline and ELNs. (docs)
- ELN Improvements:
- The mechanism for referencing something from an ELN has changed to be @ instead of /. (docs)
- Autocompletion makes finding objects to reference easier. (docs)
- Move Assay Runs and Notebooks between Projects. (docs | docs)
Release 23.6, June 2023
- See an indicator in the UI when a sample is expired. (docs)
- On Sample grids in Picklists, Workflows & Source Details pages, when only one Sample Type is included in a grid that supports multiple Sample Types, default to that tab instead of to the general "All Samples" tab. (docs)
- Track the edit history of Notebooks. (docs)
- Control which data structures and storage systems are visible in which Projects. (docs)
- Move Sources between Projects. (docs)
- Project menu includes quick links to settings and the dashboard for that Project. (docs)
Release 23.5, May 2023
- Move Samples between Projects. (docs)
- Assay Results grids can be modified to show who created and/or modified individual result rows and when. (docs)
- BarTender templates can only be defined in the home Project. (docs)
- ELN Improvements: Pagination controls are available at the bottom of the ELN dashboard. (docs)
- A new "My Tracked Jobs" option helps users follow workflow tasks of interest, even when they are not assigned tasks. (docs)
Release 23.4, April 2023
- Add the sample amount during sample registration to better align with laboratory processes. (docs | docs)
- StoredAmount, Units, RawAmount, and RawUnits field names are now reserved. (docs)
- Users with a significant number of samples in storage may see slower than usual upgrade times due to migrating these fields. Any users with existing custom fields for providing amount or units during sample registration are encouraged to migrate to the new fields prior to upgrading.
- Use the Sample Finder to find samples by sample properties, as well as by parent and source properties. (docs)
- Built in Sample Finder reports help you track expiring samples and those with low aliquot counts. (docs)
- Text search result pages are more responsive and easier to use. (docs)
- Administrators can specify a default BarTender template. (docs)
- ELN Improvements: View archived notebooks. (docs)
- Sample Status values can only be defined in the home project. Existing unused custom status values in sub-projects will be deleted and if you were using projects and custom status values prior to this upgrade, you may need to contact us for assistance. (docs)
Release 23.3, March 2023
- Samples can have expiration dates, making it possible to track expiring inventories. (docs)
- If your samples already have expiration dates, contact your Account Manager for help migrating to the new fields.
- Administrators can see the which version of Sample Manager they are running. (docs)
- ELN Improvements:
Potential Backwards Compatibility Issue: In 23.3, we added the materialExpDate field to support expiration dates for all samples. If you happen to have a custom field by that name on any Sample Type, you should rename it prior to upgrading to avoid loss of data in that field.
Release 23.2, February 2023
- Clearly capture why any data is deleted with user comments upon deletion. (docs | docs)
- Data update (or merge) via file has moved to the "Edit" menu of a grid. Importing from file on the "Add" menu is only for adding new data rows. (docs | docs | docs)
- Use grid customization after finding samples by ID or barcode, making it easier to use samples of interest. (docs)
- Electronic Lab Notebooks now include a full review and signing event history in the exported PDF, with a consistent footer and entries beginning on the second page of the PDF. (docs)
- Projects in the Professional Edition of Sample Manager are more usable and flexible.
- Data structures like Sample Types, Source Types, Assay Designs, and Storage Systems must always be created in the top level home project. (docs)
Release 23.1, January 2023
- Storage management has been generalized to clearly support non-freezer types of sample storage. (docs)
- Samples will be added to storage in the order they appear in the selection grid. (docs)
- Curate multiple BarTender label templates, so that users can easily select the appropriate format when printing. (docs)
- Electronic Lab Notebook enhancements:
- To submit a notebook for review, or to approve a notebook, the user must provide an email and password during the first signing event to verify their identity. (docs | docs)
- Set the font-size and other editing updates. (docs)
- Sort Notebooks by "Last Modified". (docs)
- Find all ELNs created from a given template. (docs)
- An updated main menu making it easier to access resources across projects. (docs)
- Easily update Assay Run-level fields in bulk instead of one run at a time. (docs)
Release 22.12, December 2022
- The Professional Edition supports multiple Sample Manager Projects. (docs)
- Improved interface for assay design and data import. (docs | docs)
- From assay results, select sample ID to examine derivatives of those samples in Sample Finder. (docs)
Release 22.11, November 2022
- Add samples to multiple freezer storage locations in a single step. (docs)
- Improvements in the Storage Dashboard to show all samples in storage and recent batches added by date. (docs)
- View all assay results for samples in a tabbed grid displaying multiple sample types. (docs)
- ELN improvements to make editing and printing easier with a table of contents highlighting all notebook entries and fixed width entry layout, plus new formatting symbols and undo/redo options. (docs)
- New role available: Workflow Editor, granting the ability to create and edit workflow jobs and picklists. (docs)
- Notebook review can be assigned to a user group, supporting team workload balancing. (docs)
Release 22.10, October 2022
- Use sample ancestors in naming patterns, making it possible to create common name stems based on the history of a sample. (docs)
- Additional entry points to Sample Finder. Select a source or parent and open all related samples in the Sample Finder. (docs | docs)
- Video Tutorial: Sample Finder
- New role available: Editor without Delete. Users with this role can read, insert, and update information but cannot delete it. (docs)
- Group management allowing permissions to be managed at the group level instead of always individually. (docs | docs)
- With the Professional Edition, use assay results as a filter in the sample finder helping you find samples based on characteristics like cell viability. (docs)
- Assay run properties can be edited in bulk. (docs)
Release 22.9, September 2022
- Searchable, filterable, standardized user-defined fields on workflow enable teams to create structured requests for work, define important billing codes for projects and eliminate the need for untracked email communication. (docs)
- Storage grids and sample search results now show multiple tabs for different sample types. With this improvement, you can better understand and work with samples from anywhere in the application. (docs | docs | docs)
- The leftmost column of sample data, typically the Sample ID, is always shown as you examine wide datasets, making it easy to remember what sample's data you were looking at. (docs)
- By prohibiting sample deletion when they are referenced in an ELN, Sample Manager helps you further protect the integrity of your data. (docs)
- Easily capture amendments to signed Notebooks when a discrepancy is detected to ensure the highest quality entries and data capture, tracking the events for integrity. (docs)
- When exploring a Sample of interest, you can easily find and review any associated notebooks. (docs)
Release 22.8, August 2022
- Aliquots can have fields that are not inherited from the parent sample. Administrators can control which parent sample fields are inherited and which can be set independently for the sample and aliquot. (docs)
- Drag within editable grids to quickly populate fields with matching strings or number sequences. (docs)
- When exporting a multi-tabbed grid to Excel, see sample counts and which view will be used for each tab. (docs)
Release 22.7, July 2022
- Our user-friendly ELN (Electronic Lab Notebook) is designed to help scientists efficiently document their experiments and collaborate. This data-connected ELN is seamlessly integrated with other laboratory data in the application, including lab samples, assay data and other registered data. (docs)
- Make manifest creation and reporting easier by exporting sample types across tabs into a multi tabbed spreadsheet. (docs)
- All users can now create their own named custom view of grids for optimal viewing of the data they care about. Administrators can customize the default view for everyone. (docs)
- Create a custom view of your data by rearranging, hiding or showing columns, adding filters or sorting data. (docs)
- With saved custom views, you can view your data in multiple ways depending on what’s useful to you or needed for standardized, exportable reports and downstream analysis. (docs)
- Customized views of the audit log can be added to give additional insight. (docs)
- Export data from an 'edit in grid' panel, particularly useful in assay data imports for partially populating a data 'template'. (docs | docs)
- Newly surfaced Picklists allow individuals and teams to create sharable sample lists for easy shipping manifest creation and capturing a daily work list of samples. (docs)
- Updated main dashboard providing quick access to notebooks in the Professional Edition of Sample Manager. (docs)
- Samples can now be renamed in the case of a mistake; all changes are recorded in the audit log and sample ID uniqueness is still required. (docs)
- The column header row is 'pinned' so that it remains visible as you scroll through your data. (docs)
- Deleting samples from the system entirely when necessary is now available from more places, including the Samples tab for a Source. (docs)
Release 22.6, June 2022
- Save time looking for samples and create standard sample reports by saving your Sample Finder searches to access later. (docs)
- Support for commas in Sample and Source names. (docs)
- Administrators will see a warning when the number of users approaches the limit for your installation. (docs)
Release 22.5, May 2022
- Updated grid menus: Sample grids now help you work smarter (not harder) by highlighting actions you can perform on samples and grouping them to make them easier to discover and use. (docs)
- Revamped grid filtering and enhanced column header options for more intuitive sorting, searching and filtering. (docs)
- Sort and filter based on 'lineage metadata', bringing ancestor information (Source and Parent details) into sample grids (docs)
- Rename Source Types and Sample Types to support flexibility as your needs evolve. Names/SampleIDs of existing samples and sources will not be changed. (docs)
- Descriptions for workflow tasks and jobs can be multi-line when you use Shift-Enter to add a newline. (docs)
Release 22.4, April 2022
- In the Sample Finder, apply multiple filtering expressions to a given column of a parent or source type. (docs)
- Download templates from more places, making it easier to import samples, sources, and assay data from files. (docs)
Release 22.3, March 2022
- Sample Finder: Find samples based on source and parent properties, giving users the flexibility to locate samples based on relationships and lineage details. (docs)
- Redesigned main dashboard featuring storage information and prioritizing what users use most. (docs)
- Updated freezer overview panel and dashboards present storage summary details. (docs)
- Available freezer capacity is shown when navigating freezer hierarchies to store, move, and manage samples. (docs | docs)
- Storage labels and descriptions give users more ways to identify their samples and storage units. (docs)
Release 22.2, February 2022
- New Storage Editor and Storage Designer roles, allowing admins to assign different users the ability to manage freezer storage and manage sample and assay definitions. (docs)
- Note that users with the "Administrator" and "Editor" role no longer have the ability to edit storage information unless they are granted one of these new storage roles.
- Multiple permission roles can be assigned to a new user at once. (docs)
- Sample Type Insights panel summarizes storage, status, etc. for all samples of a type. (docs)
- Sample Status options are shown in a hover legend for easy reference. (docs)
- When a sample is marked as "Consumed", the user will be prompted to also change it's storage status to "Discarded" (and vice versa). (docs | docs)
- User-defined barcodes in integer fields can also be included in sample definitions and search-by-barcode results. (docs)
- Search menu includes quick links to search by barcode or sample ID. (docs)
- See and set the value of the "genId" counter for naming patterns. (docs)
Release 22.1, January 2022
- A new Text Choice data type lets admins define a set of expected text values for a field. (docs)
- Naming patterns will be validated during sample type definition. (docs)
- Editable grids include visual indication when a field offers dropdown choices. (docs)
- Add freezer storage units in bulk. (docs)
- User-defined barcodes can be included in Sample Type definitions as text fields and are scanned when searching samples by barcode. (docs | docs)
- If any of your Sample Types include samples with only strings of digits as names, these could have overlapped with the "rowIDs" of other samples, producing unintended results or lineages. With this release, such ambiguities will be resolved by assuming that a sample name has been provided. (docs)
Release 21.12, December 2021
- The Sample Count by Status graph on the main dashboard now shows samples by type (in bars) and status (using color coding). Click through to a grid of the samples represented by each block. (docs)
- Grids that may display multiple Sample Types, such as picklists, workflow tasks, etc. offer tabs per sample type, plus a consolidated list of all samples. This enables actions such as bulk sample editing from mixed sample-type grids. (picklists | tasks | sources)
- Improved display of color coded sample status values. (docs)
- Include a comment when updating storage amounts or freeze/thaw counts. (docs)
- Workflow tasks involving assays will prepopulate a grid with the samples assigned to the job, simplifying assay data entry. (docs)
Release 21.11, November 2021
- Archive an assay design so that new data is disallowed, but historic data can be viewed. (docs)
- Manage sample status, including but not limited to: available, consumed, locked. (docs)
- An additional reserved field "SampleState" has been added to support this feature. If your existing Sample Types use user defined fields for recording sample status, you will want to migrate to using the new method.
- Incorporate lineage lookups into sample naming patterns (docs)
- Assign a prefix to be included in the names of all Samples and Sources created in a given project. Removed in version 22.12.
- Prevent users from creating their own IDs/Names in order to maintain consistency using defined naming patterns (docs)
Release 21.10, October 2021
- Customize the aliquot naming pattern. (docs)
- Sample names can incorporate field-based counters using :withCounter syntax. (docs)
- Record the physical location of freezers you manage, making it easier to find samples across distributed sites. (docs)
- Manage all samples and aliquots created from a source more easily. (docs)
- Comments on workflow job tasks can be formatted in markdown and multithreaded. (docs)
- Redesigned job tasks page. (docs)
Release 21.9, September 2021
- Edit sources and parents of samples in bulk
- View all samples (and aliquots) created from a given Source
- Lineage graphs changed to show up to 5 generations instead of 3
- Redesigned job overview page
Release 21.8, August 2021
- Aliquot views available at the parent sample level. See aggregate volume and count of aliquots and sub-aliquots.
- Find Samples using barcodes or sample IDs
- Import aliases for sample Sources and Parents are shown in the Sample Type details and included in downloaded templates.
- Improved interface for managing workflow jobs and templates.
Release 21.7, July 2021
- Sample actions can be initiated from Freezer views
- Sample actions can be initiated from Source detail views
Release 21.6, June 2021
- Create Aliquots of samples singly or in bulk
- Create and manage Picklists of samples to simplify operations on groups of samples
- Track movement of storage units within a freezer, or to another freezer
- Minor change in freezer editing behavior that the hierarchy is initially collapsed.
Release 21.5, May 2021
- Copy a Freezer definition
- Samples and Sources can include images and other file attachments
Release 21.4, April 2021
- Barcode generation for Samples
- When inferring fields to create Sources, Sample Types, and Assay Designs from a spreadsheet, any "reserved" fields present will not be shown in the inferral, simplifying the creation process
- When importing data, any unrecognized fields will be ignored and a banner will be shown
- On the main menu, "My Assigned Work" has been renamed "My Queue"
Release 21.3, March 2021
- A new Tube Rack type of freezer storage unit has been added
- Derive samples from one or more parent samples
- Pool multiple samples into a set of new samples
- The field editor offers a new "Summary" view
Release 21.2, February 2021
- The field editor now includes checkboxes to enable deletion of multiple fields and export of subsets of fields.
- Background Import:
- Removal of previous size limits on data import
- Progress reporting for asynchronous imports
- In-app Notifications when background imports are complete
Release 21.1, January 2021
- Freezer Management
- Match your digital storage to your physical storage
- Store and locate samples with ease
- Monitor usage and capacity in graphical dashboards
- Track volume, freeze/thaw counts, and comments during check in/out events
Release 20.12, December 2020
- Detailed audit logging now shows only what has changed when data is updated.
Release 20.11, November 2020
- Batching of creating samples and sources, enabling selection of "just created" items.
Release 20.10, October 2020
Release 20.9, September 2020
- Label colors are shown in the samples section of the main dashboard.
- In anticipation of future support for Freezer Management, underlying functionality like the ability to access storage locations from the main menu have been added.
Release 20.8, August 2020
- Sample Types can have custom Label Color assignments to help users differentiate them. (docs)
- In anticipation of future support for Freezer Management, underlying functionality like the ability to see the storage location of a sample has been added. These facilities are not yet visible in the application interface.
Release 20.7, July 2020
- Improved search experience. Filter and refine search results. (docs)
Release 20.6, June 2020
- Bug fixes and small improvements
Release 20.5, May 2020
- Use Sample Timelines to track all events involving a given sample.
- Detailed audit logging has been improved for samples, under the new heading "Sample Timeline Events."
- Sample Types can be created by inferring fields from a file, or by defining fields manually. Source types offer the same convenience.
- Editing of sample parents is now available.
- The definition of Sample Types can now include "Source Alias" columns, similar to parent aliases already available.
Release 20.4, April 2020
- The creation interface for Sample Types has been merged to a single page showing both properties and fields. This makes it easier to create naming expressions that use fields in your Sample Type.
- Define Sources for your samples. The source of a sample could be an individual or a cell line or a lab. Tracking metadata about the source of samples, both biological and physical, can unlock new insights.
Release 20.3, March 2020
- Samples can be added to a workflow job during job creation. You no longer need to start a job after selecting samples of interest, but can add or update the samples directly within the job editing interface.
- Removing unnecessary fields is easier with an icon shown in the collapsed field view.
February 2020
- LabKey Sample Manager is Launched!
The symbol indicates a feature available in the Professional Edition of Sample Manager or with a Premium Edition of LabKey Server.
Release Notes: LabKey LIMS
- Generate customizable reports with ease.
- Add automation with transformation scripts for assay data.
Release 26.2, February 2026
- Column widths now adjust dynamically, allowing more columns to be visible at once with less horizontal scrolling. (docs)
- Configured URL links can now be opened in a new browser tab for easier comparison and multitasking. (docs)
- Entities you don't have access to in lineage views are now shown as restricted rather than being omitted, preserving full context without exposing details. (docs)
- Sample Status is available as a filter for "All Sample Types" in Sample Finder. (docs)
Release 26.1, January 2026
- Support for multiple unit types provides improved inventory and material management. (docs)
- Move workflow jobs to different folders to better reflect changes in projects or organization. (docs)
- Workflow tasks now support sample filters, allowing you to control which samples are included at each step. (docs)
- Improved plot customization with new layout, axis, size, color, and per-series line controls. (docs)
- Client APIs can query and update samples using the RowId value; using the LSID value is no longer required.
- Sample names (SampleId) can be updated via a file, when RowId is provided.
Release 25.12, December 2025
- Amount and Units Fields - Improvements have been made to ensure that the Amounts & Units fields function as paired fields. (docs)
- Negative Amount Values Disallowed - Sample Manager now enforces that the Amounts field cannot have a negative value.(docs)
- Identifying Fields - Identifying fields are now shown in more assay import scenarios. (docs)
Release 25.11, November 2025
- Audit log captures the method used to insert, update, and delete records. (docs)
- When an ELN notebook is recalled by an administrator, the author will now receive an email notification, improving visibility and timely follow-up.
- The Customize Grid View and Filter pop-up dialogs now list fields alphabetically, making it faster and more intuitive to find and select fields.
- Error bars are available on Bar and Line charts. (docs)
- Multiple charts can be displayed above data grids. Select up to 5 charts to display. (docs)
Release 25.10, October 2025
- Amounts and Units Changes - Amount and Unit fields are now enforced as a pair—both must be completed together or left empty. (docs)
- Required Fields in Workflow Jobs - Required fields in workflow jobs are now enforced during job creation, instead of during job completion. (docs)
- Identifying Fields - Administrators can now set up to 6 identifying fields. (docs)
Release 25.9, September 2025
- Improved Audit Logging Behavior - The LabKey Client APIs will now respect the audit level configured by the system to improve adherence to compliance and ease development. When both the system and API parameters specify an auditing level, the higher, more detailed level is applied. (docs)
- Several improvements were made to overall system reliability and performance.
Release 25.8, August 2025
- Improved ELN Editing: ELN editors now get faster feedback when pasting images into an ELN: files pasted into an ELN now fail immediately if they can't be loaded. SM-VAL-25.7-B (docs)
- The CheckedOut date/time stamp is now an available column in sample grids. SM-VAL-25.7-D
- You can now view all audit events for a transaction in one place. SM-VAL-25.8.A (docs)
- The audit log now records original file names when duplicates are automatically renamed. SM-VAL-25.8-B (docs)
Release 25.7, July 2025
- Improvements were made to address overall system reliability and performance.
- Continued investment in automated testing and internal quality checks to support ongoing feature development.
Release 25.7.8, September 2025
- Selection order is retained when editing in a grid. SM-VAL-25.7-A
- Improved ELN Editing: ELN editors now get faster feedback when pasting images into an ELN: files pasted into an ELN now fail immediately if they can't be loaded. SM-VAL-25.7-B (docs)
- Improved import feedback: When attachment fields are supplied with data in a file import or file update for Sources, users will be provided an error message that Attachment data cannot be provided via a file. SM-VAL-25.7-C
- The CheckedOut date/time stamp is now an available column in sample grids. SM-VAL-25.7-D (docs)
- We have addressed an issue with moving assay runs. Moving assay runs that have multiple file fields now associate correctly. SM-VAL-25.7-E
- We have addressed an issue with cross-sample-type import or cross-folder sample import, where the CheckedOut column was being ignored. SM-VAL-25.7-F
- We have addressed an issue with cross-sample-type import and cross-folder sample import, where Yes/No text fields were being inadvertently converted to Boolean values. SM-VAL-25.7-G
- We have addressed an issue where samples being removed from storage could not be assigned a Locked sample status type. SM-VAL-25.7-H
Release 25.6, June 2025
- Lineage details can be used in aliquot naming patterns. SM-VAL-25.6-A (docs)
- Users can enter a reason when they make changes to a Sample Type, Source Type, or Assay Design. SM-VAL-25.6-B (docs)
- Fields of type "Sample" can be set to validate that values already exist in the system. SM-VAL-25.6-C (docs)
- Several improvements were made to address overall system reliability and performance.
Release 25.5, May 2025
- Several improvements were made to address overall system reliability and performance.
- Continued investment in automated testing and internal quality checks to support ongoing feature development.
Release 25.4, April 2025
- Several improvements were made to address overall system reliability and performance.
- Continued investment in automated testing and internal quality checks to support ongoing feature development.
Release 25.3, March 2025
Maintenance Release 25.3.2, April 2025
- Bulk edit grids now show amounts as entered, regardless of selected units.
- Field names longer than 40 characters are now supported, though not recommended.
Release 25.3.0, March 2025
- You can now use numeric positions for boxes, plates, and tube racks (instead of xy coordinates) when that will better match your lab. SM-VAL-25.3-A (docs)
- Improved support for using special characters in column names and data. SM-VAL-25.3-B (docs)
- Updates to our content security policy (CSP) to enforce strong settings that will block serious cybersecurity threats. Administrators can add allowed external resources if needed. SM-VAL-25.3-C (docs)
Release 25.2, February 2025
- Assay transform scripts can be configured to run when data is imported, updated, or both. (docs)
- Use conditional formatting to selectively highlight field values in grids. (docs)
Release 25.1, January 2025
- A trendline option has been added to the chart builder for Line charts. (docs)
- Customize the downloadable template file for Samples, Sources, and Assay Designs. (docs)
Release 24.11, November 2024
- The Chart Builder can be used from within the application to add and edit charts on grids. (docs)
Release 24.10, October 2024
Release Notes: Biologics LIMS
Biologics LIMS includes all features available in LabKey LIMS, plus:
- Bioregistry and classification engine
- Media management
- Molecule property calculator
- Plate and plate set support
- Antibody screening and characterization
Release 26.2, February 2026
- GenBank import improved: nearly all information GenBank files is captured on import, including the original file.
- Improved Molecule creation: Select protein sequences to kick off the molecule creation process.
- Column widths now adjust dynamically, allowing more columns to be visible at once with less horizontal scrolling. (docs)
- Configured URL links can now be opened in a new browser tab for easier comparison and multitasking. (docs)
- Entities you don't have access to in lineage views are now shown as restricted rather than being omitted, preserving full context without exposing details. (docs)
- Sample Status is available as a filter for "All Sample Types" in Sample Finder. (docs)
Release 26.1, January 2026
- Support for multiple unit types provides improved inventory and material management. (docs)
- Move workflow jobs to different folders to better reflect changes in projects or organization. (docs)
- Workflow tasks now support sample filters, allowing you to control which samples are included at each step. (docs)
- Improved plot customization with new layout, axis, size, color, and per-series line controls. (docs)
- Client APIs can query and update samples using the RowId value; using the LSID value is no longer required.
- Sample names (SampleId) can be updated via a file, when RowId is provided.
Release 25.12, December 2025
- Amount and Units Fields - Improvements have been made to ensure that the Amounts & Units fields function as paired fields. (docs)
- Negative Amount Values Disallowed - Sample Manager now enforces that the Amounts field cannot have a negative value.(docs)
- Identifying Fields - Identifying fields are now shown in more assay import scenarios. (docs)
Release 25.11, November 2025
- Audit log captures the method used to insert, update, and delete records. (docs)
- When an ELN notebook is recalled by an administrator, the author will now receive an email notification, improving visibility and timely follow-up.
- The Customize Grid View and Filter pop-up dialogs now list fields alphabetically, making it faster and more intuitive to find and select fields.
- Error bars are available on Bar and Line charts. (docs)
- Multiple charts can be displayed above data grids. Select up to 5 charts to display. (docs)
Release 25.3, March 2025
- Rapidly find the plates and experiments in which samples have been used, and vice versa.
- Automatically generate analytics like regressions and statistics to accelerate your work
Release 25.2, February 2025
- Support for advanced plate layouts using using dilutions.
- Use the "Replicate Group" column to denote a plate well as a replicate instead of setting the well's type to "Replicate".
- Replicate wells have a type of "Sample" and the "Replicate Group" will need to be filled in.
- Add Samples to an existing Plate Set.
- Navigate from a plate set to any notebooks that reference it.
- Edits to outlier exclusions will result in the rerunning of any transform scripts that are configured to run on update.
Release 25.1, January 2025
- Users can now specify hit selection filter criteria on Assay fields. When a run is imported/edited the hit selections for the assay results will be recomputed and automatically applied based on these criteria.
- Navigate from a sample to the plate(s) it has appeared on.
- Perform many types of linear regression analysis and chart them.
- Exclude outlier plate-based assay data points and have that reflected in calculations and charts.
Release 24.12, December 2024
- Plate sets can be referenced from an Electronic Lab Notebook.
Release 24.11, November 2024
Major antibody discovery and characterization updates including:- Campaign modeling with plate set hierarchy support.
- Plan plates easier with graphical plate design and templating.
- Automate routine analyses from raw data collected.
- Perform hit selection from multiple, integrated results across plates and data types.
- Generate instructions for liquid handlers and other instruments.
- Automatically integrate multi-plate results including interplate replicate aggregation.
- Dive deeper into plated materials to understand their characteristics and relationships from plates.
Release 24.10, October 2024
- Charts are added to LabKey LIMS, making them an "inherited" feature set from other product tiers. (docs)
Release 24.7, July 2024
- A new menu has been added for exporting a chart from a grid. (docs)
Release 23.12, December 2023
- The Molecule physical property calculator offers additional selection options and improved accuracy and ease of use. (docs)
Release 23.11, November 2023
Release 23.9, September 2023
- Charts, when available, are now rendered above grids instead of within a popup window. (docs)
Release 23.4, April 2023
- Molecular Physical Property Calculator is available for confirming and updating Molecule variations. (docs)
- Lineage relationships among custom registry sources can be represented. (docs)
- Users of the Enterprise Edition can track amounts and units for raw materials and mixture batches. (docs | docs)
Release 23.3, March 2023
- Potential Backwards Compatibility Issue: In 23.3, we added the materialExpDate field to support expiration dates for all samples. If you happen to have a custom field by that name, you should rename it prior to upgrading to avoid loss of data in that field.
- Note that the built in "expirationDate" field on Raw Materials and Batches will be renamed "MaterialExpDate". This change will be transparent to users as the new fields will still be labelled "Expiration Date".
Release 23.2, February 2023
- Protein Sequences can be reclassified and reannotated in cases where the original classification was incorrect or the system has evolved. (docs)
- Lookup views allow you to customize what users will see when selecting a value for a lookup field. (docs)
- Users of the Enterprise Edition may want to use this feature to enhance details shown to users in the "Raw Materials Used" dropdown for creating media batches. (docs)
Release 23.1, January 2023
- Heatmap and card views of the bioregistry, sample types, and assays have been removed.
- The term "Registry Source Types" is now used for categories of entity in the Bioregistry. (docs)
Release 22.12, December 2022
- Projects were added to the Professional Edition of Sample Manager, making this a common feature shared with other tiers.
Release 22.11, November 2022
- Improvements in the interface for managing Projects. (docs)
- New documentation:
Release 22.10, October 2022
- Improved interface for creating and managing Projects in Biologics. (docs)
Release 22.9, September 2022
- When exploring Media of interest, you can easily find and review any associated Notebooks from a panel on the Overview tab. (docs)
Release 22.8, August 2022
- Search for data across projects in Biologics. (docs)
Release 22.7, July 2022
- Biologics subfolders are now called 'Projects'; the ability to categorize notebooks now uses the term 'tags' instead of 'projects'. (docs | docs)
Release 22.6, June 2022
- New Compound Bioregistry type supports Simplified Molecular Input Line Entry System (SMILES) strings, their associated 2D structures, and calculated physical properties. (docs)
- Define and edit Bioregistry entity lineage. (docs)
- Bioregistry entities include a "Common Name" field. (docs)
Release 22.3, March 2022
- Mixture import improvement: choose between replacing or appending ingredients added in bulk. (docs)
Release 21.12, December 2021
Release 21.11, November 2021
- Apply a container specific prefix to all bioregistry entity and sample naming patterns. (docs)
Release 21.10, October 2021
- Customize the definitions of data classes (both bioregistry and media types) within the application. (docs)
Release 21.9, September 2021
- Customize the names of entities in the bioregistry (docs)
Release 21.7, July 2021
- Removal of the previous "Experiment" mechanism. Use workflow jobs instead.
Release 21.5, May 2021
- Nucleotide and Protein Sequence values can be hidden from users who have access to read other data in the system. (docs)
Release 21.4, April 2021
- Specialty Assays can now be defined and integrated, in addition to Standard Assays (docs)
- Creation of Raw Materials in the application uses a consistent interface with other sample type creation (docs)
Release 21.3, March 2021
- Biologics LIMS begins using the same user interface as Sample Manager.
- Release notes for this and other versions prior to this change can be found in the documentation archives.
Release 17.1, March 2017
Sample Manager & LIMS Validated
Overview
Software validation is the process of ensuring that a software system meets its intended use and performs reliably within its operational environment. For regulated industries, validation provides documented evidence that the system consistently produces results that meet predetermined specifications and compliance requirements. Validation of Sample Manager/LabKey LIMS is an optional add-on to Sample Manager Professional Edition or LabKey LIMS and performed by our third-party partner, CompliancePath. The Validation Pack is updated with each ESR release of Sample Manager/LabKey LIMS and includes Installation Qualification (IQ) and Operational Qualification (OQ) documentation.This page will be updated with the most recent version of the release schedule.25.11.8 Sample Manager/LabKey LIMS Validated Release Schedule
Please refer to this schedule for the 25.11.8 validated release of Sample Manager/LabKey LIMS.- Week of January 19: Upgrade test environments to early access release of 25.11.8
- Jan 26 : LKSM Validation Pack Released (Originally scheduled for Feb 6)
- Feb 18: Target release date for LIMS Validation Pack (previously scheduled for Feb 13)
- Upgrade of production environments to 25.11.8
- Asia/Oceania - Feb 10 5:30-7:00am AEDT
- Africa/Europe - Feb 09 6:30-8:00pm GMT
- Americas - Feb 09 6:30-8:00pm EST
If you have any concerns, please contact us at help@labkey.com.
Last updated: 2026-02-05
Sample Manager & LabKey LIMS Software Validation & Release Schedule - General
This is the generalized release schedule for LabKey Sample Manager & LIMS validationPhase 1: Validation Planning and Initial Updates
Release: YY.MM.00Goal: Align on validation scope and begin valpack updates.
- Week 0: Kickoff: LabKey + CompliancePath meet to discuss approach, timelines, and scope.
- Week 1: LabKey upgrades CompliancePath server (pre-work for validation updates).
- Weeks 1-4: CompliancePath begins updating Validation Pack (URS, Risk Assessment, OQ updates).
Phase 2: Customer PQ Preparation & Staging Environment Updates
Releases: YY.MM.02 – YY.MM.06Goal: Prepare customers for PQ by upgrading test environments and communicating schedule.
- Weeks 2-6: LabKey sends communications to customers:
- Release notes for YY.MM.02–YY.MM.06
- Validation timeline & expectations
- Staging upgrade windows
- Rough plans for production upgrades
- Weeks 2-6: LabKey upgrades customer testing/staging environments.
Phase 3: Validation Pack Release & Production Upgrades
Release: YY.MM.08Goal: Deliver final validation materials and complete production rollout.
- Week 8: LabKey upgrades CompliancePath and customer testing environments to YY.MM.08.
- Weeks 9-10: CompliancePath executes OQ testing and finalizes the Validation Pack updates.
- Week 10: CompliancePath releases the completed Validation Pack to LabKey and customers.
- Week 11: LabKey upgrades customer production environments to YY.MM.08.
Get Started with Sample Manager
Welcome to LabKey Sample Manager
The resources linked here will help you get started using LabKey Sample Manager for sample tracking. First, complete the steps in the Exploring Sample Manager guide to learn to add sample information, define lineage, and understand the processes you will follow to find and use your data.Getting Started, Step by Step
More Video Overviews and Demos
Each will open in a new tab.- Upgrade your lab from spreadsheets to Sample Manager
- Spring cleaning with Sample Manager
- See how Sample Manager can level-up your lab work
- Getting started with Sample Manager
Configure Freezers and Other Storage
Matching your physical storage to virtual storage locations in the app can make it easy to know where to find the samples you need.Add Sample Inventories
Getting your sample information loaded is the heart of using Sample Manager. Define the structure of the data to describe each "type" of sample in your system. Once the types are defined, the samples can be created within the application or imported from a spreadsheet.- Create a Sample Type
- Add Samples
- Import Samples from File
- Import Samples with Storage Locations
- Import Samples with Lineage
Associate Sources
If your samples have physical or biological sources that you want to track, you can learn about adding them and associating them with samples in these topics:Define Assays (Premium Feature)
The data you obtain from running instrument tests on your samples will be uploaded as an assay to LabKey Sample Manager. These topics will guide you in designing assays and uploading your data.Outline Workflow (Premium Feature)
Your laboratory workflow can be managed by creating workflow jobs for the sequences of tasks your team performs. Add your users, set permissions, and organize your jobs and templates following these topics:Use Electronic Lab Notebooks (Premium Feature)
Collaborate and record your work in data-connected Electronic Lab Notebooks. Use templates to create many similar notebooks, and manage individual notebooks through a review and signing process.Learn more in this section:Sample Manager Dashboard
- Release Announcement Banner
- Dashboard Insights
- Sample Finder
- Locations Recently Added To
- Main Menu
- Header Menus
- Notebooks Available in the Professional Edition of Sample Manager.
- Jobs List Available in the Professional Edition of Sample Manager.
Release Announcement Banner
At the top of the dashboard, you'll see a banner announcing the latest release. Click the text "See what's new" to link to the release notes.Dashboard Insights
See the current status of the system, with several display options. By default, you see the total count of samples of each Sample Type, shaded by the label color you assign.- Sample Count by Status (Default)
- Sample Count by Sample Type
- Assay Run Count by Assay (Professional Edition Feature)
- Sample Count by Status offers:
- All Statuses
- With a Status
- No Status
- Sample Count by Sample Type -or- Assay run Count by Assay offer the timeframe over which the count of samples or assays is determined. Options:
- All
- In the Last Year
- In the Last Month
- In the Last Week
- Today
Exclude Sample Types from Dashboard Insights
If desired, an administrator can selectively exclude selected Sample Types from the Dashboard Insights panel. For example, if you have one "static" set of inventory items in your repository, but these never change, you may be interested in hiding them to make the Insights panel more usable for the types in active use.Select > Application Settings and scroll down to the Dashboard section. Note that if you are using Folders in Sample Manager, this option is on the Folders tab. In the Dashboard section, you can uncheck any Sample Types that you wish to have excluded from the Sample Insights dashboard for this folder.Sample Finder
Click Go to Sample Finder in the center of the dashboard to search for samples by properties of their parents and sources. Learn more in this topic:Locations Recently Added To
Storage is shown on the dashboard using a Locations Recently Added To widget.- The "All Recent Locations" tab is the default, showing storage locations that have had samples added to them most recently.
- You can also choose "Your Recent Locations" to see where you last added samples, giving you a quick way to jump back to where you were.
Main Menu
Throughout the application, you can access a master menu providing quick links to everything in the system.- Source Types
- Sample Types
- Storage
- Picklists
- Assays Available in the Professional Edition of Sample Manager.
- Workflow Available in the Professional Edition of Sample Manager.
- Notebooks Available in the Professional Edition of Sample Manager.
Header Menus
In the upper right, menus provide quick access to resources and actions.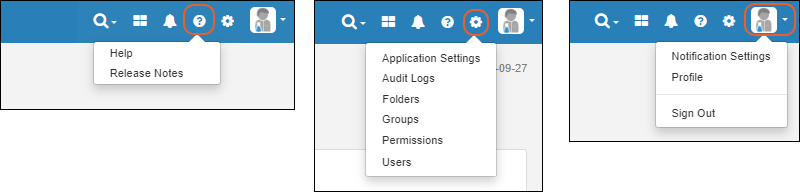
- : Search
- : Product Selection Shown when using a Premium Edition of LabKey Server.
- : Notifications
- : Documentation
- Help Opens the documentation home page for the application.
- Release Notes
- : Administration Settings
- Application Settings: Includes Display Settings, Notebook Settings, BarTender Configuration, ID/Name Settings, and Manage Sample Statuses.
- Audit Logs
- Folders If available
- Groups
- Permissions
- Users
- User avatar menu
Notebooks (Professional Edition Feature)
Collaborate and record your work in data-connected Electronic Lab Notebooks. Use templates to create many similar notebooks, and manage individual notebooks through a review and signing process. Available in the Professional Edition of Sample Manager.Learn more in this section:Jobs List (Professional Edition Feature)
At a glance, see the jobs and tasks assigned to you in Your Job Queue. A second tab will show you other Active Jobs. Learn more about jobs and workflow in this section:Related Topics
Folder Organization
- Video
- Overview
- Home and "Sub" Folders
- Manage Folders
- Select Types of Data in Folder
- Exclude Sample Types from Dashboard Insights
- Select Folder Storage
- Work within Folders
- Best Practices
Video
In this video, you will see how to configure and use Folders to work with data across different teams using Sample Manager Professional.Overview
Without the use of Folder organization, Sample Manager does not partition data for different teams. Permissions are granted to all data simultaneously in a single "container".When you enable Folders in the Professional Edition of Sample Manager, data can be grouped and partitioned by teams.
Home and "Sub" Folders
The home is the top level container and provides shared definitions and storage configurations. Data can be added directly in the home or to the individual "sub" folders as appropriate. Permissions are controlled independently in each folder, making it easy to partition separate team spaces.- Sample Types, Source Types, etc. are defined in the home folder to support lab-wide consistency. Administrators can create and/or edit these structures from within subfolders, but the changes will be made to the home definition and shared by all subfolders.
- Storage systems like freezers are also defined in the home to match the shared physical space. Individual teams can see the details of stored samples they have permission to read, but only see space allocated to other teams as "occupied".
- Reports and views of all data you have access to can be created in the home, summarizing all the data an individual user has access to.
- Create shared resources like reagents in the home so that they can be shared & viewed by all folders.
- Use folder permissions to partition each study from one another making it easy to operate in compliance with regulations.
Manage Folders
To manage Sample Manager Folders, open the application and select > Folders.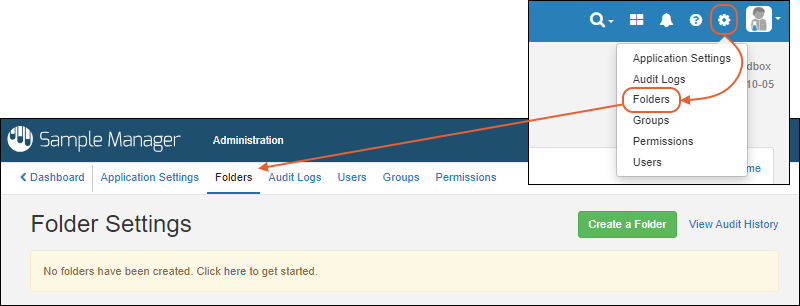 On the Folders administration page, you'll see a listing of any existing folders, with the first selected by default. You can edit existing folder details or create a new one.
On the Folders administration page, you'll see a listing of any existing folders, with the first selected by default. You can edit existing folder details or create a new one.
Create New Folder
Click Create a Folder to add a new folder. Enter the Folder Name and uncheck the checkbox if you wish to provide a different label.- Data: Select the types of data that can be viewed in this folder.
- Dashboard: Select the data types to include in the Dashboard Insights graphs.
- Storage: Select the storage that can be used by samples in this folder.
Set Folder Permissions
When first created, only administrators can view a new folder. This can be useful when configuring the folder for a team to ensure the users only access it when ready.You can configure folder permissions immediately when you create it by clicking Update folder permissions. Later you can also reach this page by clicking the for the folder on the main menu, or from anywhere in the folder by selecting > Permissions.Edit Folder
Click the name of a folder to make any changes to its settings.- Select Types of Data in Folder
- Select Types for Dashboard Insights
- Select Folder Storage
- BarTender Settings
- ID/Name Settings
- Manage Sample Statuses
Archive Folder
When users no longer need regular access to a given folder, such as when the work it records is completed, the folder can be archived. Archived folders retain all their data and access permission settings, so users with access will still see archive folder data in search results, and in the home (top-level) application. Archived folders can be viewed, but will no longer be offered as options for actions like moving samples or other information. This can simplify the selection interface for users without actually deleting the folder or any data.To archive a folder:- Click its name from the > Folders dashboard.
- Click Archive Folder.

- Click Yes, Archive Folder in the popup to confirm.
Delete Folder
To delete a folder:- Click its name from the > Folders dashboard.
- Click Delete Folder.
Select Data in Folder
An administrator can restrict the Source Types, Sample Types, and Assay Designs that are visible in a given folder. This can be configured during folder creation or later by editing folder settings.You'll see all the Source Types, Sample Types, and Assay Designs available in the home (top-level) folder. By default, everything available will be visible in the folder, but you can uncheck boxes for types of data that will not be used in the folder. If data is present, you'll see a message about what will no longer be visible.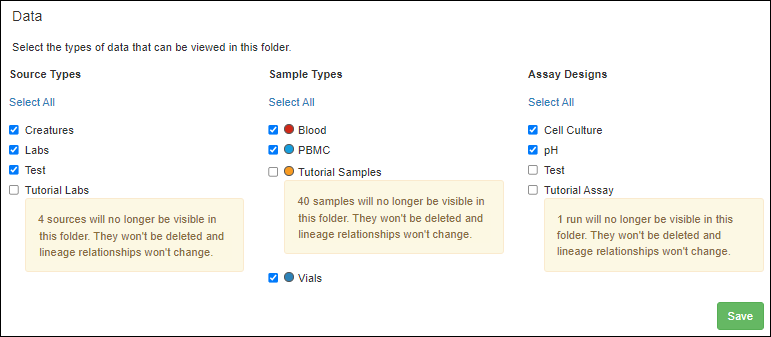 Note that 'hiding' or deselecting a data structure here does not delete any data that may exist using it. It just simplifies dropdown menus for users. The hidden entities will not be seen, but will still be present (and lineage relationships preserved). Nor does hiding a type of data prevent it from being used in the future if these settings are edited.The folders in which a given data structure is visible can also be set while creating or editing the data structure itself in the Folders panel. For example, if the "Labs" Source is not in use for "Team B", the Folders panel might look like this:
Note that 'hiding' or deselecting a data structure here does not delete any data that may exist using it. It just simplifies dropdown menus for users. The hidden entities will not be seen, but will still be present (and lineage relationships preserved). Nor does hiding a type of data prevent it from being used in the future if these settings are edited.The folders in which a given data structure is visible can also be set while creating or editing the data structure itself in the Folders panel. For example, if the "Labs" Source is not in use for "Team B", the Folders panel might look like this:
Exclude Sample Types from Dashboard Insights
When using Folders in Sample Manager, the ability to exclude Sample Types from Dashboard Insights is moved to the Folders tab. There is a Dashboard section for each folder, including for the top level home folder.Select Folder Storage
An administrator can limit the storage systems that are accessible from a given folder by using checkboxes in the Storage panel. For example, if you wanted to create a folder for a team that would only use a single freezer in their local lab, you could hide all other storage systems.Work within Folders
Once one or more folders have been defined, the main menu will include the name of the current folder and a panel on the left for selecting the home or another folder. Each user will see only the folders to which they have access.To navigate, click the name of the desired folder, then the page of interest from the main menu. When you hover over a folder name, you'll see quick links to key pages for that folder:- Dashboard
- Administration
Best Practices
- Home: Top-level folder
- All users must have Read access to the top level of the application.
- Recommended: Only assign non-admin users Read access (and Storage Designer/Editor as appropriate) in the top level Home folder. Edit/add permissions will be granted in individual folders.
- The Home is where all Source Types, Sample Types, Assays, Storage Systems and Templates for ELN and Workflow are defined.
- Administrators can create and edit Source Types, Sample Types, Assays, Storage Systems from within folders, but all changes are made in the home definitions and apply to all folders.
- Add to the Home all "shared resources" you want to be able to access in other folders, such as shared reagent lists, etc.
- Folders
- Team/Group/Study-based access to sources, samples, assay data, workflow & ELNs.
- Within a folder, users can edit/update their folder data.
- From the Home level, users can read their folder data (and all data they can access).
- For data to be readable by everyone but that does not need to integrate (or inherit from) other data, create a folder that everyone has access to.
Related Topics
Cross-Folder Actions
- Cross-Folder Actions
- Restricted Visibility of Inaccessible Data
- Add or Edit Entities Across Folders
- Move Samples, Sources, and Assay Runs Between Folders
- Move Workflow Jobs Between Folders
- Move Notebooks Between Folders
- Storage Shared Across Folders
Cross-Folder Actions
Summary of Functionality
When Folders are in use, the following summarizes the high-level behavior of actions for Samples and Sources (or other Data Classes), collectively referred to as "entities". A given user may have different permissions assigned in different folders, as well as at the application-level, i.e. in the Home folder.Reading of entities: You can read the current folder's entities as well as the ones in the Home and other folders you have permissions to read.Connecting entities with lineage relationships: Entities can have ancestor entities in the current folder and the Home, provided you have the necessary permission. Entities defined in the Home cannot have ancestors in a subfolder.Creating entities: Entities are created by default in the folder the user is viewing at the time the action is initiated. Users with appropriate permissions can choose to create entities in other folders they have access to when using the grid to create them. Importing entities from file will create them in the folder where the import is initiated, unless the file includes a "Container" column (which can also be labelled "Folder").Updating entities: An entity can be updated from the folders in which it was created. Users with appropriate permissions can also update entities across folders. The folder to which the entity is assigned is shown 'read only' when editing entities from multiple folders in a grid. Moving entities between folders should be done separately.Deleting entities: An entity can be deleted from the folder in which it was created. Users with appropriate permissions can also delete entities defined in different folders.Lookups: You can look "UP" but not down the folder hierarchy in lookup fields. In the Home folder you will see only values defined in the Home itself. In a child folder you will see values in both the current and Home folder.- Lookups from one child folder to another child folder may display as expected when viewed from the Home folder. If a user attempts to edit such a cross-folder lookup value from the Home folder, they may see an error that the target is "no longer a valid value". To support such 'sibling' folder lookups, an experimental "Less Restrictive Lookups" feature can be enabled.
Detailed Cross-Folder Actions
When Folders are in use, these notes apply to how some cross-folder actions are handled:- Samples:
- Individual sample creation in a grid allows parent types from the current folder and folders above it. You cannot select parents from a "child" folder of the current one.
- Grid edit and sample deletion: Works for entities in the current folder. With appropriate permissions users can also edit and delete entities defined in different folders.
- Sample derivation, pooling, and aliquoting: Works for entities in the current folder. Users with appropriate permissions can also select samples from the Home and up to one folder for these actions. The derivatives or aliquots will be created in the folder where they are derived/aliquoted, or if in the Home with a folder selected, they will be created in the folder.
- File import only allows sample parent type and parent selection from the current folder.
- Editing parent details for an individual sample only works when adding parents from the current folder or the Home.
- Sources (as well as Bioregistry and Media entities in Biologics LIMS):
- Creating entities, either in a grid or via file import, allows parents and related entities to be chosen from the current folder or folders above it in the hierarchy. You cannot select parents from a "child" folder of the current one.
- Editing in a grid or in bulk is allowed for entities in the current folder. Users with permission can also edit entities from any folder they have access to. You do not need to be in the folder the entity belongs to, but you must have edit permission there.
- Deleting is allowed for entities in the current folder and other folders the user has access to.
- Deriving samples from a data class (source, registry source, or some media) allows parent data class objects from only the current folder or a folder above it.
- Assay:
- Imported assay data can reference samples in the current folder or any 'higher' (parent) folder, but cannot reference samples in sibling or 'lower' (child) folder.
- Imported assay data can also reference samples defined in the /Shared project.
- When assay runs and/or results are editable, users with appropriate permissions can select from multiple folders for editing.
- Storage:
- Freezers (or other storage) can only be defined in the Home folder.
- Samples from multiple folders can be selected at once for adding to storage.
- Learn more about shared storage below
- Workflow:
- Creation of workflow jobs using samples from different folders is possible both from any folder.
- The job that is created will belong to the folder in which it was created, regardless of the samples included.
- When the job is opened, the user is navigated to the folder in which it is defined, to ensure that actions performed as part of the tasks are in the correct folder.
- Additional samples from all visible folders can be added to existing jobs.
- Picklists:
- Creation of picklists using samples from different folders is possible from any folder.
- The picklist that is created will belong to the folder in which it is created, regardless of the samples included.
- The picklist will be visible in only the folder in which it was created.
- Additional samples from all visible folders can be added to existing picklists in the current folder.
- Notebooks:
- Notebooks can reference entities from all folders visible in the current context.
- Notebooks created from a parent folder are also visible from any child folder, and vice versa.
- Notebooks created in any folder can be submitted for review, or accessed for review, from any folder.
Restricted Visibility of Inaccessible Data
When a user has access to a subset of folders, there can be situations where this user should be restricted from seeing data from other folders, including identifiers such as Sample IDs that might contain restricted details. In detail pages, listing pages, and grids, entities a user does not have access to will show "unavailable" in the display instead of a name or rowid.As an example, if a notebook contains references to any data a user cannot access, they will not be able to access that notebook. It will not be included on the "All Notebooks" listing and if they happen to directly access it, such as through a saved URL, they will see only a banner message reading "You cannot view notebook [notebook title] because it contains references to data you don't have permission to view. The references may be in active or archived entries."Add or Edit Entities Across Folders
When the user has the appropriate permissions, samples and sources can be added to a folder the user is not currently in by using the Folder dropdown to select the target folder.- When editing in bulk, the folder is not shown (and not editable).
- When editing in a grid, the folder to which the sample belongs will be shown read-only. If any selected Samples that belong to folders the user is not authorized to edit, they will not be listed for editing.
Move Samples, Sources, and Assay Runs Between Folders
If a user has access to multiple folders and accidentally adds something to the wrong folder, or later wants to change folder affiliation, they can use Edit > Move to Folder to move Samples, Sources, or Assay Runs between folders. Entities currently belonging to multiple folders can be selected at once to be moved to a new folder.Sample, Source, and Assay Run moves require that the user must have update permissions in the current container and insert permission in the target container. If selected entities cannot be moved, the user will see a warning, but authorized moves will proceed.Note that there are some situations in which data cannot be moved:- Samples with a "Locked" status
- Samples with status values that are not defined in the Home folder. This is an uncommon scenario, and is prevented in the current application.
- Assay Runs with QC states that are not defined in the Home folder.
Move from Details Page
You can also move a single Sample, Source, or Assay Run from the details page via Manage > Move to Folder.Move Workflow Jobs Between Folders
To move a workflow job to another folder:- Go to the Workflow main page.
- In the table of jobs, select one or more jobs.
- Click Move to Folder.
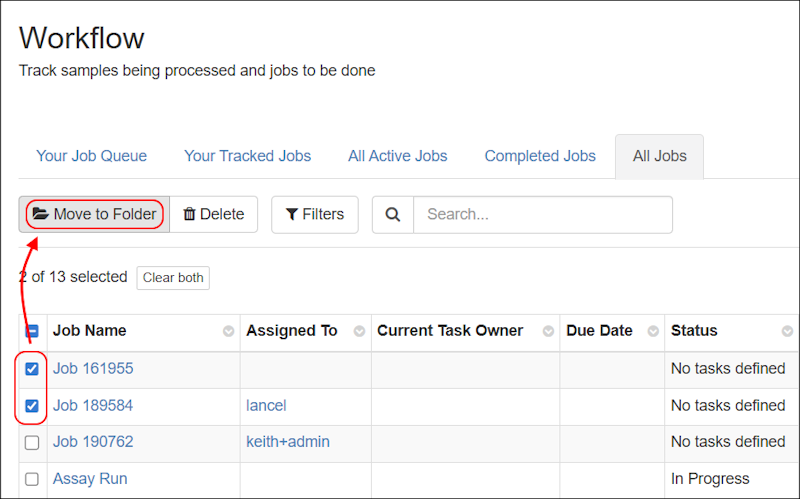
- Select the target folder and click Move.
Move Notebooks Between Folders
Notebooks can be moved from one folder to another provided the user starts in the folder where the notebook is defined and has insert permission in the target folder. One exception is that once a notebook has been approved, it can no longer be moved.Click the for the current Folder setting in the header section of the notebook.Storage Shared Across Folders
Set Up Shared Storage
To share storage, create it in the Home folder, defining the properties and hierarchies as usual. It will then be visible in all subfolders.Users with the "Storage Editor" role in any folder will then be able to use the freezer to store and manage samples. Access to sample details is always dependent upon permissions in the container where the sample is defined.In shared storage where a user has permission to see data from all folders using it, the user will see data from all containers as if all the samples were local. When viewing a sample from another folder, you'll see a note "Actions restricted in the current folder" in the hovertext.Storage Occupied by Inaccessible Samples
In shared storage where a user does not have permission to read data from other folders, storage views will show a spaces as occupied, but instead of revealing any data about the sample, will show a icon. This means the space is occupied by a "restricted sample".Related Topics
User Accounts, Groups, and Roles
- Add Users
- View User Details
- Manage Users
- User Details
- Deactivate Users
- Delete Users (Not recommended)
- View Inactive Users
- User Limit Alerts
Add Users
To add one or more new users, an administrator selects > Users, then clicks Create.- Enter one or more email addresses, each on its own line for each user you want to create.
- Select the desired Roles for the users you are creating:
- You can assign multiple roles at once. Delete a role by clicking the 'X' for the item.
- Learn more about the available roles here: Permission Roles
- Note that all users added at once are assigned the set of roles you select. You can change these role assignments later.
- Optional Message: You can add an optional additional message to include in the invitation email to your new users.
- Click Create Users.
View User Details
Within the application, when an administrator (or user with the "See User and Group Details") clicks a username, such as in a sample grid or ELN, they will see a popup showing the full name, email, description, as well as effective roles and groups this user is a member of.Manage Users
To manage users, an administrator selects > Users.You will see a grid of the active users already present in your Sample Manager application. You can use search, sort, and filter options on this user grid.User Details Panel
To view the details for any user in the grid, check the box for that user. Details including effective roles are shown in a panel to the right.- Reset Password: Send this user an email with a link to set a new password in order to access the site. Passwords must be strong enough to meet the requirements of the site.
- Deactivate
- Delete
Deactivate Users
Deactivated users may no longer log in, but their display name and group membership information will be retained for display and audit purposes. If the user is reactivated at a later time, this information will be restored. Deactivation is the recommended action for former employees, for example.To deactivate a user, an administrator has two options:- Check the box to select a single user you want to deactivate. Click Deactivate in the User Details panel on the right.
- You may also select one or more users simultaneously using the checkboxes in the grid, then select Manage > Deactivate Users.
Delete Users (Not recommended)
Deletion of a user is permanent and cannot be undone; it is generally not recommended. A deleted user's display name will no longer be shown with any assignments or actions taken by that user. A deleted user cannot be reactivated to restore any information.Instead of deleting, deactivation is recommended for any user who has performed any work in the system in the past.One scenario in which deletion might be appropriate is if you originally create a new user with an incorrect email address or other error.To delete a user, an administrator has the same two options as for deactivation:- Check the box to select a single user you want to delete. Click Delete in the User Details panel on the right.
- You may also select one or more users simultaneously using the checkboxes in the grid, then select Manage > Delete Users.
View Inactive Users
Notice that the grid reads Active Users by default. To view the grid of deactivated users instead, select Manage > View Inactive Users.User Limit Alerts
When the number of active users approaches the application limit, administrators will see a warning message on the > Users page.Related Topics
My Account
- Sign In and Out
- Session Expiration
- Edit Your Profile
- API Keys (Professional Edition Feature)
Sign In and Out
From within the LabKey Sample Manager application, you can log in and out via the user avatar menu in the upper right. When you are logged in, there will be a Sign Out link on the user menu where Sign In was before.
When you are logged in, there will be a Sign Out link on the user menu where Sign In was before.Session Expiration
If your session expires while you are using LabKey Sample Manager, you will see a notification popup with a button to Reload Page. You will be asked to log in again before completing the action.Edit Your Profile
Once logged in, you can manage your account information by selecting Profile from the user avatar menu in the upper right.Edit User Details
On your profile page, you can edit your display name, as well as your first and last name and description. You cannot edit your email address here; contact your administrator if you need to change your email address.Upload Avatar
Drag and drop an image into the drop area to use a custom avatar on your profile. The avatar image must have a height and width of at least 256 px. If you upload a rectangular image, it will be cropped to fit the square.Once you have uploaded an avatar, you can reedit your profile and click Delete Current Avatar to revert to the default.Change Password
To change your password, click Change Password in the upper right. In the popup, enter your old password, and the new password you want to use twice. Passwords must be strong and complex enough to meet the requirements set for the site. For Sample Manager clients, the "Strong" set of password rules applies automatically. Click Submit to save the new password.API Keys (Professional Edition Feature)
Developers using the Professional Edition of Sample Manager who want to use client APIs to access data in the application can do so using an API key to authenticate interactions from outside the server. A valid API key provides complete access to your data and actions, so it should be kept secret. Learn more in the core LabKey Server documentation here: Generate an API key from your Profile page.- Scroll down to the API Keys section.
- Click Generate API Key to obtain one.
- Provide an optional Description so you can keep track of when each key you create expires.
- Click Generate API Key again.
- The new key will be shown in the popup and you can click the button to copy it to your clipboard for use elsewhere to authenticate as your userID on this server.
- Important: the key itself will not be shown again and is not available for anyone to retrieve, including administrators. If you lose it, you will need to regenerate a new one.
- You'll see the creation and expiration dates for this key in the grid. The expiration timeframe is configured at the site level by an administrator and shown in the text above the grid. A key without an expiration date was configured when keys were set to never expire.
- The last usage date and any description you entered will also appear here.
Related Topics
Administrators can manage user accounts and permissions as described in this topic:Permission Roles
Available Roles
Once users and groups have been defined, an administrator can assign them one (or more) of the available permission roles:- Administrators: Have full control over the application including user management, permission assignments, storage editor and designer tasks, creating and editing sample types, assays, and job templates.
- Application Administrator is the name of this role when Sample Manager is used as a standalone application.
- When Sample Manager is used with a Premium Edition of LabKey Server, there are two levels: Project Administrators and Folder Administrators. Learn more in the core LabKey Server documentation.
- Editors: May add, edit, and delete data related to samples, assays, and jobs, but not storage information. May assign workflow jobs and tasks to other users, but not to project groups.
- Editor without Delete: May add and edit, but not delete, data related to samples, assays, and jobs. May not edit storage information. Learn about limited exceptions here.
- Readers: Have a read-only view of the application.
- Storage Editors: Have read permission for sample data. They may also add, edit, and remove samples from storage. They can move storage units, provided they have the role in the top level folder. They can create, update, and delete their own picklists, and participate in workflow jobs. Learn more here.
- Storage Designers: May read, add, and edit data related to storage locations. They may delete empty locations. Learn more here.
- Workflow Editor: This role includes the ability to read all data, and allows users to be able to add, update, and delete picklists and workflow jobs. Workflow editors can assign jobs and tasks to groups as well as individual users.
- It does not include the ability to add or edit job templates or any sample, source, or assay data.
- Note that this role does allow a user to delete tasks from a workflow job, as part of having the ability to edit the job.
- Sample Type Designer: Create and design new Sample Types or change existing ones.
- Source Type Designer: Create and design new Source Types or change existing ones.
- Assay Designer: Create and edit Assay Designs.
Assign Roles to Users and Groups
Open permissions management by selecting > Permissions.- First click the role section. You'll see the current members of that role.
- Click the Add member or group dropdown and start typing a user's email address or the name of a user group.
- Click the user email or group name to add to the role.
- Selected users and groups will be shown in the panel for the role as you go.
- Each time you select a user, the details for that user will be shown on the right to assist you.
- In the image below, the Editor role is being granted to users named "team lead" and "lab technician"; the Reader role is being granted to the "reviewer".
Remove Users and Groups from Roles
To remove a level of access for a given user or group, reopen the interface for granting that role and click the X for the user or group you want to delete from the role. Removing a user from a role does not deactivate or remove the user account itself.Related Topics
- User Accounts, Groups, and Roles
- Biologics Administration
- Security Roles Reference: More detail about all the available security roles.
- Freezer Storage Roles
- Sample Manager Permissions with Premium Editions of LabKey Server
Manage Groups
- User Groups
- Create Group
- Add Users to Groups
- Grant Permission Roles to Groups
- The "Users" Group
- Delete Group
User Groups
To access the group management page, select > Groups. All existing application groups will be shown.Create Group
To create a new group, type the name of the group and click Create Group.Add Users to Groups
Expand the group by clicking anywhere in the tile and use the Add member dropdown to add members. You can add individual users, or other groups, to a group. Each time you add a new member, you'll see User Details on the right.Grant Permission Roles to Groups
User groups can be added to permission roles just as users can. Learn more in this topic: When using the Professional Edition, if you want to be able to assign notebook review to a group, that group must be granted the "Editor" role (or higher).Users with the "Workflow Editor" role can assign workflow jobs and tasks to groups, as well as add groups to the notify list.The "Users" Group
Note that there is always a "Users" group predefined, sometimes used to represent every user with any access to the application, though this is not automatic. Users or groups must be explicitly added to this group. For example, you might add all the groups you define to the "Users" group in order to assign "Reader" permissions and ensure a minimal level of read access to every group member.Delete Group
To delete a group, you must first delete the members by clicking the X for each, then click Delete Empty Group.Related Topics
Manage Notifications
In-app Notifications
Notifications of interest to each user are shown in a menu in the header bar. If any background imports are in progress, the bell will be replaced with a . The number of waiting notices will be shown in orange, superimposed on the bell or spinner icon.Email Notifications
Users of the Professional Edition of Sample Manager can control whether they receive email for either Notebooks or Workflow or both. Email notifications allow users to learn about important events without having to manually check for them.- Notebook notifications are sent when notebooks the user is a part of are submitted, approved, or have changes requested.
- Workflow notifications are sent about events that occur regarding jobs or tasks assigned to the user or which they are following.
- Users can also find a dashboard of workflow jobs they are on the Notify List for on the Your Tracked Jobs dashboard.
- A task assigned to me is ready to be completed
- A job that includes tasks assigned to me is initiated
- A job that I owned and completed was reactivated by another user
- A comment was added to a task assigned to me
Receive Email Notifications
By default, all users will receive email notifications about Workflow and Notebook events that occur that they are part of, or are following. Users can choose to opt out of all email notifications by disabling these settings.Select Notification Settings from the user menu, then check the box for the notifications you want. Uncheck the box to disable receiving email notifications for the category.Related Topics
Sample Manager - FAQ
- Data Ownership and Audit
- Sample Identification
- Freezer Management
- Data Structures for Assays and Samples
- Jobs, Tasks, and Templates
- Sample Manager and LabKey Server
- Future Plans
Data Ownership and Audit
If I store my data in LabKey Sample Manager do I still own the data? If I choose to end my subscription later, will I be able to get it back?
Absolutely. Whether cloud-based or on premise, you always own your own data. If you stop using Sample Manager, you will receive a full export of all your data.Will privacy be maintained if I use Sample Manager?
Absolutely, the LabKey security model guarantees privacy and security using our role based access model.However, if you need to store PHI and/or are interested in HIPAA compliant protection of your data, contact us to discuss whether another LabKey product might better meet your compliance needs.Do samples have an audit trail for chain of custody tracking?
Yes! Every action is tracked in a set of audit logs on a row by row level. Enhanced chain of custody tracking features are available in a Timeline for samples. Learn more in this topic:Sample Identification
Is the sample ID assigned by the system unique to just one lab? Can they be shared?
Yes, right now because there is a single Sample Manager application serving each lab, if you ask the application to generate sample IDs for you, they will be unique within that single lab. However, letting the system assign sample IDs is not the only option.If you wanted to share sample information across multiple labs, you could override the automatic assignment option by providing your own unique sample IDs, such as by using a sample manifest. The distinct Sample Manager applications at many lab sites could accept and use these sample IDs that are drawn from a master assignment list ensuring that they are both unique within a single application and unique across multiple locations.Does Sample Manager handle replicates?
Yes, sample replicates can be identified using custom naming conventions, i.e. S101-1 and S101-2 are replicates of one original sample.Does Sample Manager track the subject of study, i.e the source of the sample.
Yes, you can identify and track many types of sample Sources within the application. Learn more in this section:Does Sample Manager support using a barcoding system?
Yes, you can create your own field (either text or integer) to hold your own barcode values. Or, with the addition of a "UniqueID" column, Sample Manager will generate unique barcodes for samples for you. These barcodes are read-only, simple, and easy to use. Learn more in this topic: Sample Manager also supports integration with BarTender for groups to use for printing labels with sample details. Learn more in these topics:Freezer Management
Does Sample Manager provide a freezer management solution?
Yes! LabKey Sample Manager includes a robust and flexible freezer management solution. Design virtual storage to match your physical storage. Easily find samples, or empty space for storing new samples. Track volume and check-in and check-out to support your workflow. Control access with specific storage roles. Learn more in this section:Data Structures for Assays and Samples
How do I get my existing sample data into the system?
LabKey Sample Manager is specifically designed to make data import easy. Design the structure of existing data and drag and drop to upload it simply and efficiently. Note that very large uploads may need to be split into batches to upload successfully.Use custom templates to make it easier to format your data.How does a user know what columns are expected?
The handy Download Template feature gives the user a blank template for what columns are expected. The user can either add their data to this template or simply confirm that they have the correct columns prior to import.Can I build in customized data integrity checks?
Absolutely. Every field can have data validation performed, such as ensuring correct formats, valid ranges, and other such measures. You can also use controlled vocabularies for text fields, i.e. presenting uploading users with pulldown menus of options instead of free text entry fields.Do you have support for tracking study visits, where multiple samples of different types are collected from one subject at once?
We don't have a built-in mechanism for tracking study visits at this time, but by defining additional custom columns for your sample types, you can track the individual and date of collection for matching. For example, using a required column for study visit, you would capture this information.What export types are supported?
Currently you can export data as CSV, TSV, and Excel. Multi-tabbed sample lists can be exported as multi-tabbed Excel spreadsheets.Are there predefined templates for data in Sample Manager?
There are no predefined templates. Users have full control of creating data templates for your own needs. During the definition of assays, Professional Edition users have the option to let the application infer fields in the data structure from the columns in a spreadsheet.Are the Assays customizable? Can I create my own assays? (Professional Edition Feature)
Absolutely, with the Professional Edition of Sample Manager. Our demos and examples include some possible ways to structure typical assays, but when you define your own, you have complete control over the fields and types of data collected. The only requirement is that assay data needs to provide a column linking to the sample.What happens if you import assay data but it has a column name that doesn't match?
The assay data import process will read only the "expected" columns from your data. If you have additional columns, they will be ignored. If you have a difference in column naming, you may be able to make use of column aliases to import data from a column of a mismatched name.When importing data, you will see a preview of the first few rows to aid you in correcting issues or adding aliases.Jobs, Tasks, and Templates (Professional Edition Feature)
Does each workflow job depend on the completion of the previous job? Or can you have multiple jobs underway simultaneously? Can you configure which job is dependent on which other job?
Each workflow job can begin/proceed independently of all other jobs. You can have as many jobs underway simultaneously as you like. If you want to have actions that proceed in a sequence, consider whether these should be defined as tasks within a single larger job, rather than separate jobs.In the future, we hope to add an administrative option to make a job dependent upon completion of another job, but at present this is not supported. In the meantime, you could also consider having a 'check for previous job completion' task at the start of the job you want to happen 'next'.Sample Manager and LabKey Server
Can assay results loaded via Sample Manager be linked to LabKey Studies?
Yes, if you are using the Professional Edition of Sample Manager as part of a Premium Edition of LabKey Server, your application will be running on the same server as your other LabKey projects. After loading assay data into Sample Manager, you can access it via traditional LabKey Server folder management tools and link that data into your study on the same server.Can Sample Manager make use of assays already defined in my LabKey Server?
Yes, if you have defined Standard Assays in the scope available to your integrated Professional Edition of Sample Manager, you will see them in the list of assay designs. You may need to map one of the columns to your sample information before you can use them.Future Plans
We are very interested in hearing your feedback about what is important to you. Future development of new features for LabKey Sample Manager is already underway.Do you need other software to do data analysis and generate reports?
Yes, at this time, users of Sample Manager export their sample data for analysis and reporting. In the future, analysis and reporting will be added within the application.Note that LabKey Server itself is a candidate for such analysis and reporting, and in fact, users of Premium Editions of LabKey Server can access data from Sample Manager directly from the traditional LabKey user interface.Does Sample Manager track reagents, vendor batch number, etc.?
Not explicitly at this time. You can use custom columns to track this information yourself. One option is to use a controlled vocabulary text field to track information and let users select from lists instead of free entering values.Related Topics
More Questions?
Have more questions or need additional information? Please get in touch with us:Request Support
New to Sample Manager?
Already a Subscriber?
Return to Documentation
Contact Us
Explore Sample Manager
- Take a self-guided tour of the application
- Sign up for a personalized demo to ask your questions
- Get started with your own trial version of the application where you can experience working with your own data
Ontology Concept Annotations
Once ontologies have been loaded and enabled in your folder, you can use Concept Annotations to link fields in your data with their concepts in the ontology vocabulary. A "concept picker" interface makes it easy for users to find desired annotations.
Browse Concepts
Reach the grid of ontologies available by selecting > Go To Module > More Modules > Ontology.
- Click Browse Concepts below the grid to see the concepts, codes, and synonyms loaded for any ontology.
- On the next page, select the ontology to browse.
- Note that you can shortcut this step by viewing ontologies in the "Shared" project, then clicking Browse for a specific row in the grid.
- Type into the search bar to immediately locate terms. See details below.
- Scroll to find terms of interest, click to expand them.
- Details about the selected item on the left are shown to the right.
- The Code is in a shaded box, including the ontology prefix.
- Any Synonyms will be listed below.
- Click Show Path or the Path Information tab to see the hierarchy of concepts that lead to the selection. See details below
Search Concepts
Instead of manually scrolling and expanding the ontology hierarchy, you can type into the search box to immediately locate and jump to concepts containing that term. The search is specific to the current ontology; you will not see results from other ontologies.
As soon as you have typed a term of at least three characters, the search results will populate in a clickable dropdown. Only full word matches are included. You'll see both concepts and their codes. Click to see the details for any search result. Note that search results will disappear if you move the cursor (focus) outside the search box, but will return when you focus there again.
Search terms will not autocomplete any suggestions as you type or detect any 'stem' words, i.e. searching for "foot" will not find "feet".
Path Information
When you click Show Path you will see the hierarchy that leads to your current selection.
Click the Path Information for a more complete picture of the same concept, including any Alternate Paths that may exist to the selection.
Add Concept Annotation
- Open the field editor where you want to use concept annotations. This might mean editing the design of a list or the definition of a dataset.
- Expand the field of interest.
- Under Name and Linking Options, click Select Concept.
- In the popup, select the ontology to use. If only one is loaded, you will skip this step.
- In the popup, browse the ontology to find the concept to use.
- Use the searching and browsing options to locate the concept you want.
- Click it in the left panel to see the code (and any synonyms) on the right.
- Click Apply.
- You'll see the concept annotation setting in the field details.
- Save your changes.
View Concept Annotations
In the data grid, hovering over a column header will now show the Concept Annotation set for this field.
Edit Concept Annotation
To change the concept annotation for a field, reopen the field in the field editor, click Concept Annotation, make a different selection, and click Apply.
Related Topics
Help Links
Click to Explore
Samples
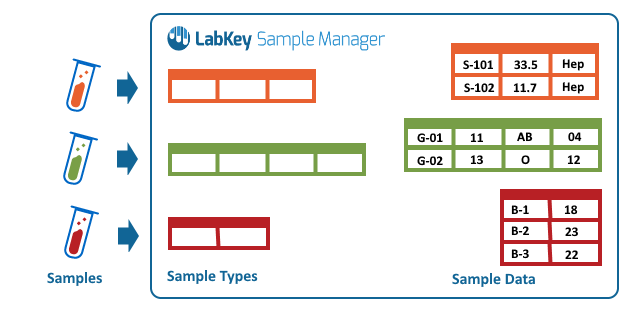 This section covers the creation and management of sample types and samples.
This section covers the creation and management of sample types and samples.
Topics
- Create a Sample Type
- Add Samples:
- Sample ID Naming
- Sample Lineage / Parentage
- LIMS: Samples
- Sample Expiration
- Sample Timeline
- Manage Samples
- Manage Sample Status
- Edit Selected Samples
- Sample Finder
- Sample Search
- Print Labels with BarTender
- Barcode Fields
- Sample Picklists
Related Topics
- Sources: Sources can be physical, biological, or a combination of both.
- Storage Management
Create a Sample Type
- Video Overview
- Create a New Sample Type
- Define Sample Type Properties
- Define Sample Type Fields
- Configure Sample/Aliquot Field Inheritance
- Finish Creating Sample Type
- Edit Sample Type Design
- Resource: Sample Naming Pattern Examples
Video Overview
In this video you will learn how to create a new Sample Type.Create a New Sample Type
Create a new Sample Type by clicking Sample Types on the main header menu, then selecting Create > Sample Type.Define Sample Type Properties
- The Name of the Sample Type is required and must be unique.
- The Name must start with a letter or number character, and avoid special characters and some reserved substrings listed here.
- You can edit the Name later if needed, unless an admin has disabled this option.
- Entering a Description is optional. A description can help others understand the usage.
- Naming Pattern: Every sample must have a unique name or identifier, known as a "Sample Id". Unless you will always provide these sample IDs yourself you should provide a pattern to generate them for you.
- Hover over the to see an example name using the current pattern.
- Details about customizing naming patterns are below.
- Aliquot Naming Pattern: For aliquots created from samples of this type, you can leave this blank to accept the default naming pattern, or customize it if desired.
- Provide any Lineage Settings needed.
- Set Storage Settings.
- Define a Barcodes field if desired.
- Before you click Finish Creating Sample Type, determine whether you need to add Fields.
Naming Pattern
Every sample must have a unique name or identifier, known as a "Sample Id". For each Sample Type, determine whether you will provide these sample IDs or whether you want the system to generate them for you. This decision determines what, if anything, you will enter in the Naming Pattern field. Learn more in this topic: Sample ID Naming.- If you will provide the sample IDs, and they are in your data in a column named "SampleID", you can delete the default naming pattern (and ignore the grayed out placeholder text).
- If you will provide the sample IDs, but they are in a column named something other than "SampleID", you provide a naming pattern like this pattern ${COLUMN_NAME}. See below.
- If you want sample identifiers generated for you by the system, provide a naming pattern using the guidance in this topic: Sample ID Naming. A few examples are at the bottom of this page.
- The default naming pattern is "S-${genId}", meaning the letter S followed by a dash and an incrementing counter.
Aliquot Naming Pattern
By default, the name of the aliquot will use the name of its parent followed by a dash and a counter for that parent's aliquots.${${AliquotedFrom}-:withCounter}${${AliquotedFrom}.:withCounter}Lineage Settings
- To declare a field that will identify a parent sample, click Add a Parent. Select the Sample Type of the parent and enter the File Import Column Name.
- If there are Sources defined in the system, you will have a similar option to Add a Source here.
Storage Settings
- Label Color: You can assign a color to this type of sample to help make quick visual identification easier in various application views.
- Amount Type
- Any: Amounts can be entered in any Mass, Volume, or Other unit listed below. Amounts entered won't be converted for storage or display.
- Mass: Samples may be registered with these units: ng, ug, g, mg, or kg. Amounts entered will be converted for storage and display. For storage in the database, user-entered amounts are converted to the base mass unit mg. For display, the amounts will be converted to the display unit configured by the administrator.
- Volume: Samples may be registered in these units: uL, mL, or L. Amounts entered will be converted for storage and display. For storage in the database, user-entered amounts are converted to the base volume unit mL. For display, the amounts will be converted to the display unit configured by the administrator.
- Other: Samples may be registered in these units: unit, pieces, packs, blocks, slides, cells, boxes, kits, tests, bottles. Amounts entered won't be converted for storage or display.
- Amount Display Units: Amounts will be converted to this unit for display.
Label Color
To assign a Label Color when you are creating or editing your sample type, click the selection area to open a color picker:- Click a block of color to select it.
- Type a hex code next to the # sign or individual RGB values into the panel for more color control.
Define Sample Type Fields
Click the Fields section to open it.Default System Fields
Every Sample Type is created with several default fields (columns) built in. For example, like any data structure, "Created/CreatedBy/Modified/ModifiedBy" record the userID and time of creation/modification respectively. In addition, you will see the Sample Type Default System Fields at the top of the Fields panel.
- SampleID: The name of the sample.
- Status (SampleState): Tracks the status of the sample.
- Description: This is a description field for an individual sample, not the description of the Sample Type as a whole that is shown in the properties section.
- Expiration Date (MaterialExpDate): Tracks the date this sample expires on.
- Amount (StoredAmount): The amount of this sample.
- Units: The units associated with the Amount value for this sample. This may be different from the "Amount Display Units" set for the Sample Type.
- Aliquots Created Count (AliquotCount)
- Freeze/Thaw Count (FreezeThawCount)
- Storage Location, Row, and Column
Add Custom Sample Type Fields
Add any additional fields that you want in your Sample Type definition as follows:- Click the Fields section to reopen it if needed.
- You can collapse the Default System Fields section to hide it.
Import or Infer Custom Fields from File
- This will populate the fields section based on inferrals from the data.
- Note: If your spreadsheet contains columns for built-in fields, such as "SampleID" and "Description", they will not be shown here.
- You can also import field definitions from a prepared JSON file if you have one.
Manually Define Fields
- For each additional field in your sample data, click Add Field:
- Enter the Name. Names should not contain spaces or special characters; instead you can use such characters in the label for the field. Learn more in this topic.
- Select the Data Type. Learn more about types and their properties in this topic.
- Check the box if the field is required.
- Click the (expansion) icon to define more properties of your new field as needed.
- You can reorder the fields by dragging and dropping with the six-block handle.
- Add all the fields you need. If you add an extra and need to delete it, click the on the right.
Barcodes
If a field of type Unique ID is included in your Sample Type, the system will generate barcode values for you.When you are defining a new sample type, you will see in the initial setting of the Barcodes property that no Unique ID field exists. You will be prompted to include one after inferring or manually creating other fields.Configure Sample/Aliquot Field Inheritance
For each field included in the Sample Type, whether inferred or manually created, you can configure whether the field is settable (and editable) for samples, aliquots, or both.- Click the to expand each field's details.
- Under Sample/Aliquot Options, select one:
- Editable for samples only (default): Aliquots will inherit the value of the field from the sample.
- Editable for aliquots only: Samples will not display this field, but it will be included for aliquots.
- Separately editable for samples and aliquots: Both samples and aliquots can set a different value for this property.
Finish Creating Sample Type
- Click Finish Creating Sample Type when finished.

Edit Sample Type Design
If necessary, you can return to edit properties or fields/columns later via Manage > Edit Sample Type Design.Resource: Sample Naming Pattern Examples
A few examples of how you might choose to name your samples and how to design a naming expression. You can use values from your data, dates, and counters to ensure that every sample name is unique. Separators like '-' and '_' are commonly used but optional.| Description | Example SampleIDs | Naming Pattern |
|---|---|---|
| The character 'S' followed by a counter (Default) (always unique within this type of sample) | S-1 S-2 S-3 | S-${genId} |
| The word 'Blood' followed by a 2 digit counter | Blood-01 Blood-02 Blood-03 | Blood-${genId:number('00')} |
| The value of the "Lab" column + '_' + a counter | LabA_1 LabB_2 LabC_3 | ${Lab}_${genId} |
| Use a default value if the "Lab" column is empty (+ '_' + a counter) | LabA_1 LabB_2 LabUnknown_3 | ${Lab:defaultValue('LabUnknown')}_${genId} |
| 'S' plus current date followed by a counter that resets daily (always unique; tracks date of entry into system) | S-20200204-1 S-20200204-2 S-20200205-1 | S-${now:date}-${dailySampleCount} |
| 'S' plus values from two columns in my data: the Study and ParticipantID (only unique if one sample per participant) | S-Study101-pt231 S-Study101-pt232 S-Study101-pt233 | S-${Study}-${ParticipantId} |
| Values of "ParticipantID" and "Draw Date" columns, then a counter (always unique and tracks draw date, not date of entry into system) | pt231-20191203-1 pt231-20191214-2 pt231-20191214-3 | ${ParticipantID}-${DrawDate}-${genId} |
| The values already present in the "SampleID" column in my data | unique values provided | No naming pattern required |
| The values already present in the "Identifier" column in my data | unique values provided | ${Identifier} |
Related Topics
Add Samples
- Create New Samples
- Assign Amount and Units
- Work with New Samples
- Create Derivatives, Pooled Samples, and Aliquots
- Aliquot the Selected Sample(s)
- Derive from Selected
- Pool Selected
Create New Samples
To add new samples, you either navigate to the type of sample you are adding and use the Add menu above the grid, or go to the home page or the Sample Types dashboard and click Add Samples. In both places you choose either:Create Samples From Grid
Using the Add Manually option gives you several options for entering sample information into the application directly, as opposed to uploading a file of data. You can manually enter values for the fields in a grid format, or use bulk insert and bulk update to streamline entry of similar values.Learn more in this topic: Create Samples from GridImport Samples From File
The other method for entering samples, particularly useful for a large group or when sample information is available in a spreadsheet already, is to import directly from a file.Learn more in this topic: Import Samples from FileAssign Amount and Units
During Sample import, you can provide Amount and Units values for each sample.The Amount and Units fields are enforced as a pair — both must be provided together or both left blank. This rule applies in all contexts, including file import, grid editing, bulk updates, and manual entry. In the field designer, when you select or deselect one of these fields, the other field will be selected or deselected accordingly. In pop-up editing boxes, these fields appear on the same line, emphasizing that they are paired fields.Negative values cannot be entered in the Amount field. Users cannot manually specify a negative amount value when inserting or updating. Also, amounts cannot be decremented to a negative value.When the Amount Type is configured as Mass or Volume, user-entered amounts are converted to the "base unit" for storage in the database: milligrams (mg) for Mass, milliliters (mL) for Volume. User-entered amounts are also converted for display. For example, if the display unit is mL, and the user enters amount 1 and unit L, then 1000mL will be displayed. Learn more about display values for amounts and units here.Original user-entered amounts are retained and can be viewed in the sample Timeline and Audit Log.Whether a Sample is or is not in freezer storage within the application, you'll be able to see the amount displayed in an Inventory/Storage Details panel of the sample details.Sample amount values can be used to generate "low volume" reports in the Sample Finder.Also see: Create a Sample TypeWork with New Samples
After creating samples, the banner message indicates how many were created. In that message, click select them in the grid to select this recently created batch for actions like adding them to a picklist or workflow job. You can also click Add them to storage to immediately add the new samples to freezer storage locations.Create Derivatives, Pooled Samples and Aliquots
Once you have created the Samples you want to use as parents, you can also create new samples as derivatives, pooled samples, and aliquots. Learn more in these topics:Related Topics
- Create a Sample Type
- Create Samples from Grid
- Import Samples from File
- Manage Sample Status
- Attach Images and Other Files
- Sample Lineage / Parentage
- Associate Samples with Sources: Create new samples from Sources of those samples.
- LIMS: Samples
- Data Import Guidelines
- Store Samples
- Migrate Storage Data into LabKey
Create Samples from Grid
Add Manually
Open the grid method for creating new Samples from the desired Sample Type by selecting Add > Add Manually. You can also choose Add Samples > Add Manually from the main dashboard.Select Sample Type and Number
In the popup, select the Sample Type and provide the Number of Samples.Add Samples of Multiple Types
You can simultaneously add samples of multiple types by using Add Another Sample Type, providing a second type and number of samples.Add Parent
If the Sample Type has one or more Parent samples, you will see a prepopulated parent field. In cases where the parent field is marked as Required you will need to provide a value, otherwise it is optional. To add more, click Add Parent and select the Sample Type of the parent(s). If you will have parents of different types, add one for each type of parent.Add Source
If the Sample Type has one or more Sources, you will see a prepopulated source field. In cases where the source field is marked as Required you will need to provide a value, otherwise it is optional. Click Add Source and select the type of Source you want to associate with samples of this type.Add Sample Data in Grid
The data entry grid will have the number of samples you requested in the modal. To add more rows, you can enter the number and click Add Samples. Remove excess rows by selecting them and clicking Delete.- Sample ID:
- If you provided a Naming Pattern with the Sample Type, the Sample ID can be generated, and this will be noted in the grid. Hover over the to see an example of a generated name.
- You could override this value by manually typing in the box, but remember that all sample IDs must be unique.
- Parent and Source Fields:
- If the Sample Type includes any parent or source lineage settings, there will be columns for the Sample or Source Types of these fields included by default.
- Enter values for remaining fields using the guidance in this topic
- Choose a value from fields with a .
- Enter data directly into the grid. Copy and paste of cells is supported, as is dragging to propagate text or number series values.
- When ready, click Finish Creating ## Samples.
Bulk Add
Instead of directly populating a grid, you can click Bulk Add to create many samples at once. This is particularly convenient when samples share the same settings for some or all columns.- Specify the number of New Samples.
- Note that if you have included a parent field, you will have additional options here.
- Enter values for any or all of the properties listed that you want to be applied to all the new samples.
- Click Add Samples To Grid.
Bulk Insert with Parent(s)
If you added one or more parent fields before clicking Bulk Add, you will have an additional selection to make in the bulk creation popup. Choose whether you want to use the parent sample(s) to create:- Derivatives, specifying the number of derivatives per parent.
- Pooled Samples, specifying the number of new samples to create for the pooled parent(s).
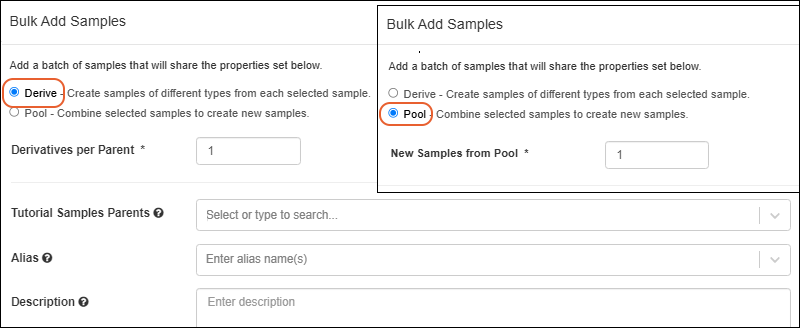
- Provide values for properties that will be shared by the newly created samples. Including 'parents' as applicable.
- Hover over the for the Status field to see a legend of available statuses.
- Click Add Samples to Grid and continue to complete the grid with any properties not shared by all the new samples.
- Click Finish Creating ## Samples when done.
Edit in Bulk
While creating new samples in the grid, you can use the Edit in Bulk option to assign a common value to a selected set of rows. As for bulk update of assay data rows, you control which fields are Enabled for update to a new common value you provide. The disabled fields will retain their original values.Related Topics
Import Samples from File
Obtain File Import Template
An import template shows all the expected columns for your data structure, making it easier to ensure your imported data will conform to system expectations. Templates are available for Sample Types, Source Types, and Assays from the main dashboard for the type of structure.From the main menu, select the target Sample Type. Click the Template button to download the template for this type of sample.
- Use this template as a guide for ensuring your data matches the expected columns.
- Note that any "File" fields will be omitted from the downloaded template.
- When populating the template, you may not need to include all columns. For example, if you have defined lineage import columns (aliases) for sources or parents, all possible columns will be included in the template, but you only need to include the ones you want to use.
Import Samples From File
You can start from the home page of the application or the Sample Types dashboard and select Add Samples > Import from File or from the overview page for a specific Sample Type, select Add > Import from File.- Set the Sample Type for the Samples you will import.
- If you started from a specific sample type grid, the type will be prepopulated and you will not see the dropdown.
- If you want to import samples of multiple types, select Import multiple sample types. Learn more below.
- If you are including storage information for the samples you're importing, select the appropriate radio button:
- Add samples to existing storage only: all storage locations must exist already.
- Add samples to existing or new storage (created during import): a combination of existing and new storage units are allowed.
- Use the Template button if you didn't already download a template file for your data.
- Drag and Drop the file into the target area to upload it.
- You will see the sample information obtained from the file in the panel.
- Only fields included in the definition of the Sample Type will be imported. In this example, the Sample ID will be generated for you, so is not included in the imported file or shown here. If any fields are unrecognized, they will be ignored and a banner will be shown.
- Note that you cannot edit the values here. If you see an error, you can use the X to delete the loaded file, make changes offline, and reselect the revised file.
- Click Import to create the samples from the data in the file.
Import Samples Across Multiple Sample Types
To import samples across multiple Sample Types, select the Sample Type option Multiple sample types.Import Samples and Create New Storage
The default Storage Option when you import samples from file is to Add samples to existing storage only, provided the location details are included in the file. Learn more about the details that you need to provide here.To create any new storage locations while importing samples from file, i.e. to assign new samples to either new or existing storage locations within the same file, a user with the "Storage Designer" role can select the option Add samples to existing or new storage (created during import).- If any locations in that path do not exist, they will be created.
- The "StorageLocation" field value should be slash-separated and must end with a terminal storage unit.
- If any levels in this field value do not exist, they will be created.
- You can also create an entirely new storage system (freezer) with this process, using a value like "MyNewFreezer/Shelf1/Box1" where "MyNewFreezer" does not already exist.
- You can also precede new storage with a / to make it clear that a new top level system is to be created, i.e. "/MyNewFreezer/Shelf1/Box1".
- If the terminal storage unit (final element in the field value) does not already exist, it will be created as the type of unit specified in the "Storage Unit" field.
- This means that only the first row for a new storage unit needs to provide the correct type of unit to create.
- If all units in the full "StorageLocation" field already exist, the "Storage Unit" value is not read for that row.
Update Existing Samples
To update existing samples, or merge a combined spreadsheet of new and existing samples, select Edit > Update from File. The update page is very similar to the Import from File page, with additional Update Options. The default is Only update existing samples.- Any data you provide for the rows representing samples that already exist will replace the previous values.
- When updating, you only need to provide the columns you wish to update. Existing data for other columns will be left as is. If you wish to update the SampleId values, then you must include the RowId column in your file.
- Important: All columns you upload will be updated in the system. If you include an empty column, all data for that data will be deleted.
- Note that you cannot update an existing sample to make it an aliquot of another sample.
- Before clicking Import, provide a Reason for Update if desired or required.
Merge by Allowing New Samples
The Create new samples too update option controls whether you will be able to merge the incoming file or simply update existing rows.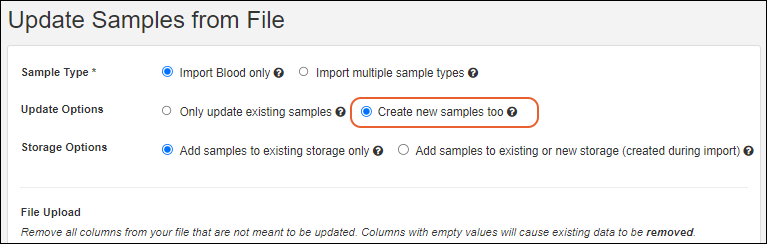
- When this option is not selected, the default, the update will fail if rows are included that do not match existing samples already in the system.
- When selected, any rows for samples that do not yet exist will add them as new samples.
- As for updates without merge, before clicking Import, provide a Reason for Update if desired or required.
Related Topics
Sample ID Naming
- Unique Sample Names are Provided: Unique sample names are provided to the system
- Generate Names with Naming Patterns: Generate unique names within the system
- Default Naming Pattern
- See and Set genId
- Date Based Naming
- Incorporate Column Values
- Incorporate Lineage Lookups
- Include Ancestor Names/Properties Using "~"
- Include Grandparent Names/Properties Using ".."
- Naming Pattern Elements/Tokens
- Incrementing Sample Counters
- Naming Pattern Validation
- Examples
Unique Sample Names are Provided
If your data already includes the unique sample names to use, identify the column name that contains them.Naming Column is "SampleID" or "Name"
If the name of this column is "SampleID" or "Name", these default column names are automatically recognized as containing sample names. To confirm that they are used, be sure to Delete the default naming pattern that is provided in the user interface (and ignore the grayed out placeholder text that remains).Naming Column is Something Else
If the column containing unique sample names is named something else, you provide that column name using a simple naming pattern expression that specifies the name of the column to use, rather than an expression to generate one.For example, if the sample names are in a column named "Identifier", you would enter the naming pattern:${Identifier}Generate Names with Naming Patterns
If your data does not already contain unique names, the system can generate them upon import using a naming pattern that contains tokens, including counters to ensure names are unique. The system can build a unique name from syntax elements, such as:- String constants
- Incrementing numbers
- Dates and partial dates
- Values from columns in the imported data, such as tissue types, lab names, subject ids, etc.
- Separators such as '_' underscores and '-' hyphens
- Note that if you use a hyphen '-', you will want to use double quotes when you later search for your samples. An unquoted search for Sample-11 would interpret the hyphen as a minus sign and seek pages with "Sample" without "11".
Default Naming Pattern
The default naming pattern in Sample Manager generates names from two elements: the prefix "S-" plus an incrementing integer.S-${genId}S-1
S-2
S-3
and so on...
See and Set genId
The token genId is an incrementing value which starts from 1 by default, and is maintained internally for each sample type (or source type) in a container. Note that while this value will increment, it is not guaranteed to be continuous. For example, any creations by file import will 'bump' the value of genId by 100, as the system is "holding" a batch of values to use.When you include ${genId} in a naming pattern, you will see a blue banner indicating the current value of genId.Date Based Naming
Another possible naming pattern for samples is to incorporate the date of creation. For example:S-${now:date}-${dailySampleCount}- The S- prefix is simply a string constant with a separator dash. Using separators like "-" and "_" is optional but will help users parse sample names.
- The now:date token will be replaced by the date of sample creation.
- The dailySampleCount token will be replaced by an incrementing counter that resets daily.
Incorporate Column Values
If you want to use a column from your data as part of the name, but it does not contain unique values for all samples, you can incorporate it in the pattern by using the column name in token brackets and also including an additional uniqueness element like a counter. For example, if you want to name many samples for each participant, and the participant identifier is in a "ParticipantID" column, you could use the pattern:${ParticipantID}-${genId}${ParticipantID}-${CollectionDate}-${LabName}-${now:date}-${genId}Incorporate Lineage Lookups
More general syntax to include properties of sample sources or parents in sample names is also available by using lookups into the lineage of the sample.- Specific data type inputs: MaterialInputs/SampleType1/propertyA, DataInputs/DataClass2/propertyA, etc.
- Import alias references: parentSampleA/propertyA, parentSourceA/propertyA, etc.
- In some scenarios, you may be able to use a shortened syntax referring to an unambiguous parent property: Inputs/propertyA, MaterialInputs/propertyA, DataInputs/propertyA
- This option is not recommended and can only be used when 'propertyA' only exists for a single type of parent. Using a property common to many parents, such as 'Name' will produce unexpected results.
Blood-${DataInputs/Mouse/Strain}-${genId}Blood-${parentBlood/SampleID:first}-${genId}Include Ancestor Names/Properties Using "~"
To reference an ancestor name (or other property), regardless of the depth of the ancestry, you can use syntax that includes a tilde and will 'walk the lineage tree' to the named Source or Sample Type regardless of depth of a lineage tree (up to a maximum depth of 20). Note that this type of syntax may be more resource intensive, so if you know that the ancestor will always be the direct parent or at another specific/consistent level, you should use another lineage lookup for efficiency.For example, consider a "Participant" Source Type and also a Sample Type like "Blood" that could be either a direct 'child' of the source, or a grandchild (of an intermediate sample like "Tissue"), or any further descendent. You can include properties of the Participant source of a "Blood" sample with a naming pattern like this:${~DataInputs/Participant/Name}
${~DataInputs/Participant/OtherProperty}${~MaterialInputs/SampleTypeName/Name}
${~MaterialInputs/SampleTypeName/OtherProperty}${${~DataInputs/Participant/Name}-:withCounter}Include Grandparent Names/Properties Using ".."
If you know that the ancestor of interest is a fixed number of generations above the direct parent, i.e. the grandparent or great-grandparent generation, you can use ".." syntax to walk the tree to that specific level. Here are a few examples of syntax for retrieving names from the ancestry of a sample. For simplicity, each of these examples is shown followed by a basic ${genId} counter, but you can incorporate this syntax with other elements. Note that the shortened syntax available for first generation "parent" lineage lookup is not supported here. You must specify both the sample type and the "/name" field to use.To use the name from a specific grandparent sample type, use two levels:${MaterialInputs/CurrentParent/..[MaterialInputs/GrandParentSampleType]/name}-${genId}${MaterialInputs/CurrentParent/..[MaterialInputs/GrandParentSampleType]/propertyColumn}-${genId}${parentAlias/..[MaterialInputs/GrandParentSampleType]/name}-${genId}${MaterialInputs/CurrentParent/..[MaterialInputs/GrandParentSampleType]/..[MaterialInputs/GreatGrandSampleType]/greatGrandFieldName}-${genId}${plasmaParent/..[MaterialInputs]/name}-${genId}
${MaterialInputs/Plasma/..[MaterialInputs]/name}-${genId}
${MaterialInputs/..[MaterialInputs]/name}-${genId}Naming Pattern Elements/Tokens
The following elements, or "tokens" are available for building naming patterns.| Name Element | Description | Scope0000000000000000000000000 |
|---|---|---|
| genId | An incrementing number starting from 1. This counter is specific to the individual Sample Type in a given container. Not guaranteed to be continuous. | Current Sample Type |
| sampleCount | A counter incrementing for all samples of all Sample Types, including aliquots. Not guaranteed to be continuous. Learn more below. | All Sample Types in the application (home plus any folders) |
| rootSampleCount | A counter incrementing for non-aliquot samples of all Sample Types. Not guaranteed to be continuous. Learn more below. | All Sample Types in the application (home plus any folders) |
| dailySampleCount | An incrementing counter, starting with the integer '1', that resets each day. Can be used standalone or as a modifier. | All Sample Types and Source Types on the site |
| weeklySampleCount | An incrementing counter, starting with the integer '1', that resets each week. Can be used standalone or as a modifier. | All Sample Types and Source Types on the site |
| monthlySampleCount | An incrementing counter, starting with the integer '1', that resets each month. Can be used standalone or as a modifier. | All Sample Types and Source Types on the site |
| yearlySampleCount | An incrementing counter, starting with the integer '1', that resets each year. Can be used standalone or as a modifier. | All Sample Types and Source Types on the site |
| randomId | A four digit random number for each sample row. Note that these random numbers are not guaranteed to be unique. | Current Sample Type |
| batchRandomId | A four digit random number applied to the entire set of incoming sample records. On each import event, this random batch number will be regenerated. | Current Sample Type |
| now | The current date, which you can format using string formatters. | Current Sample Type |
| Inputs | A collection of all DataInputs and MaterialInputs for the current sample. You can concatenate using one or more values from the collection. | Current Sample Type |
| DataInputs | A collection of all DataInputs for the current sample. You can concatenate using one or more values from the collection. | Current Sample Type |
| MaterialInputs | A collection of all MaterialInputs for the current sample. You can concatenate using one or more values from the collection. | Current Sample Type |
| <SomeDataColumn> | Loads data from some field in the data being imported. For example, if the data being imported has a column named "ParticipantID", use the element/token "${ParticipantID}" | Current Sample Type |
Formatting Values
You can use formatting syntax to control how the tokens are added. For example, "${genId}" generates an incrementing counter 1, 2, 3. If you use a format like the following, the incrementing counter will have three digits: 001, 002, 003.${genId:number('000')}:defaultValue Modifier
When you are using a data column in your string expression, you can specify a default to use if no value is provided. Use the defaultValue modifier with the following syntax. The 'value' argument provided must be a String in ' single quotes.${ColumnName:defaultValue('value')}:minValue Modifier
Tokens including genId, sampleCount, and rootSampleCount can be reset to a new higher 'base' value by including the :minValue modifier. For example, to reset sampleCount to start counting at a base of 100, use a naming pattern with the following syntax:S-${sampleCount:minValue(100)}S-${sampleCount:minValue(100):number('0000')}Names Containing Commas
It is possible to include commas in Sample and Source names, though not a best practice to do so. Commas are used as sample name separators for lists of parent fields, import aliases, etc., so names containing commas have the potential to create ambiguities.If you do use commas in your names, whether user-provided or LabKey-generated via a naming pattern, consider the following:- To add or update lineage via a file import, you will need to surround the name in quotes (for example, "WC-1,3").
- To add two parents, one with a comma, you would only quote the comma-containing name, thus the string would be: "WC-1,3",WC-4.
- If you have commas in names, you cannot use a CSV or TSV file to update values. CSV files interpret the commas as separators and TSV files strip the quotes 'protecting' commas in names as well. Use an Excel file (.xlsx or .xls) when updating data for sample names that may include commas.
Incrementing Sample Counters
The genId token is a basic incrementing value, but you can incorporate other ways of including counts in naming patterns. Some auto-incrementing counters calculate the next value based on all samples and sources across the entire application, while others calculate based on only the current Sample Type in the current container. See the Scope column for the specific incrementing behavior. When the scope is application-based, within a given container values will be sequential but not necessarily contiguous.sampleCount Token
When you include ${sampleCount} as a token in your naming pattern, it will be incremented for every sample created in the application (the home and any folders it contains), including aliquots. This counter value is stored internally and continuously increments, regardless of whether it is used in naming patterns for the created samples.For example, consider a system of naming patterns where the Sample Type (Blood, DNA, etc) is followed by the sampleCount token for all samples, and the default aliquot naming pattern is used. For "Blood" this would be:Blood-${sampleCount}
${${AliquotedFrom}-:withCounter}| Sample/Aliquot | Name | value of the sampleCount token |
|---|---|---|
| sample | Blood-1000 | 1000 |
| aliquot | Blood-1000-1 | 1001 |
| aliquot | Blood-1000-2 | 1002 |
| sample | DNA-1003 | 1003 |
| aliquot | DNA-1003-1 | 1004 |
| sample | Blood-1005 | 1005 |
Blood-${sampleCount} <- for samples
Blood-${sampleCount} <- for aliquots| Sample/Aliquot | Name |
|---|---|
| sample | Blood-1000 |
| aliquot | Blood-1001 |
| aliquot | Blood-1002 |
| sample | DNA-1003 |
| aliquot | DNA-1004 |
| sample | Blood-1005 |
- Addition of samples (and/or aliquots) of any type anywhere in the application will increment the token(s).
- Any failed sample import would increment the token(s).
- Import using merge would increment the token(s) by more than by the new number of samples. Since we cannot tell if an incoming row is a new or existing sample for merge, the counter is incremented for all rows.
rootSampleCount Token
When you include ${rootSampleCount} as a token in your naming pattern, it will be incremented for every non-aliquot (i.e. root) sample created in the application (the home and any folders it contains). Creation of aliquots will not increment this counter, but creation of any Sample of any Sample Type will increment it.For example, if you use the convention of using the Sample Type (Blood, DNA, etc) followed by the rootSampleCount token for all samples, and the default aliquot naming pattern, for "Blood" this would be:Blood-${rootSampleCount}
${${AliquotedFrom}-:withCounter}| Sample/Aliquot | Name | value of rootSampleCount | value of sampleCount |
|---|---|---|---|
| sample | Blood-100 | 100 | 100 |
| aliquot | Blood-100-1 | 100 | 101 |
| aliquot | Blood-100-2 | 100 | 102 |
| sample | DNA-101 | 101 | 103 |
| aliquot | DNA-101-1 | 101 | 104 |
| sample | Blood-102 | 102 | 105 |
Date Based Sample Counters
Date-based sample counters are available that will be incremented based on the date when the sample is inserted. These counters are incrementing, but since they apply to all sample types, within a given Sample Type, values will be sequential but not necessarily contiguous.- dailySampleCount
- weeklySampleCount
- monthlySampleCount
- yearlySampleCount
- As standalone elements of a name expression, i.e. ${dailySampleCount}, in which case they will provide a counter across all sample types and source types based on the date of creation.
- As modifiers of another date column using a colon, i.e. ${SampleDate:dailySampleCount}, in which case the counter applies to the value in the named column ("SampleDate") and not the date of creation.
:withCounter Modifier
Another alternative for adding a counter to a field is to use :withCounter, a nested substitution syntax allowing you to add a counter specific to another column value or combination of values. Using :withCounter will always guarantee unique values, meaning that if a name with the counter would match an existing sample (perhaps named in another way), that counter will be skipped until a unique name can be generated.The nested substitution syntax for using :withCounter is to attach it to an expression (such as a column name) that will be evaluated/substituted first, then surround the outer modified expression in ${ } brackets so that it too will be evaluated at creation time. The counter is applied to the case-insensitive version of the inner expression, such that if there is a case-only mismatch in a column (Blood/blood), the counter will not 'restart' for the differently cased variation.This modifier is particularly useful when naming aliquots which incorporate the name of the parent sample, and the desire is to provide a counter for only the aliquots of that particular sample. The default naming pattern for creating aliquots combines the value in the AliquotedFrom column (the originating Sample ID), a dash, and a counter specific to that Sample ID:${${AliquotedFrom}-:withCounter}${Blood-${Lot}-:withCounter}${Lot}-${Blood-:withCounter}${${Date:date}-${Lot}-:withCounter}${${AliquotedFrom}-:withCounter(42,'000')}Naming Pattern Validation
During creation of a Sample Type, both sample and aliquot naming patterns will be validated. While developing your naming pattern, the admin can hover over the for a tooltip containing either an example name or an indication of a problem.- Invalid substitution tokens (i.e. columns that do not exist or misspellings in syntax like ":withCounter").
- Keywords like genId, dailySampleCount, now, etc. included without being enclosed in braces.
- Mismatched or missing quotes, curly braces, and/or parentheses in patterns and formatting.
- Use of curly quotes, when straight quotes are required. This can happen when patterns are pasted from some other applications.
Caution: Escaping Special Characters
When field names or data type names contain special characters, including but not limited to {,}, and \, these characters must be escaped with a backslash \.Caution: Using Numbers-Only as Sample IDs
Note that while you could create or use sample names that are just strings of digits, you may run into issues if those "number-names" overlap with row numbers of other samples. In such a situation, when there is ambiguity between sample name and row ID, the system will presume that the user intends to use the value as the name.Examples
| Naming Pattern | Example Output | Description |
|---|---|---|
| S-${genId} | S-101 S-102 S-103 S-104 | S- + a simple sequence |
| ${Lab:defaultValue('Unknown')}_${genId} | Hanson_1 Hanson_2 Krouse_3 Unknown_4 | The originating Lab + a simple sequence. If the Lab value is null, then use the string 'Unknown'. |
| S-${now:date}-${dailySampleCount} | S-20170202-1 S-20170202-2 S-20170202-3 S-20170202-4 | S- + the current date + "-" + daily resetting incrementing integer |
| S-${Column1}-${Column2} | S-Plasma-P1 S-Plasma-P2 | Create an id from the letter 'S' and two values from the current row of data, separated by dashes. |
Example String Modifiers
The following naming patterns show usage of string modifiers.| Naming Pattern00000000000000000000000000000000000000000000000 | Example Output000000000000000000 | Description |
|---|---|---|
| S-${Column1}-${now:date}-${batchRandomId} | S-Blood-20170103-9001 | |
| S-${Column1:suffix('-')}${Column2:suffix('-')}${batchRandomId} | S-Blood-PT101-5862 | |
| ${Column1:defaultValue('S')}-${now:date('yy-MM-dd')}-${randomId} | Blood-17-01-03-2370 S-17-01-03-1166 | ${Column1:defaultValue('S')} means 'Use the value of Column1, but if that is null, then use the default: the letter S' |
| ${DataInputs:first:defaultValue('S')}-${Column1} | Nucleotide1-5 S-6 | ${DataInputs:first:defaultValue('S')} means 'Use the first DataInput value, but if that is null, use the default: the letter S' |
| ${DataInputs:join('_'):defaultValue('S')}-${Column1} | Nucleotide1_Nucleotide2-1 | ${DataInputs:join('_'):defaultValue('S')} means 'Join together all of the DataInputs separated by underscores, but if that is null, then use the default: the letter S' |
Related Topics
Aliquot Naming Patterns
Aliquot Naming Patterns
To set a naming pattern, open your Sample Type design within LabKey Biologics or Sample Manager. This option cannot be seen or set from the LabKey Server interface.Aliquot Pattern Validation
During pattern creation, aliquot naming patterns will be validated, giving the admin an opportunity to catch any syntax errors. Users will be able to see the pattern and an example name during aliquot creation and when viewing sample type details. Learn more in this topic:withCounter Syntax
Using nested substitution syntax, you can include a counter specific to the value in the AliquotedFrom column (the originating Sample ID). These counters will guarantee unique values, i.e. will skip a count if a sample already exists using that name, giving you a reliable way to create unique and clear aliquot names.Prefix the :withCounter portion with the expression that should be evaluated first, then surround the entire expression with ${ } brackets. Like sample naming patterns, the base pattern may incorporate strings, and other tokens to generate the desired final aliquot name.The following are some examples of typical aliquot naming patterns using withCounter. Learn more in this topic: Sample ID NamingDash Count / Default Pattern
By default, the name of the aliquot will use the name of its parent sample followed by a dash and a counter for that parent’s aliquots.${${AliquotedFrom}-:withCounter}Dot Count
To generate aliquot names with "dot count", such as S1.0, S1.1, an admin can set the Sample Type's aliquot naming pattern to:${${AliquotedFrom}.:withCounter}Start Counter at Value
To have the aliquot counter start at a specific number other than the default 0, such as S1.1001, S1.1002., set the aliquot naming pattern to:${${AliquotedFrom}.:withCounter(1001)}Set Number of Digits
To use a fixed number of digits, use number pattern formatting. For example, to generate S1-001, S1-002, use:${${AliquotedFrom}-:withCounter(1, '000')}Include Lineage Elements
As for Sample IDs, Aliquot Naming Patterns can include elements like names and properties from the AliquotedFrom sample's lineage. Learn more about this syntax here:- Incorporate lineage elements from parent samples or sources.
- Include ancestor elements from any lineage depth using "~".
Examples
| Aliquot Naming Pattern | Generated aliquot names |
|---|---|
| [None provided] | S1-1, S1-2 |
| ${${AliquotedFrom}-:withCounter} | S1-1, S1-2 |
| ${${AliquotedFrom}.:withCounter} | S1.1, S1.2 |
| ${${AliquotedFrom}-:withCounter(1001)} | S1-1001, S1-1002 |
| ${${AliquotedFrom}-:withCounter(1, '000')} | S1-001, S1-002 |
| ${${MaterialInputs/DNA}-:withCounter} | DNA1-1, DNA1-2 |
| ${${MaterialInputs/DNA/propName}-:withCounter} | prop1-1, prop1-2 |
| ${${~DataInputs/Participant}-:withCounter} | Participant1-1, Participant1-2 |
| ${${~MaterialInputs/Blood/Barcode}-:withCounter} | 123456-1, 123456-2 |
Related Topics
ID and Name Settings
- User-defined IDs/Names: Force the use of naming patterns for consistency
- Naming Pattern Elements/Tokens: See and update Sample count tokens
User-defined IDs/Names
To maintain consistent naming, you may want to force usage of naming patterns for samples and sources rather than allow users to enter their own names, risking inconsistencies. This requires that all types have a naming pattern that can be used to generate unique names for them.When users are not permitted to create their own IDs/Names, the ID/Name field will be hidden during creation and update of rows, and when accessing the design of a new or existing Sample Type or Source Type.Additionally:- Attempting to import new data will fail if an ID/Name is encountered.
- Attempting to update existing rows during file import will also fail if an unrecognized or new ID/Name is encountered.
- Select > Application Settings.
- Scroll down to ID/Name Settings.
- Uncheck the box Allow users to create/import their own IDs/Names.
- Note that to complete this change, all entities in the system must have a valid naming pattern. You will see a warning if any need to be added.
Naming Pattern Elements/Tokens
The sampleCount and rootSampleCount tokens are used in Naming Patterns across the application. In this section, you'll see the existing value (based on how many samples and/or aliquots have already been created). To modify one of these counters, enter a value higher than the value shown and click the corresponding Apply New... button.Related Topics
Sample Lineage / Parentage
- Add a Parent
- Add a Source
- Define Lineage When Creating Samples
- Create Aliquots, Derivatives, and Pooled Samples
- Include Ancestor Metadata in Sample Grids
- Explore Sample Lineage Graph
- Explore Sample Lineage Grid
Add a Parent
Within the definition of a Sample Type, you can indicate one or more columns that will generate a parentage relationship with the samples being created. The parent(s) can exist in the same or different Sample Type than the one you are creating. Adding a parent and providing a File Import Column Name (aka a "Parent Alias") in the definition of a Sample Type creates the linkage to parent samples during import. You can opt to make this relationship Required if desired. Note that the column alias/import column name is not actually added to the Sample Type, instead the system pulls data from it to determine parentage relationships.For example, if you have a group of "Vial" samples like the following, where v1 has two 'children', v1.1 and v1.2:| SampleID | MyParent |
|---|---|
| v1 | |
| v1.1 | v1 |
| v1.2 | v1 |
- You can add the parent relationship:
- During initial Sample Type creation.
- After creation by reopening the type for editing via Manage > Edit Sample Type Details.
- Click Add a Parent.
- Parent Type: Select the sample type of the parent. This may be in the "(Current Sample Type)" or another one that has been defined.
- File Import Column Name: Enter the parent column name for import. The column name is case sensitive.
- Check the Required box if you want every sample of this type to require a parent of this type.
- You can add additional parentage columns if applicable by clicking Add a Parent and repeating the process.
- Click Save.
Add a Source
Within the definition of a Sample Type, you can indicate one or more columns that will indicate a Source of the samples being created. Adding a source of a given type in the definition of a Sample Type creates the linkage to sources during import, rather than requiring sources be added later.Define Lineage When Creating Samples
When creating samples, you can include parent/source information based on the Lineage Settings in the sample type, i.e. for any parents or sources included. When you use a template to import from file columns for all parent/source lineage relationships will be included. As shown below, the "MyParent" column contains the parent identifier.Require Lineage Relationships (Optional)
If you want to require that all samples of a given type have a given type of parent or source entity, use the checkbox in the Sample Type definition:Create Aliquots, Derivatives, and Pooled Samples
Once you have created the parent samples in the system and want to create new samples with specific parents (i.e. aliquot, pool, or derive new samples), you can do so from the Samples grid or by importing from a file referencing the parent sample IDs.Create from Samples Grid
Within the application, you can create new samples from any grid of parent samples. Note that if you select more than 1000 rows, the option to create samples with those parents is disabled.- Select the Sample Type of interest from the main menu.
- Select the parent sample(s) using the checkbox(es).
- Select the desired option from the Derive menu. On narrower browsers, this will be a section of the More menu:
- Aliquot Selected: Create aliquot copies from each selected sample.
- Derive from Selected: Create multiple output samples per selected parent sample. Select the type to derive.
- Pool Selected: Created one or more pooled outputs from the selected samples.
- Add the desired number of new samples and click Go to Sample Creation Grid.
Create from Selected Sources
If you have already created the Sources in the system and want to create new samples from specific sources, you can do so from the Sources grid.- Select the Source Type of interest from the main menu.
- Select the desired source(s) using the checkbox(es).
- Select Derive > Samples.
- Choose the Sample Type and enter the desired number of samples in the popup.
- Click Go To Sample Creation Grid.
- You will see the selected Sources prepopulated in the column for that type of source in the grid.
- Enter remaining sample information before clicking to Finish Creating the samples.
Import from File
After creating the parent samples, you can create new derivatives, pooled samples, and aliquots by importing from a file, referencing the parent samples you already created. Obtain the expected import format template, then populate it to indicate the relationships.- Select the Sample Type of interest from the main menu.
- Select Add > Import from File.
- Click Template to download the template of fields.
- Populate the spreadsheet as needed to indicate the intended relationships, then import it to create the new samples.
Include Ancestor Metadata in Sample Grids
You can include fields from ancestor samples and sources in customized grid views. Any user can make their own named custom views, selectively sharing them with the rest of the team, and administrators can customize the default view everyone sees.For example, if you had a "PBMC" sample type and wanted to include in the grid both the "Draw Date" field from the parent "Blood" sample type AND the "Strain" field from a "Mouse" source, you would add the following:| Sample Type: PBMC | |||
|---|---|---|---|
| Parent Type | Parent Field | Label | Position |
| Blood (a Sample Type) | Draw Date | Blood Draw Date | right of Processing Operator |
| Mice (a Source Type) | Strain | Mouse Strain | right of Blood Draw Date |
Explore Sample Lineage Graph
Viewing any sample detail page, you will see a tab for Lineage in the header bar. For example:- Click anywhere in the graph and drag to reposition it in the panel.
- Click any node to see details in the panel to the right.
- Double-click any node to shift focus to that node, which will also adjust to show up to 5 generations from that node.
- This allows you to view the "siblings" of a sample of interest by first double-clicking the common "parent", and then single clicking each sibling node for details.
- Zoom out and in with the and buttons in the lower left.
- Step up/down/left and right within the graph using the arrow buttons.
- Refresh the image using the button. There are two options:
- Reset view and select seed
- Reset view
- On the right of the graph, a side panel lists the children as links.
- Click any sample to reset the graph focus in the left hand panel.
- Hover over a link to reveal direct links to the overview page or lineage view for that child. A tooltip will also show the Sample Type name.
Lineage Generations in Graph
When lineage is complex, different "generations" are shown horizontally aligned.Lineage Grid
The lineage grid can be especially helpful when viewing lengthy lineages or derivation histories. By default, the children of the currently selected sample, aka the "seed", are shown.Troubleshooting
In situations where sample names are only strings of digits, you may see unexpected lineage results if those "number-names" overlap with row numbers of other samples. In such a situation, when there is ambiguity between sample name and row ID, the system will presume that the user intends to use the value as the name.Related Topics
Aliquots, Derivatives, and Sample Pooling
- Aliquots: Split a sample into identical aliquots which retain the properties of the parent sample and may have additional properties.
- You may also split an aliquot into a next generation of "subaliquots" by the same process.
- Derivatives: Derive new samples from one or more existing samples, possibly with new properties or treatments.
- Samples may also be derived from Sources in the system.
- Pooled Samples: Group together existing samples to make a combined pool sample for actions on the aggregate.
- Create New Samples from Selected Parents
- Derive from Selected Samples
- Pool Selected Samples
- Aliquot Selected Samples
- Import Aliquots, Derivatives, and Pooled Samples from File
- Work with Aliquots
Create Samples from Selected Parents
If you have already created the parent samples in the system and want to create new samples from these specific parents (whether by aliquoting, deriving, or pooling), you can do so from the Samples grid. (Note that if you select more than 1000 rows, the option to create samples with those parents is disabled.)- Select the Sample Type of interest from the main menu.
- Select the parent sample(s) using the checkbox(es).
- Select Derive > , then the type of derivation. Note that on narrower browsers, the "Derive" section will be under the More > menu.
- Aliquot Selected
- Derive from Selected
- Pool Selected: Put multiple samples into pooled outputs.
Derive from Selected Samples
When you select Derive from Selected, you'll see the popup with the Derivatives option chosen.- Select the Derivative Type.
- Provide the desired number of Derivatives per parent.
- Click Go to Sample Creation Grid.
- You will see the Samples you had selected prepopulated as parents in the grid, with as many rows per parent as you specified.
- Enter remaining sample information before clicking to Finish Creating the samples.
Pool Selected Samples
To pool a set of samples into pooled outputs, such as for testing on an aggregate of many samples, follow these steps:- Select the Sample Type of interest from the main menu.
- Select the 'parent' samples using the checkboxes.
- If you only select one parent sample, the pooling option will not be shown.
- Select Derive > Pool Selected.
- In the popup, choose the Sample Type for the new pooled samples, enter the desired number of New samples from Pool, and click Go to Sample Creation Grid.
- You will see the Samples you had selected prepopulated as parents in every row of the grid.
- Enter remaining sample information before clicking to Finish Creating the samples.
Aliquot Selected Samples
To create aliquot portions of samples, follow the same initial process. The aliquots will share (inherit) some properties from the parent sample and may have additional aliquot-specific and/or aliquot-editable properties. If you select several samples first, each one will be aliquoted individually, i.e. you will see the desired number of rows per 'parent' with each only having the single parent.- Select the Sample Type of interest from the main menu.
- Select the 'parent' sample(s) using the checkbox(es).
- Select Derive > Aliquot Selected.
- Add the desired number of Aliquots per parent by entering the number and clicking Go to Sample Creation Grid.
- You will see the Samples you had selected prepopulated as Aliquoted From parents in every row of the grid.
- You will also see any field properties that are editable for aliquots ("Provider" and "Color" as shown here).
- You cannot change parent or source types when creating aliquots.
- Each aliquot row will have the AliquotedFrom field set to the selected 'parent' (which may itself have additional parent and source information in the system).
- Enter values in the rest of the columns as needed before clicking to Finish Creating the aliquots.
Aliquot Naming
Like any other sample, aliquots must each have a unique name within the sample type. It is best practice for aliquots to include the name of the 'parent' sample, i.e. the value from the ${AliquotedFrom} column. You can accept the default naming pattern for aliquots or include a custom pattern in the sample type definition.The default aliquot naming pattern is the parent sample name followed by a dash and incrementing counter:${${AliquotedFrom}-:withCounter}${${AliquotedFrom}.:withCounter}Aliquot Information in Grids
When viewing a grid of samples, you can expose various calculated columns related to aliquots. These columns are populated only for the original samples; they are set to null for aliquots themselves.- Aliquot Total Amount: Aggregates the total available amount of all aliquots and subaliquots of this sample.
- Available Aliquot Amount: Aggregates the volume of all aliquots with a status of type "Available".
- Aliquots Created Count: The total number of aliquots and subaliquots of this sample.
- This column is populated only for samples, and set to zero for samples with no aliquots.
- Available Aliquot Count: The total number of aliquots (and subaliquots) that have a status of type "Available".
Import Aliquots, Derivatives, and Pooled Samples from File
After creating the parent samples, you can create new derivatives, pooled samples, and aliquots by importing from a file, referencing the parent samples you already created. Obtain the expected import format template, then populate it to indicate the relationships.- Select the Sample Type of interest from the main menu.
- Select Add > Import from File.
- Click Template to download the template of fields.
- Populate the spreadsheet as needed to indicate the intended relationships. Key details follow.
- Leave the SampleID column blank to have a name created using the aliquot naming pattern.
- Drop the completed spreadsheet into the upload window, then click Import.
Aliquots
Populate the AliquotedFrom column with the Sample ID of the parent Sample. Leave the SampleID column blank to have a name created using the aliquot naming pattern. For example, to create 3 aliquots of sample "S-001":| SampleID | ...Other columns... | AliquotedFrom |
|---|---|---|
| ...Other values... | S-001 | |
| ...Other values... | S-001 | |
| ...Other values... | S-001 |
Derivatives
Populate the columns defined as parent aliases with the Sample IDs of the parent(s). Leave the SampleID column itself blank to have a name created using the sample naming pattern. For example, to create two derivatives of two parent samples, where IDs will be generated and the parent alias column is named "ParentSample":| SampleID | ...Other columns... | ParentSample |
|---|---|---|
| ...Other values... | S-001 | |
| ...Other values... | S-001 | |
| ...Other values... | S-002 | |
| ...Other values... | S-002 |
Pooled Samples
Similar to the above, populate the columns defined as parent aliases with the Sample IDs of the parents. For example, to create two pooled samples from two parent samples:| SampleID | ...Other columns... | ParentSample |
|---|---|---|
| ...Other values... | S-001, S-002 | |
| ...Other values... | S-001, S-002 |
Work with Aliquots
View Aliquots of a Sample
When viewing sample details (click the name of the sample on the grid) you will see a panel showing Aliquots, if any aliquots of this sample exist.- Total Aliquots Created: includes any subaliquots.
- Available Aliquot Count: Availability means that an aliquot is marked with a sample status of the "Available" type. This may include custom statuses created of that type represented by a green tag.
- Available Aliquot Amount: based on the combined amounts for all 'Available' aliquots.
- Jobs with Aliquots: Count will link to a grid filtered to show the set of jobs.
- Assay Data with Aliquots: Count will link to a grid filtered to show the set of runs.
Find Original Sample Details for an Aliquot
When you are viewing sample details for an aliquot it will look like any sample, with Aliquot Data on the Overview panel, as well as Original Sample Data for the sample it was aliquoted from. A link to the details page for that originating sample is included.Aliquots in Lineage
The Lineage tab for a sample that has aliquots will show them as 'children' in the graph. Use the Filter menu to select whether to include Derivatives, Sample Parents, and Aliquots in the display.Related Topics
- Sample Lineage / Parentage
- Manage Samples: Edit lineage for a single sample.
- Edit Selected Samples: Edit parents and sources for multiple selected samples.
- Manage Sample Status
Sample Expiration
- Enable Expiration Date Field
- Assign Expiration Dates to Samples
- Find Expired Samples or Samples About to Expire
Enable Expiration Date Field
When you define or edit a Sample Type, you will see the MaterialExpDate (Expiration Date) column listed with the Default System Fields.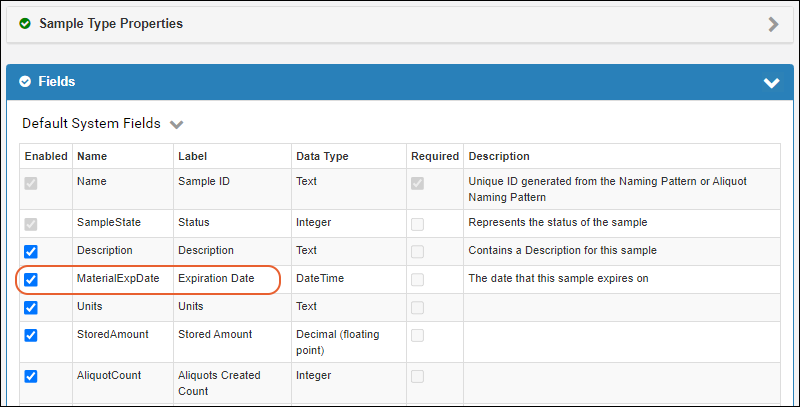 Enable it by checking the box (it is enabled by default).You can also check the box in the Required column to require that every Sample of this type have an expiration date. Note that if you have existing Samples without expiration dates, they must be populated before the field can be required.
Enable it by checking the box (it is enabled by default).You can also check the box in the Required column to require that every Sample of this type have an expiration date. Note that if you have existing Samples without expiration dates, they must be populated before the field can be required.
Disable Expiration Date for a Sample Type
If you uncheck the box for this field, you and your users will not see, be able to set, or use Expiration Dates for this Sample Type. Note that if you use this field at first and it contains data, disabling it will not delete the data. If you later re-enable it, the previous data will be returned to view.Assign Expiration Dates to Samples
When Samples are added or updated, either manually using a grid or via import from file, the Expiration Date can be supplied. In a file import, the column name should be MaterialExpDate.Find Expired Samples or Samples About to Expire
Samples that have already expired are marked with an indicator in the UI. Look for a red triangle in the corner of the grid value or the storage cell location. You'll also see these indicators in the detail panel for a sample.Related Topics
Sample Timeline
- Where is my sample located right now?
- Who has it now and when did they take possession of it?
- What has been done to my sample before I received it?
View Sample Timeline
To see the sample timeline, click the name of the sample in any grid where it is listed. You can find it on the grid for the Sample Type, access it from any assay data uploaded for it, or find it listed for a workflow job.Click the Timeline tab along the top row. You will see all events for that specific sample.Current Status
The Current Status section in the upper right includes summary details like where the sample is now, who was the last to handle it (and when) and active status values.Sample status, storage status, and job status of samples are highlighted with color coded indicators:- Available (green)/Consumed (yellow)/Locked (red)
- Not in storage/in storage
- No job assigned/Assigned to job
Event Details
Click any event on the timeline to see the Event Details in a panel on the lower right. For example, when the event is "Sample was updated," the details section will say when and by whom.Listing Order
By default, the timeline will Show Oldest first. Use the dropdown at the top to switch to Recent first instead.Filter Timeline
When the list of timeline events is long, it can be helpful to filter for events of interest. Click Show filters above the listing to open the filters panel (the link will now read "Hide filters").- Sample Events
- Assay Events
- Job Events
- Storage Events
Export Timeline
Click the (Export) button to download the timeline as an Excel file. The exported file will include:- Date
- Type of event
- Event name
- User
- Old Event Details
- New Event Details
Related Topics
- Audit Log: View all events for all samples.
- Storage Events Timeline for a Sample
- Manage Sample Status
Manage Samples
- View All Sample Types
- View Samples of One Type
- View Individual Sample Details
- Delete Samples
- Move Samples (Premium Feature)
View All Sample Types
To view all the Sample Types defined in LabKey Sample Manager, click Sample Types on the main menu. The Sample Types dashboard lets you:
The Sample Types dashboard lets you:
- Add Samples
- Create Sample Type
- View All Samples of all types in a tabbed grid.
- Manage > Sample Statuses
View Samples of One Type
You can open the grid of all samples of a specific type in several ways:- Click the type name in the Sample Types grid.
- Click the graph bar for that Sample Type on the main dashboard.
- Click the sample name directly from the top level menu from anywhere in the application.
Sample Type Insights
The panel above the grid gives a quick visual summary of Storage Status, Sample Status, and Aliquots. Color coded bars show the relative prevalence of each state. Hover for details about any colored segment. Click any bar segment to filter the samples grid to show only those samples.Manage Menu
The Manage menu in the upper right offers these options:- Edit Sample Type Design: Reopen the existing details and fields; edit using the same interface as you used to create the Sample Type.
- Edit Identifying Fields: Add or edit additional fields to be shown to users selecting these samples.
- Manage Templates (LIMS Feature): Users of LabKey LIMS and Biologics LIMS can customize the import templates offered to users.
- Delete Sample Type: Note this will delete all sample data and dependencies as well. Deletion cannot be undone. The administrator deleting a Sample Type can provide a Reason for Deleting if required or desired, and must confirm the action.
- View Audit History: Administrators can view the audit history for this Sample Type from this link.
View Individual Sample Details
Click the Sample ID for any sample to view the details about it in the system in a series of panels. Tabs along the top offer more details about the sample. From the Manage menu you can create new samples of any type, delete this sample, add it to a picklist, or upload assay data for this sample. Storage editors can add to storage, check it out, or remove it from storage. You can also view the audit history for this specific sample.- Lineage: View a lineage graph or grid.
- Aliquots: See aliquots and subaliquots of this sample.
- Assays: All assay data available for this sample (and any aliquots).
- Jobs: Find all jobs involving this sample (and any aliquots).
- Timeline: See a detailed timeline of all events involving this sample.
Panels on the Overview Tab
Panels on the Overview tab include:- Storage: This panel shows the amount of the sample (if provided). If the Sample is already in storage, it also shows the current location and checkout status, if any. Otherwise, you'll see a link to add the sample to storage
- Aliquots: Details about any aliquots created of this sample. Note that to be "available" in this panel, an aliquot must have a Stored Amount > 0.
- Notebooks: Links to any notebooks that reference this sample will be shown here. Click the name to open the notebook
- Details: Details about this sample.
- For an aliquot, you will see both Aliquot Details and Original Sample Details in separate panels.
- Source Details: Information about sources of this sample. Note that only one generation is shown; see the lineage tab for more.
- Parent Details: Information about parents of this sample. Note that only one generation is shown; see the lineage tab for more.
Edit Sample Details
To edit the details for this sample, click the (Edit) icon for that section. Make changes using dropdowns and selectors similar to when you originally assigned the values.Note that you can edit the SampleID (Name) here, keeping in mind that all Sample IDs must remain unique. This is useful in a situation where the original name may have included a typo or other error. You cannot, however, edit SampleIDs using any bulk method, including 'Edit in Grid' and import from a file.You can provide a Reason for Update if desired or required before clicking Save.Edit Sample Lineage
You can edit the immediate source and parent information (i.e. the first generation of lineage) directly from the sample details page. Use the (Edit) icon for the Source Details or Parent Details section as needed. Make changes using dropdowns and selectors similar to when you originally assigned the values.You can provide a Reason for Update if desired or required before clicking Save.More Sample Editing
Learn more about editing samples in this topic:Delete Samples
Deletion of samples may be necessary for a variety of reasons, and once completed, deletion cannot be undone.Deletion Prevention
For samples with data dependencies or references, deletion is disallowed. This ensures integrity of your data in that the origins of the data will be retained for future reference. Samples cannot be deleted if they:- Are 'parents' of other derived samples or aliquots
- Have assay runs associated with them
- Are included in workflow jobs
- Have a status that prevents deletion
- Are referenced by Electronic Lab Notebooks
Delete One Sample
For samples which can be deleted, you can delete a single sample while viewing it's details page.- From the detail page for the sample to delete, select Manage > Delete Sample.
- In the popup, you can enter the Reason for Deleting if required or desired.
- Confirm the deletion by clicking Yes, Delete.
Delete More Samples
You can also delete one or more samples from the grid view for the Sample Type. Note that you can only delete up to 10,000 samples at one time, so if you need to delete more than that, perform the deletion in batches.- From the grid view for the type of sample to delete, select one or more checkboxes for the sample(s) you wish to delete. To select all samples, use the checkbox at the top of the column.
- Select Edit > Delete.
- In the popup, you can enter Reason for Deleting if required or desired.
- Confirm the deletion by clicking Yes, Delete.
Partial Deletion
If deletion is disallowed for any of the samples you attempt to delete, the popup will give you more details and ask you to confirm or cancel the partial deletion of any samples without dependencies.Move Samples (Premium Feature)
When using the Professional Edition of Sample Manager (or any edition of LabKey Biologics LIMS), users with the appropriate permissions will be able to move eligible samples between Folders.Learn more in this topic: Note that some Sample Statuses, including "Locked", prevent samples from being moved.Related Topics
Manage Sample Status
- Sample Status Types
- Manage Sample Statuses
- Assign Sample Status
- Sample Status Legend
- View Sample Status
- Sample Count by Status Dashboard
Sample Status Types
The Sample Manager application has 3 built-in status types, also represented by assignable statuses. Additional named statuses can be created of any of these types.- Available: Allows any action. This status type is also used to determine calculations of available amounts.
- Consumed: Used up, and no longer can be aliquoted or checked out, but remains in the system for analysis and tracking. Storage updates are prevented.
- Locked: While locked, all actions are prevented except adding the sample to a picklist. A sample cannot be edited, have its storage updated, be moved between folders, or be added to workflows. The status can later be changed to another type of status.
Manage Sample Statuses
From the main menu, click Sample Types then select Manage > Sample Statuses. You can also reach this option using > Application Settings.View or Edit Status Details
Click the name of a status to see details on the right. Here you may edit the Label, Color, Description, and for statuses not currently in use, you can change the Status Type.Add New Status
The three main statuses correspond to the three main "status types" with different actions allowed for each type. Adding additional custom status settings can support your lab's procedures and user expectations. Click Add New Status and populate the fields on the right.Choose Status Color
For both built in and custom statuses, you can customize the color by selecting from the color picker. You may choose to assign the same color to all status of a given type, such as green for all "Available" statuses, or can choose something else to make specific statuses stand out in the system.Assign Sample Status
Users can assign statuses to their samples throughout the system:- At sample creation
- During file import of new or updated samples
- When updating a single sample's details
- In a bulk sample update
Change Status to Consumed
When you change the Sample Status to "Consumed" (i.e. to any named status of the "Consumed" type), this typically means that there is no more of the sample to be used. If the sample is in storage, the user will also be asked whether they also want to remove it from storage.When editing a single sample's details, use the checkbox to also Remove sample from storage?, adding a reason for the update if desired or required:Sample Status Legend
To see the set of available status values, hover over the for the Status column or field to see a legend listing all the available status values and their color coding.View a Sample's Status
Sample status information will be displayed in sample grids and on the sample overview and storage sample details pages. Color-coded blocks and the legend make it easy to see status at a glance. The name of the specific status will be shown, using the color-coding you have assigned.Sample Status and Storage Status
Sample Status can be viewed for a sample in storage, and is not the same as the Storage Status (In Storage, Checked Out, etc.), described further in this section: Storage ManagementSample status values are available in storage views as part of the sample details. Hovering will show the description of the status value, as shown below for a locked sample.Sample Count by Status Dashboard
On the main dashboard, you can see an overall picture of the status of your samples using the Sample Count by Status chart. A bar for each Sample Type is color coded to show how many samples of that type in each status.Related Topics
Edit Selected Samples
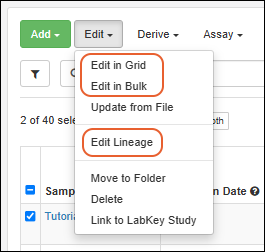 To edit selected samples, select them on a grid or picklist using the checkboxes. If you are viewing a grid that contains samples of several types, you will first need to switch to the tab for the specific sample type before you can edit properties. Once you've selected the samples to edit, use the Edit menu to select one of the options:
You can also bulk edit samples using the Edit > Update from File option.
To edit selected samples, select them on a grid or picklist using the checkboxes. If you are viewing a grid that contains samples of several types, you will first need to switch to the tab for the specific sample type before you can edit properties. Once you've selected the samples to edit, use the Edit menu to select one of the options:
You can also bulk edit samples using the Edit > Update from File option.
Edit in Grid
Learn more about options for editing data in grids in this topic: See the current values for the selected samples, including storage amount and units.- You cannot edit the sample ID or fields like generated Barcode values in the grid.
- If you have selected a mixed set of samples and aliquots, you will also see fields grayed out when they are not editable for both categories. For example, a field that is inherited by aliquots from the originating sample (i.e. only editable for samples) will not be editable here.
Edit in Bulk
Sample details, including amount and units, can be edited in bulk. If any Aliquots are selected when you edit samples in bulk, only the aliquot-editable fields will be shown in the modal and available for bulk editing. To bulk edit sample fields that are not aliquot-editable, be sure to only select non-aliquot samples before selecting this option.In the update panel, click the slider to enable the fields that you want to assign the same value for all the selected samples. Hovering over the will give you more detail about any field, such as for the Status field, you'll see a legend of available statuses. Shown below, the "Status" will be changed but the other fields will be left unchanged.Edit Lineage
Edit the sources and parent samples for the selected samples. Lineage for aliquots cannot be edited.The grid will show the current source and parent settings for the selected samples.- To edit or add a selection to an existing column, click the or start typing, you will see a dropdown of matching options to choose from.
- To add a new parent or source association for these samples, click Add Parent or Add Source, select the desired type for the new association, then select values for the rows in the new column that will be added to the grid.
Edit Sample Name
Note that you cannot edit the SampleID using any grid, bulk or file import method. To change the SampleID, you can edit the individual sample's Details. Keep in mind that sample names must remain unique.Related Topics
Sample Finder
- Video Tutorial
- Open the Sample Finder
- Find Derivatives in Sample Finder
- Sample Finder
- Filtering Criteria
- Choose Values
- Filter With Expressions
- Find Samples with Multiple Common Ancestors
- Edit Criteria
- Search Results
- Saved Searches
Video Tutorial
In this video, you will see how to create persistent, reusable reports using the Sample Finder.Note that updates since the making of this video mean that you select Reports > Find Derivatives in Sample Finder to open this report (instead of clicking "Find Derivatives"). Also, criteria can now include both common and user-defined properties of samples, parents, and sources.Open the Sample Finder
To open the Sample Finder, select it from the search menu anywhere in the application. You can also click Go to Sample Finder from the main dashboard.Find Derivatives in Sample Finder
You can prepopulate the Sample Finder with a starting search filter by selecting a set of samples or sources and selecting Reports > Find Derivatives in Sample Finder. On some grids, this option will be under the More menu. For example, to find all the samples created from any "Mouse" source, select all rows on the Source page for Mice. When you select Find Derivatives in Sample Finder, you'll jump to the sample finder with all sample "children" from your selections where you can add more search criteria.Sample Finder
When you open the Sample Finder without preselecting samples, you'll see the tile dashboard:Filtering Criteria
- Sample Properties: Find Samples based on properties defined in the Sample Type, whether built-in (common to all Sample Types) or user-defined. You can either narrow to a specific type or search across common properties of all Sample Types.
- Ex: Find all Samples with an expiration date in the next 7 days.
- Parent Properties: Find Samples based on the properties of their parent Samples.
- Both built-in and user-defined properties of the parent Sample Type are available for searching here.
- Ex: Find all child Samples derived from any Blood Sample parent with a Draw Date during a certain timeframe.
- Source Properties: Find Samples based on properties of a Source parent:
- Both built-in and user-defined properties of the Source Type are available for searching here.
- Ex: Find Samples taken from male, BALB/c mice.
- Assay Properties (Professional Edition Feature): Find Samples based on Assay Result data.
- Ex: Find Samples with a Platelet count on a CBC assay in specific range of values.
- You an also use a checkbox to find Samples without any results for a selected assay.
Choose Values
For some fields with a limited set of value options, you can use a checkbox interface to select one or more values to include. For example, to choose only samples with a Blood parent where the ParticipantID is either "PT-101" or "PT-102".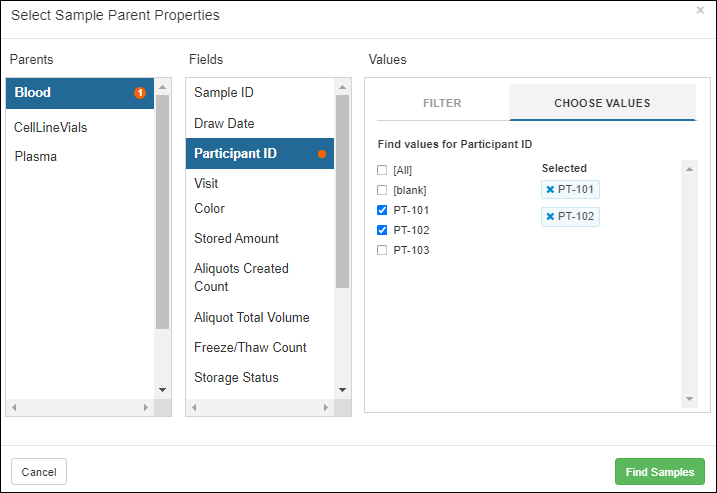
Filter with Expressions
You can also use the Filter tab to specify a filtering expression and enter a value. If desired, you can add a second filter to this column to indicate a range or other combination of expressions that will be AND-ed together.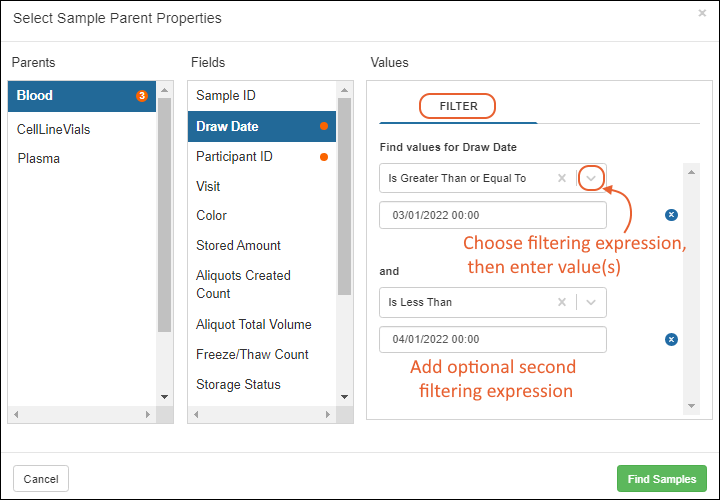 If your first filtering expression fully constrains the result (such as an "Equals" filter) you will not have the option to add a second expression.
If your first filtering expression fully constrains the result (such as an "Equals" filter) you will not have the option to add a second expression.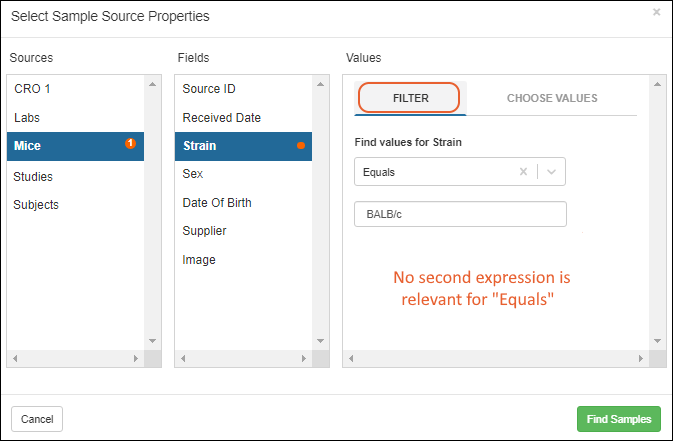
Find Samples with Multiple Common Ancestors
Use the specialty Equals All Of filter operator to find samples that share all of a provided set of multiple ancestors of a given type in common. This option is only available on the Parent Properties and Source Properties cards and can only be used for ID fields, including Sample ID and Source ID. For example, if you wanted to find samples who had both Mouse-1 and Mouse-2 as ancestors, you could use this filter operator. Up to 10 IDs can be provided separated by either new lines or semicolons.Edit Criteria
Once you've defined any sample finding criteria, you can use the buttons in the upper right to add more filters on other columns of any other available type.Each type that you filter on will have a tile, listing the active filters on columns in that type.You can use the for any tile to edit the criteria it will use to find samples. For example, you could add additional filters on other fields of a parent or source filter you already created.To add search criteria for another type of source or parent, start from the Source/Parent Properties button.- Each type will have a separate "tile" showing filters applied. When a filter includes "does not equal", you will see the values with strikethrough styling.
- To set a filter to simply require that the sample have any parent of a given type, use "SampleID is not blank" for that filter.
Search Results
As you build your set of criteria, you will see the results grid below the tiles. There is a tab for All Samples as well as individual tabs for all defined sample types. Each shows the count of samples of that type that were found using your criteria.- Creating new derivatives or aliquots (only available on the tab for a specific sample type)
- Importing assay data
- Adding to picklists or workflow jobs
- Storage Editors can manage storage status, including checking out the found samples
Saved Searches
To save a Sample Finder search, click the Save Search button (only shown when you have generated a set of search criteria).- Most Recent Search:
- The date and time of your most recent search is shown; click to open it.
- Saved Searches:
- You'll see a list of any named saved searches you've created here.
- Note that saved searches are not shared with other users.
- Other Reports
- Built in sample reports are listed here. Learn more in this topic: Built-in Sample Reports
- Manage saved searches
- Save as custom search
Save Search Button
Switching to a previous search result (such as "Searched 2022-06-21 10:27" in the above) will add a Save Search button to the header.Manage Saved Searches
When you click Manage Saved Searches you can edit the name of other searches, or delete them. You cannot edit (or delete) the currently active search; it is shown with a lock icon.Related Topics
Built-in Sample Reports
- All Samples Created by Me
- All Samples Created by Other Users
- All Samples Created in the Last 7 Days
- Samples Expiring in the Next 7 Days
- Samples with Fewer than 5 Aliquots Available
All Samples Created by Me
This report is the equivalent of filtering Sample Properties for All Sample Types where Created By is the current user.All Samples Created by Other Users
This report is the equivalent of filtering Sample Properties for All Sample Types where Created By is NOT the current user.All Samples Created in the Last 7 Days
This report filters All Sample Types where the Created date is greater than or equal to 7 days ago. Notice the use of the syntax "-7d" to mean 7 days ago in this report.Samples Expiring in the Next 7 Days
This report selects Samples where both conditions are true:- They are not already expired, i.e. their Expiration Date has not passed.
- They have an Expiration Date in the next seven days.
Samples with Fewer than 5 Aliquots Available
This report filters the Available Aliquot Count to find those with fewer than 5. In order to be included in this report, the aliquot(s) must have a sample status of the "Available" type.Related Topics
Sample Search
- Sample ID
- Barcode: Barcodes include both:
- Find Samples in Bulk
- Find Samples by Barcode
- Find Samples by ID
- Add More Samples
- Clear Search Results
- Work with Found Samples
Find Samples in Bulk
Click the right edge of the search bar or the icon on narrower browsers and select:- Find Samples by Barcode
- Find Samples by ID
- Barcodes: Search for values in all fields designated as "Barcode Fields" across all Sample Types. This includes:
- All fields of type Unique ID.
- Any text fields designated as Barcode Fields; i.e. those which have the box "Search this field when scanning samples" checked.
- Sample IDs: Search all fields of type Sample.
Add More Samples
Once you have found some Samples, you can click Add More Samples to add to the list. As in the original search, you can input either Barcodes or Sample IDs. This allows you to create a set of results that blend the two search methods.Clear Search Results
To clear the set of search results in the grid, click Reset. This may be a useful option in the case of entering a large number of incorrect samples or other issues.Work with Found Samples
Once you've built your list of Samples in the grid on the search page, you can use the menus for many actions on the All Samples tab. All search result Samples are selected by default.Related Topics
Print Labels with BarTender
Print BarTender Labels for Samples
Before you can print labels, an administrator must have completed the one-time setup steps in this topic, and configured LabKey to print labels. The admin can also specify a folder-specific default label file. When printing, the user can specify a different variant to use.After configuring BarTender, all users will see the options to print labels in the user interface.Print Single Sample Label
Open the Sample Type from the main menu, then open details for a sample by clicking the SampleID.Select Print Labels from the Manage menu.- Number of copies: Default is 1.
- Label template: Select the template to use among those configured by an admin. If the admin has set a default template, it will be preselected here, but you can use the menu or type ahead to search for another.
- Click Yes, Print to send the print request to BarTender.
Print Multiple Sample Labels
From the Sample Type listing, use checkboxes to select the desired samples, then select Print Label from the (Export) menu.- Number of copies: Specify the number of labels you want for each sample. Default is 1.
- Selected samples to print: Review the samples you selected; you can use the Xs to delete one or more of your selections or open the dropdown menu to add more samples to your selection here.
- Label template: Select the template to use among those configured by an admin. You can type ahead to search. The default label template file can be configured by an admin.
- Click Yes, Print to send the print request to BarTender.
Download BarTender Template
To obtain a BarTender template in CSV format, select > Download Template.
Troubleshooting
If you have trouble printing to BarTender from Chrome (or other Chromium-based browser), try again using Firefox.If you are not seeing values populate in your labels, confirm that you have configured BarTender to have the necessary data source connection.Error Reporting
If there is a problem with your configuration or template, you will see a message in the popup interface. You can try again using a different browser, such as Firefox, or contact an administrator to resolve the configuration of BarTender printing.Related Topics
BarTender Integration
- Video Tutorial
- Configure BarTender Integration
- Example BarTender Configuration File(s)
- Configure LabKey to Print BarTender Labels
- Set Up Data Source Connection in BarTender
- Error Reporting
Video Tutorial
Configure BarTender Integration
BarTender integration is supported when used with a BarTender Automation license. We have tested versions 2019, 2021, and 2022.In the BarTender application, you will identify the web service URL and create the label file(s) for printing. The label file has the extension .btw. LabKey applications accept a default label file, but also allow the user to specify a different variant at the time of printing.Include Print Portal
When you install BarTender, be sure to select Specify advanced installation options, then select the BarTender with Print Portal option. To add Print Portal to an existing installation of BarTender, re-open the original installer file, select Modify, and select BarTender with Print Portal.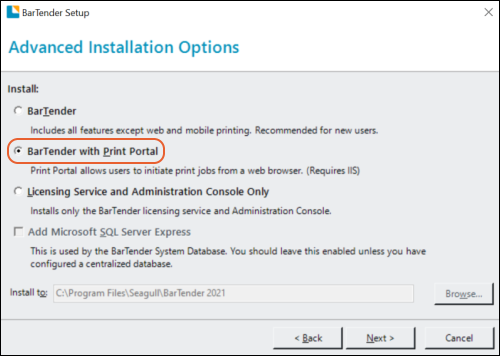 Also check the box to Add Microsoft SQL Server Express if you have not already installed it on your system.
Also check the box to Add Microsoft SQL Server Express if you have not already installed it on your system.
Create and Bind Self-Signed Certificate
The following script will:- Create the certificate with the appropriate flags, including KeyUsageProperty.
- Export the certificate.
- Import the certificate to the local CA so it is trusted by browsers.
# Create Cert
$pwd = 'password'
$cert = New-SelfSignedCertificate -FriendlyName "BarTenderCert" -DnsName $env:COMPUTERNAME -CertStoreLocation "cert:\LocalMachine\My" -KeyUsage DigitalSignature,KeyEncipherment,DataEncipherment -KeyAlgorithm RSA -HashAlgorithm SHA256 -KeyLength 2048 -KeyUsageProperty All -Provider "Microsoft Enhanced RSA and AES Cryptographic Provider" -NotAfter (Get-Date).AddYears(5)
$SSpwd = ConvertTo-SecureString -String $pwd -Force -AsPlainText
Export-PfxCertificate -Cert "Cert:\LocalMachine\My\$($cert.Thumbprint)" -FilePath bartender_cert.pfx -Password $SSpwd
Import-PfxCertificate -FilePath bartender_cert.pfx -CertStoreLocation "cert:\LocalMachine\Root" -Password $SSpwd
echo $cert.Thumbprint
Example BarTender Configuration File
Download this example to help you get started with BarTender 2021 or 2022: For versions prior to BarTender 2021, use this example: Once deployed using the BarTender deployment wizard, the LabKey application would access at the following location (unless you edit the service name):https://localhost/Integration/LabKeyBarTenderService/Execute
https://localhost/BarTender/API/Integration/LabKeyBarTenderService/Execute
Configure LabKey to Print BarTender Labels
Return to the LabKey application (Sample Manager or Biologics LIMS) and select > Application Settings.- BarTender Web Service URL: This is the URL of the web service to use when printing BarTender labels.
- Click Save to save it.
- Once you've saved the URL, you can use Test Connection to test your configuration.
Manage Label Templates
Click Add New Label Template for each template you want your users to be able to use. If you are using multiple Folders, templates can only be defined in the top-level home.- Name: Give the template an identifying display name your users will recognize.
- Description: The description can provide more detail.
- File Path: Provide the path to the label template file to use. The path should be relative to the default folder configured for the BarTender web service specified above.
- Set as Default: Click the selector if you want this template to be the default.
- You are not required to set a default, but if you do, it will be preselected when users print labels.
- For users of multiple Folders, there can be a different default template in each folder.
- Click a template name to see or update details, including whether the template is the default.
- Select and then click Delete to delete a template.
Set Up Data Source Connection in BarTender
For each template and Sample Type, you will need to set up the data source connection in BarTender as follows.- Go to the Sample Type grid and confirm that all the fields you want to include in the label are shown in the grid.
- Select (Export) > Download Template.
- In BarTender, set up the fields in the label to be connected to an External Data Source.
- Make sure to use the template file as the input (it should be a .csv file and includes the field names, rather than field labels).
Error Reporting
If there is a problem with your configuration or template, you will see a message in the popup interface allowing you a chance to verify or change the label template you've selected. If a change needs to be made to the underlying URL configuration, contact an administrator to retry the configuration process.Related Topics
Barcode Fields
- Add UniqueID Field for LabKey-Managed Barcodes
- Generate UniqueID Barcodes
- Use Existing Barcode Values
- Search by Barcode
Add UniqueID Field for LabKey-Managed Barcodes
To support barcodes generated and managed by Sample Manager, you will need a field of type "UniqueID" in your Sample Type. When you create a new Sample Type, you will be prompted to include such a barcode field.To add a "Unique ID" field to an existing sample type, open it for editing. In the Sample Type Properties section, you will see under Barcodes, a blue banner inviting you to create the field necessary. Click Yes, Add Unique ID Field.Generate UniqueID Barcodes
When you add a UniqueID field to a Sample Type that already contains samples, as you finish updating the sample type, you will be prompted to confirm that adding this field will generate barcodes for existing samples.Use Existing Barcode Values
If you have barcodes in your Sample data already, and do not want them generated or managed by LabKey, you can include a field of type "Text" or "Integer" in your Sample Type, and check the Barcode Field box for "Search this field when scanning samples".Search by Barcode
Once your samples have a barcode column of either type, you can search for the value(s) to locate the sample(s). To find by a single barcode, you can search, sort, and filter the samples grid, or use the global Search option in the header throughout the application to find a single barcode value.To more easily find samples by barcode value, you can use the Find Samples option.Related Topics
Sample Picklists
- Create a New Picklist from Samples
- Create Empty Picklist
- Add Samples to Picklist
- Manage Picklists
- View a Picklist
- Refine and Use a Picklist
- Export Picklist
- Shared Picklists and Permissions
Create a New Picklist from Samples
Select the desired samples on a grid and select Picklists > Create a New Picklist. On narrower browsers, this option will be under the More > menu:Create Empty Picklist
You can also create an empty picklist and add samples later. Click Picklists on the main menu, then click Create Picklist. Give it a name, optional description, and check the box if you wish to share it.Add Samples to Picklist
When you select samples in a grid and choose Picklists > Add to Picklist (or More > Add to Picklist), you will see all the Picklists available. If no picklists exist yet, you will be able to click create a new one here.When the number of picklists is long, you can narrow the list by typing in the "Find a picklist" box. Shared picklists are shown on a separate tab, with shared picklists you created yourself shown on the "Your Picklists" tab with a icon.Click to select the desired picklist. You'll see the existing number of samples already on the list.Manage Picklists
Click Picklists on the main menu.- Click Create Picklist to create a new empty one.
- The primary Your Picklists tab shows the picklists you have created. The Sharing column indicates whether you've shared them ("Yes" or "No").
- Click Shared Picklists to see shared ones created by you or other team members.
- Click the Name of a picklist to open it for review or editing. Learn more about using picklists below.
View a Picklist
From the Picklists dashboard, click the name of a picklist to open it in grid form. You will see the samples on the list, available on a series of tabs.Refine and Use a Picklist
Actions available for picklists include:- Manage
- Edit Picklist: Change the name, description, and whether the list is shared with your team.
- Delete Picklist: This removes the picklist, but does not delete any sample data.
- Remove from Picklist
- Use options on the Assay/Jobs/Storage/Reports menus
Export Picklists
To export a picklist, such as to use as a guide for physically removing samples from the freezer, click the icon above the list of samples in the picklist. If any sample rows are selected, only selected rows will be exported. If no rows (or all rows) are selected, the entire picklist will be exported.- CSV
- Excel: For multi-tabbed picklists, you can select which tabs to include in your Excel export.
- TSV
Shared Picklists and Permissions
When the Share this picklist box is checked for a picklist, it will appear for all team members on the Shared Picklists tab of the Picklist dashboard.- Only the creator of a picklist may edit the name, description, and whether the list is shared with the team.
- All users with the Reader role in Sample Manager can read a team picklist and export the data for completing tasks.
- Any user with Editor or Admin permissions may also add or remove samples from team picklists and add picklists to jobs.
- Storage Editors can use picklists for adding and managing freezer storage of samples.
Related Topics
Sources
- Physical Sources like labs, vendors, locations, studies, etc.
- Biological Sources like patients, mice, trees, cell lines, etc.
Topics
Create Sources
Create Source Type
Describe the type of source. Within the system, you might have several different types of source from which samples are taken, such as biological sources like animals or cell lines and physical sources like labs or vendors.From the main menu, select Source Types and click Create Source Type. When you create your first Source Type, there is a shortcut to Create a source type on the main menu. Note that you can only create Source Types in the home (top-level) folder. If the creation button is missing, navigate first to the home.- Enter a Name (Required) for this Source Type, shown here 'Creature'.
- The Name must start with a letter or number character, and avoid special characters and some reserved substrings listed here.
- You can edit the Source Type to change this name later.
- Enter an optional Description.
- Like with samples, all individual sources must have a unique name. You can either:
- Provide these with your data in a column named "SourceId" or
- Ask the system to generate them using a Naming Pattern.
- The default pattern is the word 'Source' followed by an incrementing number:
Source-${genId} - Either accept this default, specify another pattern, or delete it (and ignore the placeholder text) if you plan to supply source names.
- You can change the pattern later, but existing names of sources will not change.
- If you want to include lineage relationships (parent sources) for this type of source, click Add a Parent Source
- As for sample types, click the Fields section to open it.
- Every Source Type includes Default System Fields "SourceId" (Name) and "Description". You can use the checkbox to disable the description, but the name/sourceID is always required. In addition, fields like "Created/CreatedBy", and "Modified/ModifiedBy" are always included but not shown here. Find a list of reserved and internal field names in this topic: Data Import Guidelines
- For defining Custom Fields, you can Import or infer fields from file or manually define fields yourself. Details are found in the topic for creating fields in sample types.
- If you infer from a file that contains the built-in fields, they will not be shown as they will always be created for you.
- Note that you can create fields of the same data types as you can for sample types and assays, with the exception that you do not include fields of type "Sample". Associations of samples with sources are made from the sample definition to it's source, not the other way around.
- Click Finish Creating Source Type when finished.
Source Naming Patterns
When you create a Source Type, you decide how unique names for all the sources will be generated. You can provide them yourself or have them automatically generated with a naming pattern, similar to how samples are named.The default pattern is the word 'Source' followed by an incrementing number:Source-${genId}Add a Parent Source
Sources may have parent sources of the same or other types. For example, you might have source types for both "Studies" and "Subjects", in which all "Subjects" have a parent "Study". Just as how parent aliases can be defined for Sample Types, you can include a Parent Source for your Source Type, giving you a column name to include when uploading parents of the selected source type.Add Sources
Once you have created the Source Type, populate it with the individual sources. This process is very similar to the process of creating samples. You may want to obtain the Template to assist you in formatting your data.- Add Manually to create sources from grid or
- Import from file to import sources from file
Update or Merge Sources from File
Using the Add > Import from File option will only create new sources. If there is data for any existing Source IDs, the import will fail.To update existing sources, or import a spreadsheet merging existing sources and adding new ones, use Edit > Update from File.- Only update existing sources: (Default) Update only. Any new sources will cause the import to fail.
- Create new sources too: Merge. Both existing and new sources can be included.
Related Topics
Associate Samples with Sources
- Associate Samples with Sources During Sample Creation
- Derive from Sources
- View Source Association
- Associate Existing Samples with Sources
- Edit/Remove Source Associations
Associate Samples with Sources During Sample Creation
When you create new samples, you will see an Add Source button in the creation wizard if any source types have been created in the system. If the Sample Type you are adding has any source aliases, source columns for them will be added by default.Derive from Sources
You can also derive, new samples or new sources from selected sources. Start from the grid listing the Sources you want to derive from, then select one or more. Select Derive > Samples or Derive > Sources as desired. (Note that if you select more than 1000 source rows, the option to derive is disabled.)View Source Association
Click the name of a newly created child samples (or source) to open the overview page.Scroll down and notice that the Source Details section has been populated with details about the source(s) you linked when you created it.View Source Lineage
Click the Lineage tab to see that the source is also represented in the lineage grid. Click the node for the source to see if there are other samples derived from that specific source.Associate Existing Samples with Sources
If you create new sources after creating samples, you can add the association from the sample details page. In the next section, see how to edit existing source associations for a single sample.Open a sample type using the main menu, then an individual sample by clicking the Sample ID. You will see the Source Details (if any). Click the icon to edit the source associations for this sample.Edit/Remove Source Associations
Open a sample type using the main menu, then an individual sample by clicking the Sample ID. Use the same icon to edit existing source associations in the Source Details section.- Use dropdowns for Source IDs to add new sources of the types that are already associated.
- Click 'X' to delete individual sources or remove all sources of a given type
- Click Add Source to add new sources of another type.
Related Topics
Manage Sources
- View All Source Types
- Manage a Source Type
- Manage a Single Source
- Edit Sources
- Delete Sources
- Move Sources (Premium Feature)
View all Source Types
The top level menu lists the name of source types that are defined in the system. Click Source Types to see the full listing:Manage a Source Type
Click the name of a Source Type to manage that particular type. You can use the grid view or the top menu to switch which source page you are viewing. Use the Manage menu to select:- Edit Source Type Design: Reopen the source type creation wizard to change details or fields. Note that your changes to the design of the source type will not be 'propagated' to change the name of any existing sources. Users can provide a Reason for Update if desired or required before saving any edits.
- Edit Identifying Fields: Add or edit additional fields to be shown to users selecting sources from dropdowns.
- Delete Source Type: Delete this type of source completely. This will delete the type, all sources of this type, and all dependencies. Deletion cannot be undone. You can enter the Reason for Deleting if required or desired.
- View Audit History: See the history of actions on this source type.
- Add new sources of this type.
- Edit or delete one or more sources, or update from file.
- Derive: Create new samples or sources using any selected source row(s) as parent(s).
- Reports: Select one or more sources, then choose Find Derivatives in Sample Finder to open the Sample Finder showing all the samples created from any selected source(s). Any filters on the grid are also included in the Sample Finder.
Source Type Details
Hover over the Details link to see the naming pattern and any aliases defined for this source type.Manage a Single Source
Click the Source ID to see details for a particular source.- In the top panel you will see all the Samples generated from this specific source.
- Below, a panel lists all the details and offers authorized users an edit link.
- If this Source is referenced in any Notebooks, you'll see links to them here.
- In the Source Parent Details you'll see, and can edit, any lineage this source has, i.e. parent sources of the same or other type(s).
- Tabs along the top row let you click for all the Lineage, Samples, Assays, and Jobs that involve this source.
View Lineage, Samples, Assays, and Jobs
Use the Overview, Lineage, Samples, Assays, and Jobs tabs for a source to see the other information associated with this particular source.Lineage: See the lineage of both parent sources (if any) and derived/child samples associated with this source. Learn more about browsing lineage graphs and grids here.Samples: This tab lists the direct children, grandchildren, and further descendants from this source, including any aliquots. Learn more below.Assays: On the Assay tab, the data you see is what has been uploaded for any samples associated with this source.Jobs: See workflow jobs associated with samples from this source.Samples Created from this Source
On the Samples tab for a given source, you will see all samples (up to a maximum of 5 generations) associated with this source, including any aliquots of those samples. The All Samples tab shows samples of every type, with properties common to any sample, like storage information and status. Samples of each individual type can also be viewed on a separate tab, with the properties specific to that type.- Create Samples:
- Menu sections for Assay/Picklist/Jobs/Storage/Reports:
Edit Sources
Edit Single Source
You can edit the details of a single source by clicking the Source ID and clicking the edit icon in the Details panel.Users can provide a Reason for Update if desired or required before clicking Save.Edit Multiple Sources
You can bulk edit several sources at once by selecting them in the source type view and selecting either:- Edit > Edit in Grid: Edit values directly in a grid. Learn more about the grid editor here.
- Edit > Edit in Bulk: Assign the same value to one or more fields in the batch of selected rows. Learn more about the bulk editor here.
Update from File
Use Edit > Update from file to update source information in bulk. You can select to either:- Only update existing sources (update)
- Create new sources too (merge)
Edit Lineage
You can edit the parents of sources using Edit > Edit Lineage. In the grid, you'll see existing source parents and be able to edit or add more as needed.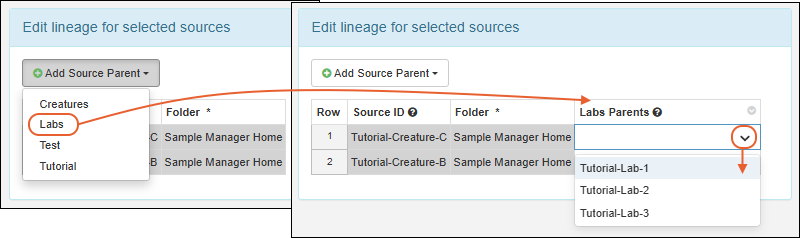
Delete Sources
A Source cannot be deleted if it has descendants, i.e. any samples associated with it (or derived from it). You also cannot delete a source that is referenced from an Electronic Lab Notebook. Deleting a source cannot be undone.Delete a Single Source
You can delete a single source from it's details page by selecting Manage > Delete Source.- In the popup, enter the Reason for Deleting if required or desired.
- Confirm the deletion by clicking Yes, Delete.
Delete Multiple Sources
Multiple sources that do not have descendants can be deleted directly from the source type page by selecting the desired sources and selecting Edit > Delete. Enter the Reason for Deleting if required or desired, then click Yes, Delete to confirm.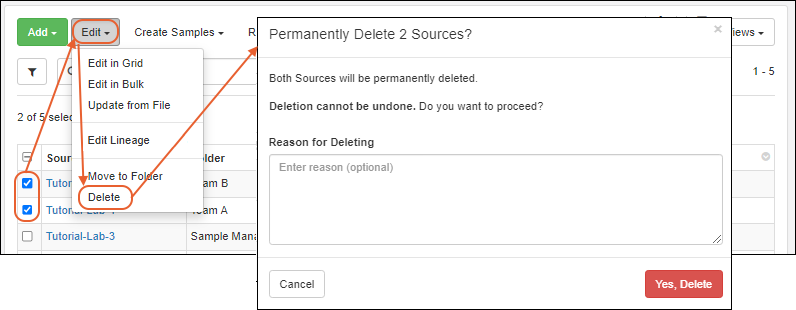 If the group of selected sources includes any that cannot be deleted, you will see a popup message indicating possible reason(s). If some sources can be deleted, you have the option to complete the partial deletion.Deletion of sources cannot be undone.
If the group of selected sources includes any that cannot be deleted, you will see a popup message indicating possible reason(s). If some sources can be deleted, you have the option to complete the partial deletion.Deletion of sources cannot be undone.
Move Sources (Premium Feature)
When using the Professional Edition of Sample Manager (or any edition of LabKey Biologics LIMS), users with the appropriate permissions will be able to move eligible Sources (including Registry Sources) between Folders.Learn more in this topic:Related Topics
Experiment / Assay Data
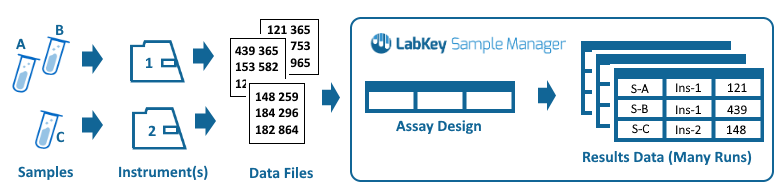 You can create as many different assay designs as you will need to describe the different types of data you will upload. In this section, learn about describing, importing, and updating assay data.
You can create as many different assay designs as you will need to describe the different types of data you will upload. In this section, learn about describing, importing, and updating assay data.
Topics
Describe Assay Data Structure
Create Assay Design
From the main menu, select Assays and then click Create Assay Design. Before any assays have been created, you will see a direct link to "Create an assay design" on the menu. Note that you can only create Assay Designs in the home (top-level) folder. If the creation button is missing, navigate home first.- Give the assay design a Name (Required).
- The name must be unique, must start with a letter or number character, and cannot contain special characters or some reserved substrings listed here.
- Enter a Description to give more information (Optional).
- Active: Check the box for assay designs in active use. When this box is unchecked, the design is archived and hidden in certain views.
- Choose Editing Settings using the checkboxes. Either runs, results, or both can be editable.
- Add the necessary fields and sample mapping described below before clicking Finish Creating Assay Design.
Add Run Fields
Run fields represent information that will be set once per run of data, such as a spreadsheet of individual result rows uploaded together. All result rows in a run will have the same value for any run fields you define.- Click the Run Fields section to open it.
- If you have a specially prepared JSON file of field definitions, you can drop it into the Import fields from file panel.
- Otherwise, click Manually Define Fields.
- Use Add Field to add each run field you need (one is created for you).
- Enter a Name (without spaces)
- Select the Data Type.
- To set more properties of the field, click the to expand it.
Add Results Fields
Results Fields represent the data information in the spreadsheet. You can define results fields in several ways:- Upload a sample spreadsheet to infer all the necessary fields. Either drag and drop a file into the upload area, or click within the same area to select a file directly.
- Import a specially designed JSON file containing field definitions by dropping or selecting it in the same panel.
- Click Manually Define Fields below the panel to define fields in the editor.
- Note that the data itself will not be imported at this time.
- Once fields have been inferred, you can make changes as needed.
- For example, if your results spreadsheet also includes columns for the run fields you defined, you may need to delete the duplicate fields. Click the to delete a field.
- If your result spreadsheet contains any reserved fields, they will not be shown on the inferral list, but they will always be created. You will see a blue banner indicating this reason for not seeing specific fields from your file.
Map to Samples
In order to associate all assay data with the sample it represents, every assay design must included a field which maps to samples in the system. The data type Sample is used to represent that mapping as a lookup into the Sample Type containing the samples.After you infer fields, if one of them is named "SampleID" (such as when you use a naming pattern), it will be mapped automatically. If not, you will see a blue message section asking you to map one of the fields to be the Sample Lookup. The pulldown menu will be populated with the results fields that were inferred. If you need to add a new field to provide the sample linkage, use Add new field. In this example, the SampleID field will be our lookup.Related Topics
Import Assay Data
Import Data
Select the assay design you want from the main menu.To start the import, click Import Data.Enter Run Details
Enter the Run Details requested.- Any fields that are required will be marked with an asterisk.
- The Assay Id field will be the name for this run of data. If you don't enter a name here, an Assay ID will be generated for you. If you upload a data file, the filename will be used. Otherwise, it will be a concatenation of the assay design name and the date and time.
- The Workflow Task field can be used to associate this assay run with a specific workflow task if appropriate.
- Assay Id: "Run1"
- Date: "2019-10-01" (the time "00:00" will autopopulate if you don't select a time)
- Instrument: "INS-01"
Enter Data (Results)
You can enter the result data in one of two ways:- Import Data from File (Default)
- Enter Data into Grid
Import Data from File
You may want to first download a template file of the expected format by clicking Template, then populate it.Drag and drop the file(s) containing your result data into the target area or click the region to select a file.Enter Data into Grid
If neither method described above is appropriate, you can use the Enter Data Into Grid tab to type directly into the entry window. Start by adding the number of rows you want to add and clicking Add row(s).Bulk Insert
You can also use the Bulk Insert button to prefill the grid with many rows of data with some or all values in common. Enter the number of rows to add and provide values that those rows should share. You do not need to enter a value for every column.Bulk Update
Once data has been entered into the grid, either directly or using bulk insert, you can select one or more rows and click Bulk Update to assign new values to all the selected rows for one or more columns. In the popup, use the slider to enable the updating of a row and enter the new value. Values in columns which are disabled in the update will remain unchanged. Any values shared by all the selected rows (such as "Hb" here) will be shown.Delete Rows
If you enter extra rows by mistake, you can select them using the checkboxes and click Delete Rows.Complete Import and Review Data
Click Import when ready to import. If there are any missing or invalid values, you will need to fix them before the import will complete.When your data has been imported, you will see the results for the specific run you just entered. Run details are at the top, results in a grid below.Related Topics
Manage Assay Designs
View All Assay Designs
The assays defined are all listed on the main menu under Assays. To see the list of assays as a grid, click the heading Assays.Manage Assay Design
Click the name of any assay from the main menu or grid to open the runs page for that assay. You will see the assay description, as well as a grid of runs.Edit Assay Design
Click the name of any assay from the main menu or assay grid to open the runs page.- To edit the design select Manage > Edit Assay Design.

- The panels for editing properties and fields in your assay will open.
- You can edit the Name if necessary, ensuring that it is unique. It is best to time such renaming for when there are unlikely to be other users importing data or viewing results, as they may encounter errors and need to refresh their browser to pick up the name change.
- Use the field editor to adjust as needed. Remember that if you delete any fields, all their data will be deleted as well.
- When finished making changes, users can provide a Reason for Update if desired or required before clicking Finish Updating....
Copy Assay Design
To copy the design select Manage > Copy Assay Design. This can be a convenient way to make many similar assay designs or add a new variation without losing the previous design.Export Assay Design
This option is only available when Sample Manager is used with a Premium Edition of LabKey Server.To export the assay design as a XAR file, select Manage > Export Assay Design.Learn more about exported assay designs in the LabKey Server documentationDelete Assay Design
Click the name of any assay from the main menu or assay grid to open the runs page.- To delete the design select Manage > Delete Assay Design.
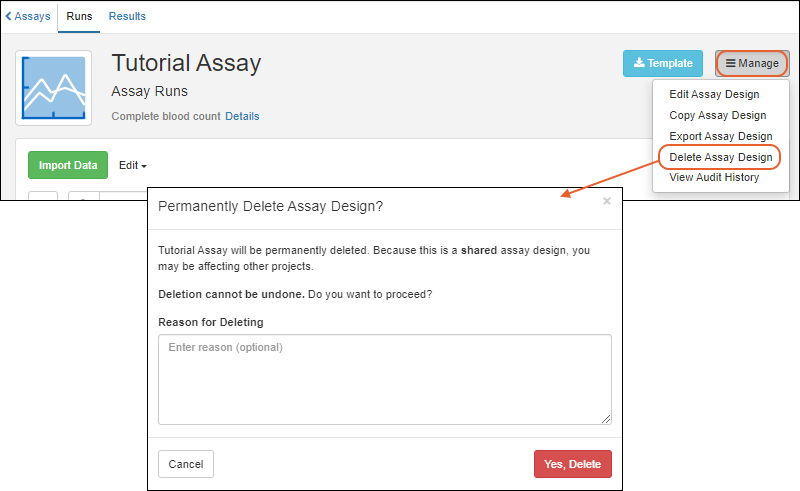
- Note that when you delete a design, all runs of data associated with it will also be deleted. Deletion cannot be undone.
- Enter the Reason for Deleting if required or desired. It will be included in the audit log.
- Confirm in the popup to complete the deletion.
- Active: When this box is unchecked, the design is archived and hidden in certain views.
Archive Assay Design
Assay Designs can be hidden from certain views by unchecking the Active checkbox in the Assay Properties panel. Archived, or inactive, designs are not shown on the main menu or available for new data entry through Sample Manager, but existing data is retained for reference.Using the archive option can be helpful when a design evolves over time. Making the older versions "inactive" will ensure that users only use the latest versions. An assay design may be reactivated at any time by returning to edit the design and checking the Active box again.When viewing all assay data for a sample, both the active and archived assays will be shown if there is any data for that sample.On the main Assays dashboard, you will be able to find inactive assays by switching to the All tab.Related Topics
Manage Assay Data
- Manage Runs
- Manage Results
- Work with Samples from Assay Results
- View Assay Data for a Sample
- View Assay Results for Selected Samples
Manage Runs
First open the assay design of interest, by using the main menu or assay dashboard then then clicking the assay design name. You'll land on the Runs tab by default.Edit Run Properties
If your assay has editable runs and you have sufficient permissions, you can edit run details.To edit run properties for a single run, click the name of the run, then use the (Edit) icon in the Run Details panel to open them for editing. You will see an entry panel you can use to make changes.Bulk Edit Run Properties
Provided your assay has editable runs and you have sufficient permissions, you can also edit run properties in bulk.- Make sure you're on the Runs tab.
- Select one or more runs, then choose either:
Re-Import Run
If you have a change of data or metadata after importing a run, and have editable runs and/or results, you may be able to make the change directly. However, if your runs/results are not editable, you can import a revised version of the run as follows. LabKey Sample Manager will track run re-imports and maintain data integrity.Opening the run details (shown above) and select Manage > Re-Import Run.- Assay Data Re-imported: This event is attached to the "old run" that is being replaced.
- Assay Data Loaded: This event is for the "new run" you import.
Delete Run
To delete a run, either:- Start from the run details page and use Manage > Delete Run
- Start from the Runs tab, select one or more runs and choose Edit > Delete.
- In the popup, enter the Reason for Deleting if required or desired. It will be included in the audit log.
- Confirm the deletion by clicking Yes, Delete.
Manage Results
The result data for your assay is available on the Results tab. Results are individual rows within runs. You cannot add results rows within the user interface. To do so, either import a new run containing the results, or add them to an existing run by reimporting the run after adding the additional rows to the run data file.You may want to include one or more of the Created/CreatedBy/Modified/ModifiedBy fields in the assay result grid view for tracking when and by whom results are edited.Edit Selected Results in Grid
If your assay has editable results, and you have sufficient permissions, you can select one or more rows using checkboxes and select Edit > Edit in Grid.Edit Selected Results in Bulk
If you are editing a number of rows to insert shared values, select the desired rows with checkboxes and select Edit > Edit in Bulk.An editing popup will let you select which field or fields you want to batch update. By default, all fields are disabled; enabling a field using the toggle will let you enter a value to assign for that field in all rows. Shown here, the MCV field will be updated with a shared value, but all other fields left unchanged.- Edit with Grid to switch to updating in a grid format (with the bulk changes you just made already applied). Use this option if you want to make individual as well as bulk row changes.
- Be sure to click Finish Updating # Results when finished with the grid update to save both the bulk changes AND individual changes you made.
- Update Results if no further editing is needed. The bulk updates will be saved.
Delete Results
To delete one or more rows of results within any run, either open the run from the Runs tab or find the desired rows on the Results tab. Use sorting and filtering to help you isolate rows of interest.Check the box(es) for the row(s) you want to delete and select Edit > Delete. In the popup, enter the Reason for Deleting if required or desired. It will be included in the audit log. Confirm the deletion by clicking Yes, Delete.Note that you can only delete 1,000 assay results in one operation. To delete more than that, perform the deletion in batches.Work with Samples from Assay Results
From the page of assay results, you can select a desired set of rows and then use the Samples menu to work with the set of samples mapped to those results.- Jobs:
- Picklists:
- Report:
- Find Derivatives in Sample Finder: Open the Sample Finder filtered to show the selected samples so that you can apply additional filtering.
- View Assay Results for Selected: View a report of results from all assays for this set of samples.
View Assay Data for a Sample
On the details page for any Sample, click the Assays tab to see data about that specific sample. There is an Assay Run Summary tab showing the number of runs per assay type. In addition, a tab for each assay type lets you browse the collected profile of results.View Assay Results for Selected Samples
From a grid of samples, including tabbed grids showing multiple sample types like picklists, you can select desired samples, then choose Reports > View Assay Results for Selected.Related Topics
Storage Management
- Match your digital storage to your physical storage.
- Represent your exact physical system, with shelves, racks, boxes, canes, bags, etc.
- For freezers and other temperature controlled units, you can track storage temperatures and freeze/thaw counts for samples.
- Track overall available storage capacity helping you decide where to store new samples.
- Answer questions about overall volume of samples by type and source; i.e. is there enough blood from patient X to complete the series of assays I want to run.
- Find specific samples and know if they are available.
- Assign samples to storage locations in groups of any size you need, including across multiple locations.
- Move samples as needed and track each location change, creating a full timeline of any sample.
- Control access to physical samples using storage-specific roles.
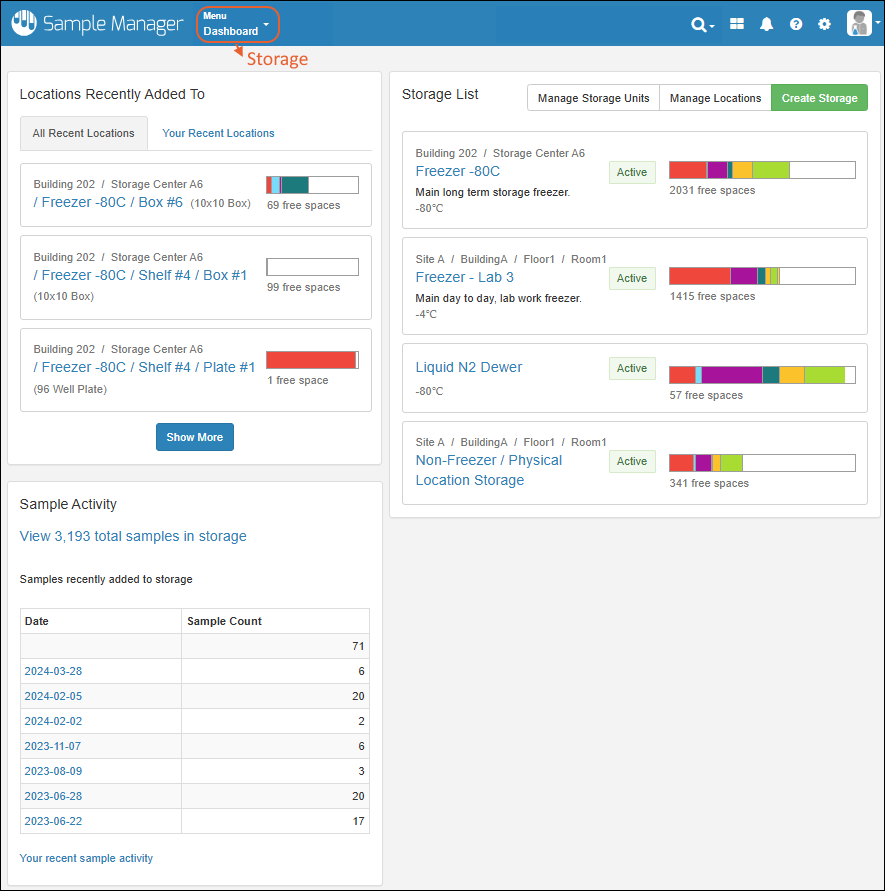
Topics
Create Storage
- Create Storage
- Storage Properties
- Storage Hierarchy
- Bulk Add Storage Units
- Create New Storage Units When Storing Samples
Create Storage
To create a new storage system, select Storage from the main menu, then click Create Storage.Storage Properties
Under Storage Properties define the following:- Storage Type: Use the selector for either:
- Temperature Controlled (Default) or
- Room Temperature
- Name: Any string name that will clearly identify the physical storage; it must be unique in the system. You'll be able to search by this name later.
- Label: Optional text description that will be shown with the name. You'll be able to search by this label later.
- Manufacturer (For temperature-controlled storage only): Optional string for the company name or maker of this unit.
- Freezer Model (For temperature-controlled storage only): Optional string for the manufacturer-assigned model number or name. Note that while the name of this field is "Freezer" model, it could also be an incubator holding samples at a non-frozen temperature.
- Temperature (For temperature-controlled storage only): Optionally include the set-point or consistent operating temperature of this storage system when in use. When giving a temperature, specify Celsius or Fahrenheit.
- Physical Location: If any physical locations have been configured, you can specify here where to find this storage system.
Advanced Storage Settings (For temperature-controlled storage only)
Click Advanced Settings to set the following properties for temperature-controlled storage. Click Apply to save any changes.- Serial Number: Unique identifier for this unit that can be useful for troubleshooting with the manufacturer.
- Sensor Name: If this system is using an alarm or monitor, you can provide the name here.
- Loss Rate: Indicates the time needed or rate at which the unit will return to an ambient temperature from its set-point.
- Status: Used to indicate if the unit is storing samples, available as a backup, or defrosting/cooling/reheating as part of a maintenance routine. Options:
- Active
- Backup
- Defrosting
Storage Hierarchy
Click the Storage Hierarchy section to open it. Add the specific Storage Units available to the panel on the right, either by:- Individually dragging them to the proper location in the hierarchy.
- Using Bulk Add to add many storage units at once.
- Non-Terminal Storage Units : These units can be contained within each other in any combination, but cannot directly contain samples. For example, a shelf can have a rack which has 2 shelves, but there must be a bag, box, or plate on those inner shelves to contain the samples themselves.
- Shelf
- Rack
- Canister
- Terminal Storage Units: These units can directly contain samples and cannot contain other units. Note that these units will only appear in the hierarchy interface when one or more types or sizes have been defined.
- Bag
- Box
- Cane
- Plate
- Tube Rack
- Bulk Add for creation of new storage units.
- Click icon to clone a unit, adding another copy of it at the same hierarchical level (including any nested storage units it contains).
- Bag: Select bag type from the dropdown.
- Box: Select box type from the dropdown.
- Cane: Select cane type from the dropdown.
- Plate: Select plate type from the dropdown.
- Tube Rack: Select the type of tube rack from the dropdown.
Storage Unit Names
By default, all the storage units you add will be named in a sequential numbering scheme, including the type of storage unit in this name. You can edit these names in any way will help your users find the correct locations, such as position/color/identifying labels you have applied or calling racks "partitions" if that is how your users refer to them. Unit names must be unique at any given level in the hierarchy. It is good practice to use unique names throughout the storage system to avoid confusion, particularly where there are many similar structures.The name is indexed making it possible to later search for units by name.Storage Description Labels
You can include a more verbose descriptive Label for every level of your storage hierarchy. When present, labels will be shown in many places to help users better identify the specific storage unit. Both as additional text in hierarchy listings, and as hover text when viewing storage location 'pathways' throughout the application.Clone Units
If you click , you can clone a unit that contains other nested units. The entire structure will be cloned at the same level as the parent. This can speed the process of describing a repetitive storage hierarchy.Finish Describing Hierarchy
Continue to add elements and rearrange them to describe your storage. Here, there are three shelves. One of the shelves has 2 bags, another shelf has two racks, each with 2 boxes of different sizes.- Once you have completed your storage hierarchy, click Finish Creating Storage.
- Note that you can return to edit your storage hierarchy later, such as after defining new storage units.
- If any storage names are duplicates within a given hierarchy, an error will be raised highlighting both of the matching names so that you can edit one or both to be unique. No error is raised for duplicate names in different parts of the hierarchy.
Bulk Add Storage Units
Instead of adding all your storage units individually, you can click Bulk Add to add many at once, streamlining the above drag and drop/copy process. For example, in this example we're adding a shelf that contains 6 racks, each containing 12 plates.Create New Storage Units When Storing Samples
When you are adding samples to storage, you can easily add a new box, bag, cane, plate or tube rack (i.e. any terminal storage unit) from within the Add Samples interface.Expand the existing storage hierarchy to find where you want to place the new unit. Click the , then type the name, select the Unit Type, and optionally include a label.Next Steps
Once you have added at least one storage system that includes some terminal storage locations, you can proceed to store samples.Related Topics
Store Samples
Start from the Storage System
Start from a List or Grid of Samples
- Select Samples to Store
- Find Storage Location for Samples
- Automatic or Manual Location Assignment
- Store Samples in Multiple Locations Simultaneously
Start from the Storage System
Find Available Space
On the home page and the storage dashboard, you will see Locations Recently Added To. Click to open the most recent locations used either by anyone or specifically by you:- Here you'll see the top level of the storage hierarchy you defined. A note of the number of free spaces in each section is provided to assist you.
- Click the to expand the nested hierarchy.
- Each level will summarize space available and in the case of terminal storage, will show the specific storage unit name (i.e. bag size or box dimension).
- As you browse, the system will remember where you were in the hierarchy if you need to go back to a different branch.
- Click any level to see more details in the panel on the right.
- Click Go to Storage View to open it.
- You can also click the location name in the 'breadcrumb' trail along the top of the panel.
Select Space to Fill
There are two categories of terminal storage units to which you can add samples.- 1. Bags and Canes use a simple numbering scheme for positions.
- For bags and canes, you don't need to do anything except click Add Samples to activate the sample panel on the right.
- 2. Boxes, Plates, and Tube Racks have a 2D layout of row and column positions (aka cells, spaces, wells, etc.) within the unit. You have two choices here:
- Click cells or drag within the layout to select a range of positions, as shown below, then click Add Samples. The selected positions will be eligible for placing samples, provided they are not already occupied.
- OR, click Add Samples without making any selection. All available positions will be eligible for placing samples.
Position Already Occupied
If you select a range of cells and one (or more) are already occupied by samples, when you click Add Samples, you will see only the unoccupied positions in the grid.Identify Samples to Store
Edit Location Properties
On the Edit Location Properties tab, you can paste into the grid from a spreadsheet or by typing directly into the grid:Search for Samples
Click the Search for Samples tab to get more assistance with finding specific samples.First, select the desired Sample Type from the dropdown.- Find the samples you want by filtering, sorting and searching.
- Click More Filters to flip to a filters panel, described below.
More Filters
Click More Filters on the Search for Samples tab to select on or more of the filter options:- Created By: Find samples added by a specific user.
- From/To: Find samples created in a specified date range.
- Click Show Results to see the samples that match your filters.
- Use checkboxes to select the samples of interest, then click Assign Positions, or return for More Filters.
- Once some filters are applied, you'll be able to click Clear all filters to clear them.
- Use Back to Grid to return to the previous set of selected samples.
Assign Positions
Once you've filtered to find the samples you want, use checkboxes to select the ones you will be adding to this storage unit.- Note: If you select more samples than there are spaces available, you will see an error message telling you to reduce the number of selected samples before proceeding.
- Position:
- In the case of a bag, the numbered position is not meaningful.
- For a cane, it could be numbered from top to bottom (or bottom to top) as your convention dictates.
- For boxes, plates, and tube racks, the "Position" column corresponds to the layout on the left, which can use either Row/Column names or a numbering system. Learn about assigning specific positions below.
- Sample ID: the selected samples are listed, but you can add more rows directly. Sample IDs must already exist in the system.
Choose Fill Order, Starting Location, and Fill Direction
When you start from the storage and search for samples to add to a structured storage unit like a box, after clicking Assign Positions, you'll be able to select the order, starting point, and fill direction.- Fill Order:
- Manual (Default)
- Sample Creation Order
- Sample ID Ascending
- Sample ID Descending
- Starting Location:
- If you preselected cells, the first available spot in your selection is used and this dropdown is not offered.
- If not, then the first available spot in a storage unit is selected by default. You can specify another starting location if desired using the dropdown.
- Fill Direction:
- Left to Right
- Top to Bottom
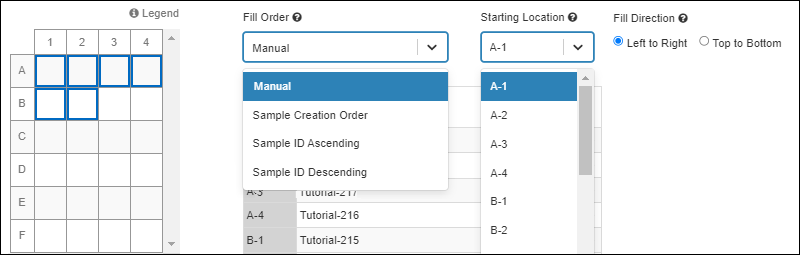
Add to Storage
Click Add to [Storage Type] to complete the position assignments.After assignment is complete, you will see the Storage View for the location.- The newly added samples will be selected in the grid.
- In the case of a box, plate, or tube rack, the cells containing the newly placed samples will also be highlighted.
Start from a List or Grid of Samples
Instead of starting from the Storage, as described above, you can add Samples to storage starting from the sample grid, either while viewing the Sample Type as a whole, or starting from a curated picklist or Sample Finder search results.Select Samples to Store
Open the grid of Samples of interest and identify the Samples you want to store. Only Samples with a Storage Status of "Not in storage" can be added, so applying this as a filter can be helpful.Use checkboxes to select the Samples, then select Storage > Add to storage above the grid. On narrower browsers, this option will be on the More > menu.Find Storage Location(s) for Samples
In the Assign Samples to Storage popup, you will see storage systems with available storage space. The application will remember where samples of this type were last added and open to that location first. You can then navigate to other locations as needed.The number of spaces available at each level will help you see easily where there is room for your new samples. To quickly find particular storage you want, use the search box at the top of the panel to find storage units by name or label.Use the icons to expand the hierarchy of the storage of your choice, seeing available spaces at each level. You'll see unit names, as well as the type/size of terminal storage units to assist you.When you find the desired terminal storage unit, click Select. If necessary, you can also add new terminal storage units by clicking the for the level where you want them.For boxes and plates, you'll see a preview of the current contents of the storage on the right.Automatic or Manual Location Assignment
Once you have selected a storage location with enough space for all selected samples, click Select Positions. Starting from a list or grid of samples gives you somewhat different options than if you had started from the storage system to store them. By default, the samples will be placed in the first available layout positions, using left to right, then top to bottom sequencing. Select either:- Automatically Fill: Customize the fill order, starting point and fill direction.
- Manually Fill: Place them manually, with the option to change the fill direction to adjust how the rows are listed in the grid.
- Populate the grid with storage properties.
- Assign positions, make any position adjustments needed.
- Click Add to [Storage Type] to complete the position assignments.
Automatic Fill Options: Fill Order, Starting Location, Fill Direction
- Fill Order:
- Samples Grid Order (Default): This option reads "Manual" if you did not start from a sample grid.
- Sample Creation Order
- Sample ID Ascending
- Sample ID Descending
- Starting Location:
- By default, the first available spot in a storage unit is selected. You can specify another starting location if desired using the dropdown.
Manual Fill Options: Fill Direction
When you choose the manual fill option, you can select the fill direction which will reorder the rows in the grid for manual filling.Store Samples in Multiple Locations Simultaneously
When you're storing more samples than can fit in one terminal storage unit, you'll be able to select multiple storage locations in the "Add Samples to Storage" modal. For example, if you have a rack of many 10x10 boxes and want to continue filling them in sequence with arbitrary numbers of samples to store in each batch, you might have a situation where there are 6 spaces left in a box. Rather than having to first pick 6 samples for that box, then the remaining samples for another box, you can select all your samples and choose multiple locations. You can even choose storage locations across different storage systems if desired.Samples are added to storage in the order they appear in the grid.As you find storage locations for your selected samples, clicking Select, you'll see a tally of the number of samples you still need to find spaces for before you can click Select Positions. Each time you choose a storage unit by clicking Select for it, that link will change to Clear in case you change your mind. In this example, we have 9, and chose a box with 6 spaces, so need 3 more from another box, but you could have hundreds and fill as many boxes as needed.Related Topics
Manage Storage
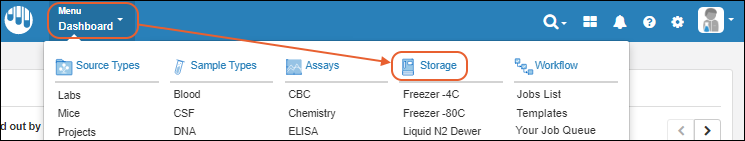 From this Storage Dashboard, you can see an overview and manage your sample storage.
From this Storage Dashboard, you can see an overview and manage your sample storage.
- Locations Recently Added To: The locations Samples were added to most recently.
- Storage List: Recent storage is listed above all other storage.
- Sample Activity
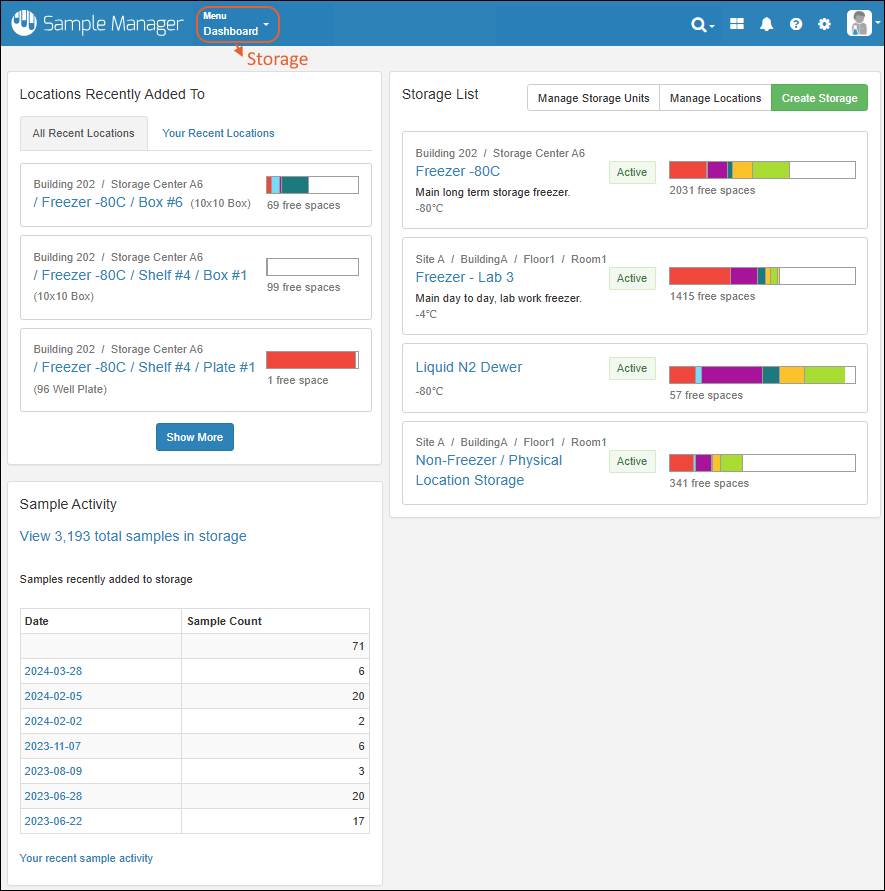
Locations Recently Added To
The panel titled Locations Recently Added To lists the storage units where samples were most recently added. There are two tabs:- The "All Recent Locations" tab is the default, showing storage locations that have had samples added to them most recently.
- You can also choose "Your Recent Locations" to see where you last added samples, giving you a quick way to jump back to where you were.
Storage List
The top of the Storage List shows Recent Storage, the most recently accessed storage systems for each individual user. Below that, All Other Storage is listed. Note that only 10 freezers are shown in this list initially; click Show More to see the rest, also 10 at a time.There are two action buttons for administrators:- Manage Storage Units: Define the "terminal" storage units in which samples are stored. Boxes, bags, canes, plates, and tube racks can be of different sizes, storage layouts, and labelling conventions.
- Create Storage: Add another storage system.
- On the left:
- Name: Click the name to open the details for this storage.
- Above the name, you'll see this storage system's location, if any has been defined.
- If set, the Status will be displayed in a colored block.
- Below the name, the label (description) may also give more identifying information to users.
- Configured set-point temperature (for temperature-controlled storage).
- On the right:
- Color coded bars represent the number of samples currently stored in that storage, scaled to give a visual indication of overall capacity free.
- The number of free spaces is below the bars.
- Click anywhere in the tile to go to that storage overview page.
Sample Activity
- View ### total samples in storage: open a filterable grid of all stored samples. Learn more here: View Storage Activity
- Click a Date to open the grid of samples added to storage by any user on that date.
Related Topics
Manage Storage Unit Types
Bags
Bags are a flexible way to store samples. Default bag sizes created in the system can hold 10, 100, 200, or 500 samples.If you want to add an additional size bag, select the Bags tab and click Add a New Bag. There is no internal limit to the number of samples a bag can hold.Boxes
Boxes can be of different sizes and layouts, accommodating many types of sample storage. Click the Boxes tab to manage box types. Some common sizes are built in to the system:- 10x10
- 10x5 (10 columns, 5 rows)
- 5x5
- 9x9
- You can add a Storage unit description to assist your users.
- Under Display Options you will see a Position display preview, i.e. how the spaces in your box will be labeled, shown here "A-2".
- Position labels: Choose either Columns and Rows or Numbered Position, which will use a standard left to right, top to bottom sequential numbering system starting with 1. For Columns and Rows make these further selections:
- Column labels: alphabetic or numeric. The default is numeric.
- Row labels: alphabetic or numeric. The default is alphabetic. If alphabetic is chosen and the number of rows is greater than 26, the rows after Z will be labeled AA, AB, AC, AD, etc.
- Label Order: Choose whether to show rows or columns first. The default is to show rows first.
Canes
Canes hold a number of samples typically in a vertical alignment. Click the Canes tab to see and add cane sizes. Built-in cane types are named for the sample capacity in the cane:- 4 Cane
- 5 Cane
- 6 Cane
- You can add a Storage unit description to assist your users.
Plates
Plates can be of different sizes and layouts, accommodating many types of instrument. Click the Plates tab to see and add plate types. Built-in plate types are:- 24 Well Plate (6x4)
- 384 Well Plate (24x16)
- 48 Well Plate (8x6)
- 96 Well Plate (12x8)
- You can add a Storage unit description to assist your users.
- Under Display Options you will see a Position display preview, i.e. how the spaces on your plate will be labeled, shown here "A-2".
- Position labels: Choose either Columns and Rows or Numbered Position, which will use a standard left to right, top to bottom sequential numbering system starting with 1. For Columns and Rows make these further selections:
- Column labels: alphabetic or numeric. The default is numeric.
- Row labels: alphabetic or numeric. The default is alphabetic. If alphabetic is chosen and the number of rows is greater than 26, the rows after Z will be labeled AA, AB, AC, AD, etc.
- Label Order: Choose whether to show rows or columns first. The default is to show rows first.
Tube Racks
Tube racks can vary greatly in size and layout. Click the Tube Racks tab to see and add the types you need. Built-in types are:- 4x5 Tube Rack
- 4x6 Tube Rack
- 6x12 Tube Rack
- 6x6 Tube Rack
- You can add a Storage unit description to assist your users.
- Under Display Options you will see a Position display preview, i.e. how the spaces in the rack will be labeled, shown here "A-2".
- Position labels: Choose either Columns and Rows or Numbered Position, which will use a standard left to right, top to bottom sequential numbering system starting with 1. For Columns and Rows make these further selections:
- Column labels: alphabetic or numeric. The default is numeric.
- Row labels: alphabetic or numeric. The default is alphabetic. If alphabetic is chosen and the number of rows is greater than 26, the rows after Z will be labeled AA, AB, AC, AD, etc.
- Label Order: Choose whether to show rows or columns first. The default is to show rows first.
Related Topics
View Storage Details
- Overview Tab
- Storage View
- Sample Information in Grids
- Select Storage to View
- Box, Plate, Tube Rack Detail Views
- View Selected Samples in Box, Plate, or Tube Rack
- Sample Actions
- Location History
- Storage Description Labels
- Manage Menu:
Overview Tab
The overview page for a storage system shows a high level view of storage status and contents.Storage Status
Storage Details
Hover over the Details link to see the properties, including the temperature (if set), usage of this storage, and any advanced settings defined for this storage.Storage Contents
Below the header, you will see a split panel where you can browse the storage hierarchy and contents.On the left, expandable sections for the top level storage units let you 'browse' the structure of this storage.- Click the to expand a section. (It will become a that you can click to collapse it again.)
- You'll see the terminal storage unit type displayed, typically the size of the unit.
- Click a section on the left to see the details for it on the right.
- A 'breadcrumb' trail along the top identifies the location of the unit being shown. Above we see a box on a rack in a shelf in a freezer.
- You can click any level of this breadcrumb to open the details for that unit on the on the Storage View tab.
- You can copy this "path" including the slash separators, and paste it to make it easier to build spreadsheets for import or other use.
- A button to Go to Storage View is displayed. Click to open this unit on the Storage View tab.
- Visual bars follow, summarizing the contents of this storage unit. They are colored using the label colors assigned to Sample Types.
- Capacity: The shaded section indicates used spaces. The number and percent available are listed.
- Sample Types: The breakdown of types of samples in this unit is shown visually using the label colors assigned. Hover over the bar to see which colors represent which types of sample.
- Samples Checked Out: The number checked out is shown, under a bar indicating the type that are checked out.
Storage View Tab
The Storage View presents a more detailed look into the contents of this storage. Below the Storage Status and Storage Details panels, the storage section lets you browse through the storage hierarchy.- Select which level of storage to view.
- Choose whether to view all samples, or only selected samples in the grid.
- Switch to a tab for a specific sample type of interest. Here you can perform the same sample menu actions as on other sample grids.
- Check the box for a specific sample to see its details on the right.
Sample Information in Grids
All storage units, from the overall storage system to a shelf to a plate or bag, show information about Samples in a grid in the Storage View, as shown above. Scroll to the right to see additional columns.Select whether to view All Samples or Selected Samples in the grid. You can also use the tabs to select whether to view samples of all types ("All Samples" or samples of a chosen type; numbers on the tabs indicate how many of each type are available at this storage level. As with any grid, you can then sort and filter them or search to find a subset of interest.The terminal storage units with assigned locations, i.e. boxes, plates, and tube racks, also include a layout interface with the sample grid below it.Select Storage to View
The top row of the panel shows what location you are viewing and lets you select into the nested levels of the storage hierarchy.- Use the to open a menu of locations available.
- Select one to view it. And if desired, go further by using the to select from the next level 'into' the storage.
- When all of the locations are of the same type, such as when Shelf #1 only contains Boxes below, the menu will have the label of the type; when storage units are mixed, "Location" will be shown.
Box, Plate, Tube Rack Detail Views
When looking at the Storage View for a Box, Plate, or Tube Rack, you will see a layout of rows and columns matching the definition of the storage unit, and color coded indications of the specifics of each cell, or place, in that layout.- Rows and columns are labeled alphabetically or numerically, depending on how you defined the storage unit.
- The 'occupied' positions will show a color-coded dot.
- Shading indicates positions that are reserved for samples that are currently checked out.
- Hover over the Legend to see it.
- Hover over any position in the grid to see a popup with details about it.
View Selected Samples in Box, Plate, or Tube Rack
- Click any position (cell) to select the sample in it and see details, as well as actions, on the right.
- The clicked/selected position is shaded blue, and if it contains a sample, the corresponding row is also selected in the data grid below the graphical layout.
- If you select a position where a sample is checked out, shaded pink, you will see details about the sample that "belongs" in that position but is currently checked out, including a link to View sample timeline where you can learn more about who checked it out, when, and why.
- If you are a storage editor or administrator, you will also see additional action buttons here.
Sample Actions
- Learn about adding samples to storage in this topic: Store Samples
- Learn more about checking samples in and out in this topic: Check Out and Check In
- Learn about moving samples and removing them from storage in this topic: Move Stored Samples
- Learn how to edit storage details in this topic: Manage Stored Samples
Location History
When in the Storage View, if no specific position is selected, use the Location History tab to see a timeline of events for the storage unit you are viewing. For example, viewing a box after it has moved, you would see two events: the original creation, and later movement to elsewhere in the hierarchy.Storage Description Labels
It may be useful to provide descriptions of individual storage units for your users. You can customize the name of the unit itself, or from within the Storage View, you can add a more verbose label or edit an existing one. For example, you could include more text from box labels here, or names of investigators or groups.Below the name of the unit, click the to edit. Type the new label (or edit the existing label). Hit enter to save.Manage Menu
Use the Manage menu for the following actions:Copy Storage Definition
Select Copy Storage Definition from the Manage menu of the storage you want to duplicate. Use this option to create a new storage system with the same initial properties and hierarchy. No storage contents will be copied.You can use this as a shortcut to creating many similar storage systems, or adding a new one as you need additional storage capacity. Once copied, customize the new storage definition. At a minimum, each storage system must have a unique name. The default name of copied storage appends "(Copy)" to the end of the original name.Delete Storage
Deletion of a storage system is permanent and cannot be undone.- The storage definition and all of its storage units will be permanently deleted.
- If any samples are stored in it, you must have an "Administrator" role to delete the storage.
- You will see a count of how many are stored. The samples themselves (the sample data) will not be deleted if you proceed, but all of the location information for those samples will be cleared.
- Non-administrators must remove all the samples from storage before they can delete a storage system.
Related Topics
Edit Storage Definition
Open Storage Editing
You can edit a storage definition by first opening the desired storage from the main menu or storage dashboard, then selecting Manage > Edit Storage Definition.Edit Storage Properties
The Storage Properties panel can be edited using the same interface as when it was originally created.Edit Storage Hierarchy
Open the storage definition for editing, then click Storage Hierarchy. The interface for editing the hierarchy is the same as when you created it. Drag and drop units to where they should be. Use to expand sections in the hierarchy.When storage is empty, i.e. before any Samples have been stored in it, you have more flexibility in changing the layout and storage location contents.Once some samples are stored, you will see a icon marking storage units that cannot be deleted. You can still add additional storage units to "locked" non-terminal storage. For example, you can add more boxes to a rack that shows as "locked" because it already contains boxes containing samples, but you cannot delete the rack entirely.Add New Storage Units
Drag and drop new storage units to the positions in the hierarchy where you want them. Adding additional storage will change the capacity and usage percentages for the storage.Adjust Existing Storage Units
Edit the Name given to storage units to help your users identify the specific locations and containers they need. You can also add a descriptive Label for each unit. Both the name and label of the box are indexed making it easier to find a desired storage unit later.Drag existing storage units to different parts of the hierarchy to move them within the current storage system. You can rearrange storage units even if they already contain stored samples.Changing the position or name of a terminal storage unit will change the current information shown for all stored samples, but does not register as a timeline event for samples provided it stays in the same storage system.Learn about moving a storage unit to a different storage system, which will be tracked as a timeline event for any samples it contains, in this topic: Move Stored Samples.Remove Storage Unit
When removing a storage unit from a storage system, first check to ensure that the physical samples stored there have been moved to new locations in the system, otherwise you will lose the tracking data associated with the first location.Learn more about moving samples in this topic: Move Stored Samples.Change Storage Unit Type
If you find you need to change the type of a storage unit, either because the wrong initial type was created or because your labelling system has been changed, first create the new/corrected unit "parallel" to the one you are replacing.- If you are changing a non-terminal storage unit (such as replacing a shelf with a rack) you can drag the contents of the old unit into the new one.
- If you are changing a terminal storage unit type, such as a box, you must specifically move the samples from the old to the new unit.
Related Topics
Manage Storage Locations
- This is distinct from the storage unit hierarchy within the storage.
- In the past, LabKey may have set these values up for you, so you will only need to follow these steps to update or add new locations.
Configure Location Options
Identify Hierarchy
To get started, identify the hierarchy of locations you need. For example, you might have two buildings with various possible storage locations in one and a single possible location in another. For example:myInstitution
│
├───Building 101
│ ├───2nd Floor
│ │ ├───Room 2A
│ │ └───Room 2B
│ └───3rd Floor
│ ├───Lab - Rm 42
│ └───Storage Center
│
└───Building 202
└───Storage Center
Manage Locations
Select Storage from the main menu, then on the Storage List, click Manage Locations.Set Physical Location
Once you have set up your Physical Locations, you can either place existing storage systems into their locations, as shown above, or edit the definitions to specify a physical location for each storage system.From the storage dashboard, click the name of the storage, then select Manage > Edit Storage Definition. Under Physical Location, click Select Location.- You could also click Manage Locations here to add or change the location hierarchy. After making changes, you'll return to editing the storage definition.
- Click to select the desired location, then click Apply.
Use Physical Locations
Unlike any location information you might place in the description field, these locations will be shown on the storage dashboard and in the selection popups when deciding where to store or move samples.Truncated Display of Long Location Paths
When a path is long, it will be truncated for easier display. The first and last locations will be shown with an ellipsis … representing all levels between them. Hovering over the truncated location will show the full location path.Related Topics
Move Stored Samples
- Move One Sample
- Move One or More Samples from Grid
- Move Samples From Storage View
- Select New Storage Location
- Move Samples to or from Multiple Locations
- Move a Storage Unit
- Move Within Current Storage System: Edit Definition Method
- Change a Storage Unit Type
Move One Sample
To move a single sample, you can select Manage > Move in Storage from the details page.Move One or More Samples from Grid
Select the sample(s) from any sample grid and select Storage > Move in Storage. On narrower browsers, this option will be under the More > menu.Move Samples From Storage View
Navigate to the Storage View of the place where the sample(s) are currently stored. If you are starting from the Sample details page, click the location box in the Storage Location panel.Select New Storage Location
The popup for moving samples uses the same interface as originally adding samples to storage. In the first step, you'll select the new storage location. Use the search bar to find locations by name or label.Move Samples to or from Multiple Locations
When selecting new storage locations for a selected group of samples, the same interface applies to adding samples to storage originally, in which if you select a new location that doesn't have room for all the samples you are moving, you'll be prompted to continue selecting additional locations for the remaining samples. Learn more in this topic:Move a Storage Unit
A storage unit can be moved to a new location either in the same storage system or in a different one by clicking Move Storage Unit. When any storage unit is moved, all the samples, as well as any other storage units contained 'within it' are also moved. When using multiple folders, moving storage units is only available to users with the Storage Editor role in the top-level home, i.e. the level where the storage system is defined.View the unit you want to move, either in the storage overview, or the storage view for that unit, then click Move Storage Unit.- If you move a storage unit containing samples, any samples it contains will stay with the same position assignment(s) within the storage unit.
- See below for an alternative method for moving within a storage system.
Move Within Current Storage: Edit Definition Method
If you are moving a storage unit within a storage system, you can also edit the definition to relocate that box to its new location, as described here:Open the storage details page by clicking the name on the main menu.- Select Manage > Edit Storage Definition.
- Click the Storage Hierarchy section.
- Use icons to expand sections.
- Lock icons are shown on any storage units containing samples, indicating that you cannot change the size or type of the units, but you can move them within the storage here.
- Locate the storage unit you want to move. Shown below, Box#1 - Green" used to be on Shelf #2/Rack #1 and is being moved to Shelf #1.
- Click Finish Updating Storage.
Change a Storage Unit Type
In order to change the type of a storage unit, such as to change the size or layout of a box, you can edit the storage hierarchy prior to storing any samples in the unit. Once samples are stored, however, you cannot edit the unit type, but instead need to accomplish this change as a sample move. First create the new storage unit, then move the samples to it. In the case of moving samples from a smaller to a larger box, you could choose to maintain the row and column layout positions, but this is not required. The new unit can be in a different storage location or could be in the same location as the previous unit.Note that while this operation is recorded as a move for the samples, there is no "location history" connection between the original box and the different sized box.Related Topics
Manage Stored Samples
- Video: Interact with Samples in Storage
- Create Samples from Stored Samples
- Create New Picklists and Jobs with Stored Samples
- Add Stored Samples to Existing Picklists and Jobs
- Edit Storage Details
- Remove Samples from Storage
- Export Storage Map
Video Walkthrough
Watch the process of working with stored samples, including checking them in and out and removing them to release the storage locations for other samples. Note that some interactions have changed since the making of this video.Create Samples from Stored Samples
When viewing samples in storage, you can create new derivatives, pooled samples, and aliquots directly. Use the grid view and select the Sample Type of the parent samples from which you want to create new ones. Choose the desired action from the Derive menu (under More > on some browser widths).Add Stored Samples to Picklists and Jobs
To simplify storage workflows, you can also select samples in storage views and create new picklists and workflow jobs including them, or add them to existing picklists and jobs. You can use the tabs for the specific Sample Type, where you'll use the usual sample grid menus. Or from the "All Samples" tab or a storage grid view, select samples and choose the desired action from the Picklists or Jobs menu. Note that on narrower browsers, these buttons will be combined into a More menu.Edit Storage Details
When a sample is initially added to storage, the initial freeze/thaw count is set to zero (for temperature-controlled storage). The storage amount can be provided when the sample is first created or later. If you need to edit these settings later, you can do so by clicking the sample, and clicking either icon in the Storage Details section to open an edit panel.Remove Samples from Storage
The action of removing a sample from storage does not result in removal of the sample information from the system, only that it is taken from an "In Storage" state to a "Not in Storage" state and the freeze/thaw count and storage location are dropped. This could be related to the consumption of all of the sample, and thus it's no longer being available, or could be temporary and the sample could be returned to storage later.If instead you want to delete all data related to a Sample, follow the instructions here: Delete Samples. Removing samples from storage will:- Remove them from your storage location views.
- No longer retain their freeze/thaw counts (or allow them to be updated).
- Prevent any further check outs or check ins.
- NOT delete the data stored in the system about the samples.
- From the Sample details page, select Remove from Storage from the Manage menu.
- From a grid of Samples, select samples using the checkboxes and select Storage > Remove from Storage.
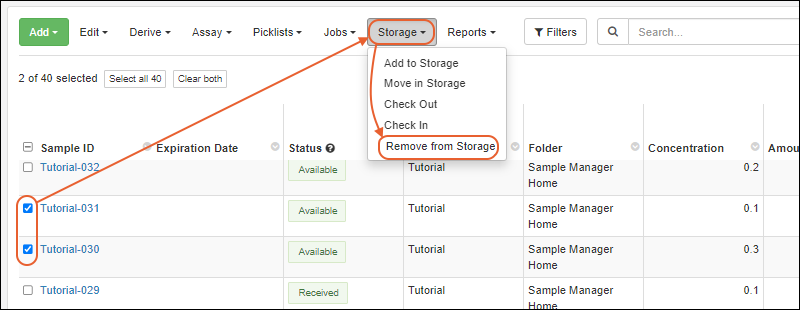
- From the Storage View of the Sample(s) location, select the sample(s), then click Remove.
Update Status and Amount of Removed Samples
In all methods of removal from storage, you will see a popup detailing the sample(s) you selected to remove. You'll see their locations, a color indicating type, and position information.Use checkboxes to optionally update Sample details upon removal:- Set Sample Status: assign a new sample status at this time. By default, the status will be set to "Consumed" (as long as that status is defined). You can change this default as desired.
- If you uncheck this box, all samples will be left with the status they had at the time of removal.
- Update Sample Stored Amount: provide a new amount for all removed samples. For example, consumed samples might now have a zero amount.
- If you uncheck this box, the samples will retain the amounts they have at the time of removal.
Export Storage Map
When viewing a terminal storage location, you can export a printable Storage Map to Excel. This can be useful when sharing with colleagues or to provide detailed data access for users going into a physical freezer location where there is no internet.From the stored samples grid, choose > Storage Map (Excel).Related Topics
Storage Roles
Storage Roles
Both storage roles include the ability to read Sample data, but not the full "Reader" role for other resources in the system (such as assay data, data classes, media, or notebooks). Neither storage role grants the ability to edit or delete Sample data. These roles supplement, but do not replace the usual role levels like "Editor" and "Administrator" which may have some overlap with these storage-specific roles.Storage roles support the ability to define users who can:- Manage aspects of the physical storage inventory and read sample data, but not read other data (assays, etc), not change sample definitions or assay definitions: Grant a storage role only.
- Manage aspects of the physical storage inventory and read all application data, including assays, etc, but not change sample definitions or assay definitions: Grant a storage role plus the "Reader" role.
- Manage sample definitions and assay designs, and work with samples and their data in workflow jobs and picklists, but not manage the physical storage inventory. Grant "Editor" or higher but no storage role.
- Manage storage inventories and also manage sample data, assay data, etc.: Grant both the desired storage role and "Editor" or higher.
- Manage storage inventories and be able to update but not delete other data (sample, assay, etc.): Grant both the desired storage role and "Editor without Delete".
Storage Editor
The role of "Storage Editor" grants the ability to read, add, and edit data related to items in storage, picklists, and jobs. The storage-related portion of this role is included with the Administrator role.A user with the Storage Editor role can:- Add, move, check in/out and remove samples from storage.
- Create, update and delete sample picklists.
- Create workflows and add or update sample assignments to jobs.
- Update a sample's status.
- Move existing storage and box locations, provided the Storage Editor role is assigned where the storage is defined, i.e. in the home project.
- Does not include permission to insert new locations (Storage designers have that permission).
- Does not include permission to read data other than sample data.
- Does not include permission to edit or delete sample data.
Storage Designer
The role of Storage Designer confers the ability to read, add, and edit data related to storage locations and storage units. Administrators also have these abilities.- Create, update, and delete storage locations, freezers, and storage units.
- They cannot delete locations that are in-use for storing samples. The Application Administrator role is required for deleting any storage that is not empty.
- Does not include permission to add samples to storage, check them in or out, or update sample status.
- Does not include permission to update picklists or workflow jobs.
- Does not include permission to read data other than sample data.
- Does not include permission to edit or delete sample data.
Related Topics
Check Out and Check In
Check Out
To check out samples, you have many options in the interface:- Navigate to the storage location, select the sample or samples you want, and click Check Out.
- From any grid of samples, use checkboxes to select the sample(s) you want and select Storage > Check Out. On a narrower browser, this option will be under the More > menu.
- From the sample details page, select Manage > Check Out to check out a single sample.
Check In
When finished with a sample, you also have several options:- Navigate to the storage location, select the space(s) reserved for the sample(s) you are checking in, and click Check In (shown below).
- You can check one or more samples in from a samples grid by selecting the row(s) for the sample(s) and choosing Storage > Check In.
- From the sample details page for a sample that is currently checked out, you can select Manage > Check In.
- In the popup:
- Enter the Amount used during checkout.
- By default, the system will Increment freeze/thaw count on check in, under the assumption that you thawed the sample to use it and it will freeze again. If this is not true, you may uncheck this box.
- Enter Reason for Check In if required or desired.
- Click Check In Sample(s).
Related Topics
View Storage Activity
Your Activity
From the main menu, click Storage, then scroll down and click Your recent sample activity.- Click a Sample Id to see the details for a sample.
- Click a storage Status ("In" or "Out") to jump directly to the Storage View of the location for that sample, whether it is currently checked in or out.
- Note that the Status column is always updated to the current storage status of the sample, not the status at the time of the activity in this grid. For example, samples that were "removed from storage" could still have the current status of "In" if they were re-added to storage later, perhaps in a different location.
Stored Items
From the Storage dashboard, click the message View #### total samples in storage in the Sample Activity panel. Here you'll see a grid of all items in storage throughout the system. You can view all samples at once, or use the tabs for each sample type to access sample grid menus for samples of that type.Sample Timeline: Storage Activity
Like other events that happen for an individual sample, storage events will be included on the Sample Timeline, including adding samples to storage, checking in and out, updating storage data, moving the storage unit that contains the sample, and removing the sample from storage.Audit Log: Storage Management Events
Storage management events are logged in the application's Audit Log. From the pages for a storage system, select Manage > View Audit History to open the audit log filtered to that specific storage system.Related Topics
Migrate Storage Data into LabKey
- Define Storage
- Templates for Sample Storage
- Adjusting Storage Fields to Match Other Systems
- Import of Samples with Storage Location Data
- Work with Stored Samples
Define Storage
You can create all the virtual storage you need in Sample Manager or Biologics LIMS first, or you can define new storage 'on the fly' during import of new samples from a file.To define storage up front, follow the steps in the following topic, taking care to use the naming and layout that will match the pathways to storage locations that will be present in the data coming from your previous system. Naming of storage, structures, and terminal locations like boxes and bags is flexible enough to match the system you were using. If necessary, you can also create custom storage unit definitions as needed. Such storage unit definitions can be specified by name in the "Storage Unit" column when adding new storage in a file.Templates for Sample Storage
Once you have defined your storage, the next step is to create the Sample Types you will be storing, if they don't already exist in the system. You can use a combination of field inferral from spreadsheets and manual refinement to match the types of samples you have stored. Once your Sample Type exists, whether or not it contains Samples, download an import spreadsheet template from the Sample Type dashboard as described here:- StoredAmount
- Units
- FreezeThawCount
- StorageLocation (A slash-delimited "path" to the terminal storage location - see below)
- StorageRow
- StorageCol
- StoragePositionNumber (an alternative to using Row/Column positions)
- StorageUnit (The type of the storage unit; required when defining new storage during import)
- StorageUnitLabel (A label for a new storage unit)
- EnteredStorage (Date)
- CheckedOut (Date)
- CheckedOutBy (UserID)
- StorageComment
Data Format Requirements
- StoredAmount
- Must be a number.
- May be negative.
- Units
- Must be one of the supported unit types listed here.
- May be supplied as either the full plural name (e.g., "grams") or the abbreviation (e.g., "g"). Casing matters.
- If the Sample Type has a default unit type provided, the units must be convertible to this unit type. (e.g., "g" and "kg" are convertible, "g" and "mL" are not).
- Note that if a value for "Units" is included in your import spreadsheet, you must also provide a storage location, even if the "StoredAmount" is blank. To import a mixed set of samples, some of which have storage locations and some of which do not, be sure to remove the value from the "Units" column for any samples not in storage.
- FreezeThawCount
- Must be a non-negative integer.
- StorageLocation
- Must contain valid names of locations in a hierarchy separated by slashes (/), for example:
- Freezer #3 / Shelf #1 / Box #1 - Green
- Names of locations may contain slashes, but if so, then they must be quoted.
- Leading and trailing spaces are tolerated and trimmed.
- A leading or trailing slash is tolerated.
- If any storage locations in this path do not exist, they will be created in the background (provided the entire sample import succeeds).
- StorageRow
- Must be either a positive integer or one of a series of letters (A-Z, a-z). Casing does not matter for the letters.
- DOES NOT check if the display format for the box type corresponds to the format provided as input. That is, if you’ve chosen to label your axes as Numeric, you can still provide alphabetic designations for the row value (A for row 1) in the input file.
- Must be in the range of the chosen terminal storage type (cannot be < 0 or larger than the number of rows configured for the terminal storage type).
- StorageCol
- Must be blank if the terminal storage type is a bag or a cane.
- If not blank, must be either a positive integer or one of a series of letters (A-Z, a-z). Casing does not matter for the letters.
- DOES NOT check if the display format for the box type corresponds to the format provided as input.
- Must be in the range of the chosen terminal storage type.
- StoragePositionNumber
- A hidden field by default. Provides an alternative to specifying a row and column.
- If not blank, must be a positive integer.
- StorageUnit
- The type of the storage unit. This value must match the name of an existing storage unit type.
- If you are creating any new storage, this field must be populated for the first row of the new storage unit.
- StorageUnitLabel
- An optional label to use in addition to the storage unit name, which is included at the end of the "Storage Location" value provided.
- EnteredStorage
- Must be a valid date or date-time value.
- Represents when the sample was stored in the physical storage system.
- If omitted, the time of the data import, i.e. the time of entering the management system is used.
- CheckedOut
- Must be a valid date or date-time value.
- If omitted and a CheckedOutBy user is provided, the current date will be used.
- CheckedOutBy
- Must be a valid email address, username, or userID for a user in the system with sufficient permissions to check out samples. Learn about adding users here: User Accounts, Groups, and Roles.
- Leading spaces and quote marks are not tolerated.
- If omitted and a CheckedOut date is provided, the sample will be checked out by the user importing the data.
- StorageComment
- Always optional.
- Will be ignored if the action for the import is not checking in, checking out, or removing from storage.
Adjusting Storage Fields to Match Other Systems
Examine the export spreadsheet from your previous storage system. It likely includes similar columns, potentially requiring name changes or other adjustments to match the expectation of the template you generated.This section will also be helpful if you are updating sample data to change storage-related fields from a spreadsheet or using the API. Some fields, such as StorageStatus cannot be user-set. They are determined by the presence or absence of values in other fields.Modify your exported data as needed, noting the following requirements for recording specific sample actions in the system:Add a sample to storage- The StorageLocation and StorageRow columns must be present and valid.
- If adding to a Box, the StorageCol column must be present and valid.
- If adding to a Bag, the StorageCol column must be absent or empty.
- Other fields may be present but are not required.
- The StorageLocation column must be present and contain the path ending in the new terminal storage for this sample. Any locations along this path, including the freezer itself, will be created if they don't exist.
- The StorageUnit column must be present and the value must match an existing storage unit type.
- The StorageRow columns must be present and valid.
- If adding to a Box, the StorageCol column must be present and valid.
- If adding to a Bag, the StorageCol column must be absent or empty.
- Other fields may be present but are not required.
- SampleId column must be present and not empty.
- At least the StorageLocation column must be present and empty.
- If the StorageRow and StorageCol columns are present they should also be empty.
- No other storage metadata columns should have values.
- SampleId column must be present and not empty.
- The corresponding metadata columns should be present.
- If the StorageLocation column is present, the StorageRow must be present and, when updating items in a box, the StorageCol column must also be present. All columns must have valid values. If updating items in a bag, the StorageCol column value should be empty if present.
- SampleId column must be present and not empty.
- The StorageLocation column and the StorageRow column must be present with valid values. If moving to a Box type, the StorageCol column must also be present with a valid value. If moving to a bag, the StorageCol column must be empty if present.
- SampleId column must be present and not empty.
- Either the CheckedOut or the CheckedOutBy column must be present and not empty.
- If the CheckedOut column is valid and the CheckedOutBy column is empty/absent, the item will be checked out by the current user.
- If the CheckedOutBy column is valid and the CheckedOut column is empty/absent, the item will be checked out by the user in the CheckedOutBy column using the current date.
- If both the CheckedOut and the CheckedOutBy columns are present, they must both have valid values.
- If both the CheckedOut and the CheckedOutBy columns are empty, the system will not recognize this as a checkout action.
- SampleId column must be present and not empty.
- One of the CheckedOut or CheckedOutBy columns must be present and empty.
- Note that with a file import, it is possible to perform multiple operations at once. For example, you may provide a row that both checks a sample out and moves it to a new location. For samples, there will be two timeline events created for this. There is no guarantee about the order of these events in the timeline.
- Since updating of storage happens only via sample import, there will always be a “Sample update” timeline event, even if nothing in the sample actually changed.
- If you are importing data for a set of samples in which some are in storage and some are not, be sure to only provide values for the storage related columns (including "units") for samples which also include storage locations.
Import of Samples with Storage Location Data
Once your spreadsheet is ready:- If you are creating all new samples with storage details, use the Add > Import from File, or Add Samples > Import From File from the dashboard.
- If you are adding storage information for existing samples, use Edit > Update from File from the sample grid.
- If your spreadsheet contains a mixture of existing and new samples, use Edit > Update from File and select the option to Create new samples too.
Work with Stored Samples
You can now use all the Storage Management features as if you had stored samples using the user interface:- Check Out and Check In
- Move Stored Samples
- View Storage Activity
- Create New Storage while Importing Samples from File
Related Topics
Workflow
- Create jobs to track and prioritize sequential tasks
- Assign work to the right users
- Use workflow templates to standardize common task sequences
- Track progress toward completion
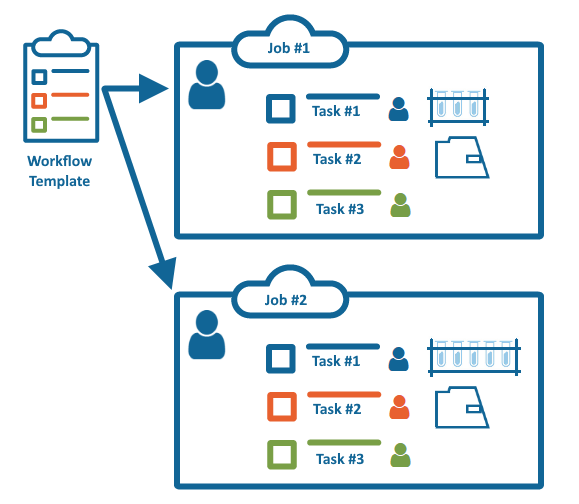 Administrators and users with the Workflow Editor role can create and edit workflow jobs and tasks. Only administrators can create job templates.
Administrators and users with the Workflow Editor role can create and edit workflow jobs and tasks. Only administrators can create job templates.
Topics
- LIMS: Workflow
- Manage Job Queue
- Complete Tasks
- Manage Notifications
- Edit Jobs and Tasks
- Create a Job Template
- Manage Jobs and Templates
Related Topics
Start a New Job
- Start a Job
- Start a Job with Selected Samples
- Job Details
- Tasks
- Input Samples
- Finish Creating Job
- Create Job From Template
Start a Job
- From the home page, click Start a New Job.
- From the main menu, choose Workflow, then click Create Job.
- Select some samples, then use Jobs > Start a New Job from any Sample or Picklist grid. On narrower browsers, this option will be under the More > menu.
Start a Job with Selected Samples
If you already know the set of samples you want to include, start from the Sample Type grid, storage view, or from a picklist containing the desired samples. This walkthrough illustrates using the Sample Type grid, but the process starting from other lists of samples is the same.- Select the sample type of interest from the Sample Types section of the main menu.
- Use filtering and checkboxes to select the Samples of interest.
- If Samples are in storage, you can use Storage Status as part of your selection criteria here. If you want to use storage location, start from the storage system where they are located instead.
- Select Jobs > Start a New Job from the menu above the grid. On narrower browsers, this section will be under the More > menu.
Job Details
On the first panel of the job creation wizard, enter details about the job:- Job Name: Provide a name for the job, or leave blank to have one generated for you.
- Description: Include a description. Note that to include a newline, you must use Shift-Enter; using Enter without Shift will save the description value as a single line.
- Job owner: This is the "owner" of overall job completion. The owner can be an individual user or a user group and may or may not be the same as the user(s) assigned to tasks in the job.
- Notify these users: Add users or groups who should get notifications as this job progresses. Users on this list will be able to follow this job on their tracked jobs list.
- Job start and due dates: Use the date picker to select the begin and end dates.
- Priority level: Use the pulldown menu to select one of the options:
- Low
- Medium
- High
- Urgent
- Attachments: Select or drag and drop any files needed for the job. For example, an SOP document, labels, or other instructions related to the job could be included here.
- Any files you upload will be listed; if you need to delete one, click the to remove it.
Tasks
Click the Tasks section to open it.Any job can be composed of several tasks to complete in sequence. Click Add Task to add each task and click to open the details panel.- Name
- Description: Note that to include a newline in your description, use Shift-Enter; using Enter without Shift will save the value.
- Assign to: Assign to a user or a user group.
- Assays to Perform (select as many as required for this task)
- Due date
Input Samples
Click the Input Samples section to open it. If you created this job from a set of samples, it will open on the Included Samples tab and you will see them listed. If not, skip ahead to add samplesIncluded Samples
- Review the listed set of included samples; if necessary select one or more rows and click Remove from Job to remove them.
- If you like, you can use the Add Samples tab (described below) to add more samples.
- When you are satisfied with the selection of samples, skip to the Finish Creating Job section.
Add Samples
- If you did not start creating this job from a selection of samples, when you open the Input Samples panel, click the Add Samples tab (if it is not open by default).
- Select the Sample Type.
- Use grid filtering, sorting, and searching to find the desired samples.
- Check the boxes to select the samples you want to include. Once samples are selected, the "Add Samples" button will be activated.
- Click Add Samples.
Finish Creating Job
Click Finish Creating Job to start the job. You will see the job overview. Note the tabs along the top edge for viewing Tasks, Samples, and Assays in addition to the Overview.Create Job From Template
You can create a job from a template you have already saved, with or without the preselection of samples. Creating a job from a template follows the same wizard process, but the details and tasks are prepopulated.- At the top of the job creation page, click Choose Job Template.
- Find the template you want to use; typing ahead will narrow the options.
- If you have already defined any tasks before applying a template, you will be warned that the template will override any existing information in your job. Cancel to retain your current tasks.
- Click to see details for any template, including creation details, number of tasks, and the description.
- Click Choose Template.
Add Job Details and Priority
The template does not prepopulate the Job Details and Priority panel; complete it as when you are creating a job without a template as described above.Define Job Tasks
The job tasks from the template are prepopulated in the job creation wizard. By default these tasks are locked. You can assign tasks to users and add due dates without editing tasks. If the template allows editing of the assays in jobs using it, the Assays to Perform box will be unlocked on all tasks so that you can define assays to run for this job.If you want to make any changes in the tasks for this job, click Edit Tasks and edit using the same interface as when you create tasks without a template. Note that editing job tasks will remove the template association from the job details as well as remove any files associated with the template from this job. You'll be asked to confirm before proceeding.Complete Wizard
When you are defining a job starting from a template, the section for Input Samples is the same as above for creating a job that did not start from a template.When you have completed all the sections of the job creation wizard, finish as described above.Jobs that were created from templates show the template name on the Overview tab of the job details. Click the job name to open the job details, then click the template name to open the template itself.Related Topics
Manage Job Queue
Jobs List Dashboard Panel
On the home page, the Jobs List shows Your Job Queue by default. This is the list of jobs that are either assigned to you or include tasks assigned to you. Filter your queue by Priority Level using the dropdown.Workflow Home
Click Workflow Home or select Your Job Queue from the main menu to see the jobs list in grid form.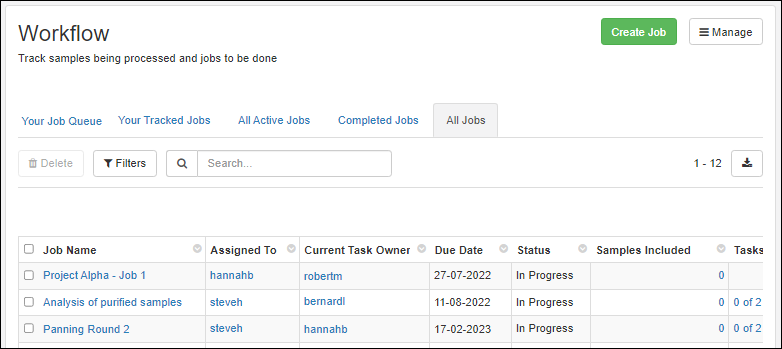 By default, you'll see Your Job Queue sorted by due date. Like other grids, you can use filtering, sorting, searches, and custom views on the grid of jobs.The tabs each show the count of jobs in the different categories:
By default, you'll see Your Job Queue sorted by due date. Like other grids, you can use filtering, sorting, searches, and custom views on the grid of jobs.The tabs each show the count of jobs in the different categories:
- Your Job Queue: Jobs and tasks assigned to you.
- Your Tracked Jobs: All Jobs in which you appear on the Notify List. This list is initially filtered to those jobs that are currently "In Progress".
- All Active Jobs: Jobs that have not been completed, including but not limited to the ones in your own queue.
- Completed Jobs: Jobs that have been completed.
- All Jobs: All of the above.
Your Job Queue
At any time in the application, you can select Your Job Queue from the Workflow section of the main menu to jump to a detailed view of your own work assignments.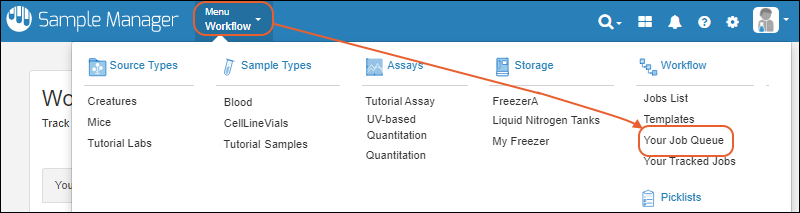
Your Tracked Jobs
You can access a grid of all jobs which include you on the Notify List by selecting Your Tracked Jobs from the Workflow section of the main menu.Related Topics
Complete Tasks
- On the Jobs panel on the home dashboard under Your Queue if it is assigned to you. You can filter by priority to find the most pressing work.
- If the job or current task is not assigned to you, you can click the Active Jobs tab on the home page.
- Use the main menu from anywhere in the application. Click Jobs List under Workflow for the grid of all jobs.
Topics
Overview Tab
On the Overview tab, you will see the job details, assignments, and list of tasks with status of each task. Status values include:- In Progress: the current/active task to be completed
- Pending: future tasks that need to wait for the 'in progress' task
- Complete: completed tasks.
Tasks Tab
Switch to the Tasks tab to see a more detailed view of job progress. Each task is listed on the left, with details of the selected task on the right.Completed tasks show a green check. Tasks that involve assays include links to View Data | Import Run.Comments
On the Tasks tab, you can enter Comments to accompany any task, including the ones marked completed. Click Start a thread to add a new comment.- You can use markdown formatting, with the help of buttons for bold, italic, link, list, and numbered list formatting.
- Switch the Markdown mode dropdown to Preview to see the formatted result.
- Click the to attach files to comments.
- Click Add Comment to save.
Complete Task
If the 'In Progress' task is assigned to you, and you have performed the actual work described, you can click Complete Current Task on the Overview tab (or Complete Task on the Tasks tab) to mark it completed. The task status changes to Complete and the next task on the list is now In Progress.Complete Tasks with Samples
When your task involves samples, you can access and review them on the Samples tab. Samples of all types are shown on a shared All Samples tab, with common properties like storage information and status. The samples of an individual type are available on additional tabs ("Plasma" and "Serum" shown here), with the properties specific to that type.- Add Samples
- Remove Samples
- More: Other menus vary based on whether you are viewing all samples or samples of only one type. Learn more about the options in this topic:
- Data Grid Basics
Complete Tasks involving Assay Import
On the Tasks tab, select the task that involves an assay. Under Assays, you'll see a pair of links for each assay involved in this task.- View Data: Once there has been data uploaded for this assay in this task, this link will take you to the filtered run.
- Import Run: Open the importer for the requested assay.
- If there are samples assigned to the job, by default you will see the Enter Data into Grid option, with rows prepopulated for the samples assigned, making it easy to manually enter values.
- When the assay is set to associate samples only with a single sample type, the samples will be filtered so that only the relevant samples of that type are included in the grid.
- You can also use bulk insert, update, and Import Data from File methods if desired.
Reassign Task
From the Tasks view, users with sufficient permissions can reassign tasks that have not been completed by changing the Task Owner. Use the dropdown to select the new task owner.Reopen Task
If you find you have closed a task by mistake, you can reopen it using the Reactivate Task button.Related Topics
Edit Jobs and Tasks
- Update Job Details
- Manage Menu
- Edit Job Tasks: Add, Reorder, Edit, Delete
- Edit Samples Assigned to a Job
- Reactivate Closed Task
- Reactivate Completed Job
- Delete Job
- Move Job
Update Job Details
In the left-hand panel you can edit many common job details including Owner, Notify list, Priority Level, Start and Due Dates, and the Description. Use dropdowns or icons to edit. Click outside the selection area for an element to save changes.Rename a Job
To change the name of the job, click the next to the name, type the new name and hit return (or click away) to save.Add or Delete Files from a Job
Files attached to a job are shown in the Attachments section on the Overview tab.- Add a new file using the Select file or drag and drop here box.
- Use the menu to:
- Download
- Remove Attachment
Manage Menu
Edit Tasks: Add, Reorder, Edit, Delete
- Click the Tasks section.
- Any tasks that have been completed (or are part of a template) cannot be changed and will be shown with a icon.
- Add more tasks via Add Task. (You cannot add tasks to a job created from a template.)
- Reorder tasks by dragging and dropping the six-block handle on the left.
- Delete a task by clicking the icon on the right.
- Click a task panel to open the details for editing.
- If the job was created from a template that allows editing the assays, you'll be able to add or remove Assays to Perform for the task.
Edit Samples Assigned to a Job
While editing job tasks, you can use the Input Samples section to adjust the set of samples within the job. You can also click the Samples tab from within the job and click Add Samples to open the same interface.Follow the instructions in this topic to add samples within the job wizard. If you only need to add or remove samples, it may be easier to follow the steps below to add or delete samples from jobs.Add Samples to a Job
From the main menu, click the name of the type of samples you wish to add to open the grid of available samples. Alternatively, choose Picklists from the user menu and click the name of the curated picklist you want to use. Select the desired sample(s) using the checkboxes and select Jobs > Add to Job. In the popup, select the desired job by name from the dropdown. Click Add to Job.Filter Samples in a Task
You can define sample filters to specific tasks, which ensures that only the specified samples are included when importing assay data for that task. This gives you more precise control over data import and improves workflow accuracy and efficiency.Task-level filters can be applied to individual jobs or to workflow templates.When applied to workflow templates, even if there are pre-existing jobs associated with them. Administrators can define if the sample filter is editable in jobs based on the template. If Allow Edit of Filters in Job is checked, then during job creation & during execution, the sample filters may be changed. If left unchecked, then during job creation & execution, the sample filters may not be changed.To add a task-level filter, navigate to the task, and click Add Filter.
Delete Samples from a Job
Open the job and click the Samples tab. Select the sample(s) you wish to delete and click Remove Samples.Reactivate Closed Task
To reactivate a closed task, click the Tasks tab in the upper left, then click the name of the closed task to select it. It will be shown with a green check icon. In the following image, "Prepare Samples" is in a closed state. Click Reactivate Task.Reactivate Completed Job
If you need to reactivate a job that was marked completed, find it by opening the Workflows page from the main menu and clicking Completed Jobs. Click the desired job name to open it.Delete Job
Learn about deleting a job in this topic:Move Job
Learn about moving a workflow job in this topic:Related Topics
Create a Job Template
Create a Job Template
To create a new job template:- Select Templates from the main header menu Workflow section.
- Click Create Template.
Template Details
Enter the Template Name and Description. The name should be unique enough to help users find it on a menu.Tasks
Click the Tasks tab and define the tasks. For each, enter the Name and Description and select any Assays to Perform as part of that task. If you want to allow jobs that use this template to be able to change which assays should be performed, check the box to Allow Edit of Assays in Job.Finish Creating Template
Click Finish Creating Template when finished. You'll see the new template overview.Custom Fields
Workflow job templates can include custom fields to support standardizing various workflows. These custom fields are defined using the standard field editor when creating or editing a template. You might include fields you want every user to complete, such as a billing code or site identifier. Custom fields can be required, and can be of any standard field type, including lookups and text choice fields for limiting user choices.- During job creation when users choose a template, they'll see a summary of fields.
- The name of the section includes the name of the template.
- Required fields need to be populated at this time; they must have values before you can create the job.
- On the job details panel, you'll see and can populate custom fields by clicking the .
Related Topics
Manage Jobs and Templates
Manage Jobs
From the home dashboard, click Workflow Home or from anywhere in the application, select from the main menu:- Jobs List: Open the dashboard on "All Jobs".
- Templates: Open the template management page.
- Your Job Queue: Open the dashboard on "Your Job Queue".
- Your Tracked Jobs: Open the dashboard on "Your Tracked Jobs".
Edit Jobs
To open an individual job for editing, click the Job Name. Details of editing jobs and tasks are covered in this topic:Delete Jobs
To delete one or more jobs, use the checkboxes on the Job List to select jobs for deletion, then click Delete.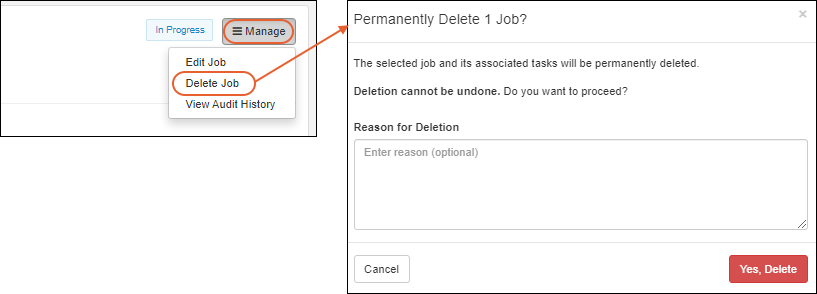
Manage Templates
Templates are managed by clicking Templates in the Workflow section of the main menu, or by selecting Manage > Job Templates from the workflow dashboard:Edit Template
To edit a template, open it as shown above and select Manage > Edit Job Template.For any template, you can edit the Name and Description, and add (or remove) custom fields.Copy Template
To make a new template using an existing one as a starting point, select Manage > Copy Job Template.Delete Templates
To delete a single template, open the details page as shown above and select Manage > Delete Job Template. Once a job has been created from a template, you can no longer delete it and this option will be inactive.To delete one or more templates, select them on the Workflow > Templates dashboard page using the checkboxes, then click Delete.Related Topics
Electronic Lab Notebooks (ELN)
Notebooks
On the dashboard, you will see summary cards for your most Recently Opened notebooks, a summary of actions For You, and many ways to filter and find specific notebooks of interest.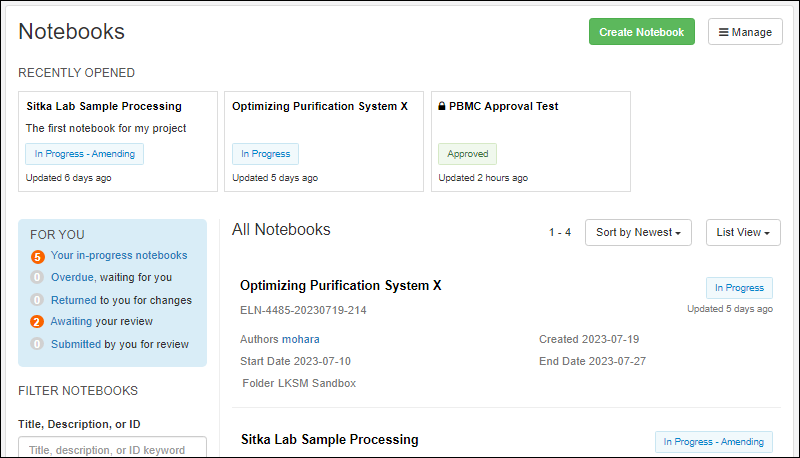 Learn more in this topic: Notebook Dashboard
Learn more in this topic: Notebook Dashboard
Notebook Notifications
To enable or disable email notifications, select Notification Settings from the user menu. Use the checkbox to subscribe:- Send me notifications when notebooks I am part of are submitted, approved, or have changes requested
- When a user starts a thread on a notebook:
- The email subject field will follow the pattern: <USERNAME> commented in <NOTEBOOK NAME>: "<FIRST FIVE WORDS>..."
- Recipients include: notebook author(s)
- The thread author will not be in the notification list. If the thread author happens to be the same as the notebook author, this author will not get an email notification.
- When a user comments on a thread:
- The email subject field will follow the pattern: <USERNAME> replied in <NOTEBOOK NAME> on thread: "<FIRST FIVE WORDS OF ORIGINAL THREAD> ..."
- Recipients include: notebook author(s) and the person who started the initial thread.
- The current commentator shall not be in the notification list. If the commentator happens to be the same as the notebook author, this person will not get an email notification.
- If a user's "Messages" notification setting at the folder level is set to "Daily digest of my conversations", then:
- The individual email subject headers within the digest will follow the pattern: "<USERNAME> commented [...]" as specified above.
- The subject-line of the digest email itself will not include comment details.
- A notification will be included in a user's digest if they have participated in the thread, if they are on the notify list, or if they are among the authors of a notebook that has been commented on.
Topics
Notebook Dashboard
- Notebook Dashboard
- Find and Filter Notebooks
- All Notebooks Listing
- Working with Notebooks in Folders
Notebook Dashboard
On the dashboard, you will see:- Recently Opened: Summary cards for your most recently viewed notebooks. Click any card to see the full notebook.
- For You: A summary of actions for you, with alert-coloring and the number of notebooks you have in each group. Click the link to see the
- Your in progress notebooks
- Overdue, waiting for you
- Returned to you for changes
- Awaiting your review
- Submitted by you for review
- Filter Notebooks: Use many ways to filter and find specific notebooks of interest.
- All Notebooks: All notebooks in the system, with any filters you apply narrowing the list.
- By default, this list is filtered to focus on your own notebooks.
- If you have not authored any notebooks, you'll see an alert and be able to click to clear the filter to see others' notebooks.
Find and Filter Notebooks
There are many ways you can filter notebooks to find what you need:- Title, Description, or ID: Type to search for any word or partial word or ID that appears. When you enter the value, the list of notebooks will be searched. Add additional searches as needed; remove existing ones by clicking the 'x' in the lozenge for each.
- Contains Reference To: Type to find entities referenced in this notebook. You'll see a set of best-matches as you continue, color coded by type. Click to filter by a specific entity.
- Created: Enter the 'after' or 'before' date to find notebooks by creation date.
- Folders: When Folders are in use, you can filter to show only Notebooks in specific Folders.
- Created from Templates: Type ahead to find templates. Click to select and see all notebooks created from the selected template(s).
- Authors: Type ahead to find authors. Click to select.
- By default the notebook listing will be filtered to show notebooks where you are an author.
- Add additional authors as needed to find the desired notebook(s).
- Remove a filter by clicking the 'x' in the lozenge.
- Reviewers: Type ahead to find reviewers. Click to select. Add additional reviewers to the filter as needed; remove a filter by clicking the 'x' in the lozenge.
- Review Deadline: Enter the 'after' or 'before' date to find notebooks by review deadline.
- Status: Check one or more boxes to show notebooks by status:
- Any status
- In progress
- Submitted
- Returned for changes
- Approved
All Notebooks Listing
The main array of notebooks on the page can be shown in either the List View or Grid View. You'll see all the notebooks in the system, with any filters you have applied. By default this list is filtered to show notebooks you have authored (if any). When the list is long enough, pagination controls are shown at both the top and bottom of either view.Sort notebooks by:- Newest
- Oldest
- Due Date
- Last Modified
Working with Notebooks in Folders
When you are using Folders, you'll see the name of the Folder that a notebook belongs to in the header details for it. Filtering by Folder is also available.Note that if a user does not have read access to all the Folders, they will be unable to see notebooks or data in them. Notebooks that are inaccessible will not appear on the list or grid view.Authors can move their notebooks between Folders they have access to by clicking the icon. Learn more here.Related Topics
Author a Notebook
- Create New Notebook
- Add to a Notebook
- Add to an Entry
- Rename an Entry
- Add References
- Add a New Day
- Comment on an Entry
- Add Attachments
- Add a Table
- Adjust Wide Tables
- Edit a Notebook
- Manage Entries
- Archive Notebook
- Submit for Review
Create New Notebook
To create a new notebook, click Create Notebook from the Notebooks Home dashboard.- Enter a Notebook Title.
- If you are an administrator, you can click Create new tag to add a new one.
- Enter a Description.
- Select Start and end dates.
- By default, You are an author. Use the selection menu to add more Co-authors.
- If you have templates defined, you can choose one to use by clicking Browse Templates.
- Click Create Notebook.
Notebook ID
An ID will be generated for the notebook and shown in the header.Expand and Collapse Detail Panel
The left panel of the notebook screen displays a table of contents, full listing of references to this notebook as well as items this notebook references, and attachments for the notebook and any entries. This section can be collapsed by clicking the and expanded again by clicking the .Table of Contents
The Table of Contents for a notebook lists all the entries, and for each entry, the days and any heading text are listed, indented to make the general structure of the document visible at a glance and easy to navigate.Create New Tag (Biologics LIMS Only)
During notebook creation in Biologics LIMS, the author can select an existing tag. If that author is also an administrator, they have the option to click Create new tag to add a new one.Learn more about tags in this topic:Add Custom Fields (Experimental Feature, Biologics LIMS Only)
To define and use custom fields for your Biologics LIMS notebooks to support additional classification that is meaningful to your team, you must first enable the experimental feature "Notebook Custom Fields".Once enabled, you will see a Custom Fields section. Use Add custom field to select an existing field to add to this notebook. Type ahead to narrow the list of existing fields to find the one you want. Shown here, the "ColumnTemperature" field is added and you can now provide a value for this field for this notebook.Rename a Notebook
A notebook author can click the to rename a notebook. Non-authors will not see the edit icon. While the system does not require names to be unique, you will want to choose something that will help your colleagues identify it on lists and dashboards.Copy a Notebook
Once you have created and populated a notebook, you can copy it to create a new notebook with the same details. This is similar to creating a new notebook from a template, except that templates do NOT include the value of any custom fields, and copied notebooks do include these values. You do not need to be a notebook author to make a copy of it. Select Save As > Copy.Add to a Notebook
A notebook lets you record your work in a series of entries, each of which can have a custom name, span multiple days, include references to data in the application, and support entry-specific comment threads.The collapsible detail panel on the left lists the Table of Contents, references (both to this notebook and from this notebook to other entities), attachments, and includes an edit history timeline. On the right, the header section lists the name and ID, the authors, shows the tag (if any), creation details, and status. A new empty notebook looks like this:Add to an Entry
The New Entry panel is where you can begin to write your findings and other information to be recorded.The formatting header bar for an entry includes:- Insert: Select to add:
- Styling menu: Defaults to Normal and offers 3 heading levels. Heading sections are listed in the Table of Contents for easy reference.
- Font size: Defaults to 14; choose values between 8 and 96 point type.
- Special characters: Add mu, delta, angstrom, degree, lambda, less than or equal, greater than or equal, and plus minus characters.
- Bold, Italics, Underline, Strikethrough
- : Link selected text to the target of your choice.
- / : Make a text selection a super- or sub-script.
- Text color. Click to select.
- Alignment selections: Left, center, or right alignment; indent or dedent.
- : Numbered (ordered) lists
- : Bullet (unordered) lists
- : Checkboxes
- and : Undo and Redo.
- : Clear formatting.
Rename an Entry
Click the next to the "New Entry" title to rename it.Add References
Within the entry panel, you can use the Insert > Reference menu, or within the text, just type '@' to reference an available resource.Add a New Day
Place the cursor where you want to add a marker for a new date. Select Insert > New Day. A date marker will be added to the panel for today's date.Comment on an Entry
Click Start a thread to add a comment to any entry. Each entry supports independent comment threads, and an entry may have multiple independent comment threads for different discussions.By default you will enter comments in Markdown Mode; you can switch to Preview mode before saving. Other dashboard options include bold, italic, links, bullets, and numbered lists. Click the button to attach a file to your comment.Type and format your comment, then click Add Comment.- Edit comment (including adding or removing files attached to comments)
- Delete thread
Add Attachments
Attachments, such as image files, protocol documents, or other material can be attached to the notebook.- To add to the notebook as a whole, click the Attachments area in the detail panel on the left (or drag and drop attachments from your desktop).
- To add an attachment to an entry, select Insert > Attachment. You can also paste the image directly into the entry; it will be added as an attachment for you.
- Copy link: Copy a link that can be pasted into another part of an ELN, where it will show the attachment name already linked to the original attachment. You can also paste only the text portion of the link by using CMD + Shift + V on Mac OS, or CTRL + Shift + V on Windows.
- Download
- Remove attachment (available for authors only)
Add a Table
You can add a table directly using the Insert > Table menu item, directly entering content. You can also paste directly from either Google Sheets or Excel either into that table, or into a plain text area and a table will be added for you.Adjust Wide Tables
After adding a table to an ELN, you can use the table tools menu to add/remove columns and rows and merge/unmerge cells. Drag column borders to adjust widths for display.Edit an Entry
Entry Locking and Protection
Many users can simultaneously collaborate on creating notebooks. Individual notebook entries are locked while any user is editing, so that another user will not be able to overwrite their work and will also not lose work of their own.While you are editing an entry, you will periodically see that it is being saved in the header:Edit History
The edit history of the notebook can be seen at the bottom of the expanded detail panel on the left. Versions are saved to the database during editing at 15 minute intervals. The last editor in that time span will be recorded. Past versions can be retrieved for viewing by clicking the link, shown with the date, time, and author of that version.Manage Entries
Your notebook can contain as many entries as needed to document your work. To add additional panels, click Add Entry at the bottom of the current notebook. You can also add a new entry by copying an existing entry.Use the menu to:Archive Entry
You cannot completely delete an entry in a notebook. Archiving an entry collapses and hides the entry.- You can immediately Undo this action if desired.
- Note that when an entry is archived, any references in it will still be protected from deletion. If you don't want these references to be 'locked' in this way, delete them from the entry prior to archiving. You could choose to retain a plain text description of what references were deleted before archiving, just in case you might want to 'unarchive' the entry.
Copy Entry
Create a duplicate of the current entry, including all contents. It will be placed immediately following the entry you copied, and have the same name with "(Copy)" appended. You can change both the name and position.Reorder Entries
Select to open a panel where you can drag and drop to rearrange the entries. Click Save to apply the changes.Archive Notebook
You cannot delete a notebook (just as you cannot fully delete a notebook entry, but you can Archive it so that it no longer appears active. Archived notebooks will be locked for editing, removed from your list of recently accessed notebooks, and will no longer show in your active notebooks. They can be restored at a later time.To archive a notebook, select Archive > Archive Notebook from the top of the notebook editing page.Access Archived Notebooks
From the Notebook Dashboard, select Manage > Archive. You'll see a grid of archived notebooks making it easier to restore them or otherwise access their contents.Submit for Review
Learn about the review process in this topic:Related Topics
Notebook References
Add a Reference
Within the entry panel, you can use the Insert > Reference menu, or within the text, just type '@' to reference an available resource.- Keep typing to find a single item to reference
- Or, the @ menu lets you reference multiple items in a given category
Auto-Complete References
When you type a reference name, the full-text search results will be shown under where you typed it. The best match is listed first, with alternatives below. You can hover to see more details. Click the desired item to complete the reference.Add References in Bulk
When you type the '@' and don't continue typing, you'll see a menu of categories and can add multiple references in bulk by pasting a list of references by ID (Sample ID, Assay Run ID, ELN ID, etc.).For example, if you click Samples you will next be able to click one of the existing Sample Types, then can paste in a list of Sample IDs, one per line.- Correct any errors and click Recheck References
- Or click to Skip and Add References, skipping any listed and proceeding with only the ones that could be found.
Bulk Reference Assay Batches
When referencing Assays, if the Assay Type you choose includes Batch fields, you'll have the option to reference either runs or batches.View References
Once a reference has been added, you can hover over the color coded lozenge in the text to see more details, some of which are links directly to that part of the data, depending on the type of reference. Click Open in the hover details to jump directly to all the details for the referenced entity.Related Topics
Notebook ID Generation
Notebook ID
When you create a new notebook, a unique ID value is generated for it. This can be used to help differentiate notebooks with similar names, and cannot be edited later. The default format of this ID is:ELN-{userID}-{date}-{rowId}Customize Notebook Naming
When the Professional Edition of Sample Manager, LabKey LIMS, or Biologics LIMS are used with a Premium Edition of LabKey Server, an administrator can adjust the pattern used for generating Notebook IDs by selecting from several options. The selected pattern applies site-wide, i.e. to every new Notebook created on this server after the selection is made. IDs of existing Notebooks will not be changed.- Navigate to the LabKey Server interface via > LabKey Server > LabKey Home.
- Select > Site > Admin Console.
- Under Premium Features, click Notebook Settings.
- Select the pattern you want to use. Options can be summarized as:
- ELN-<USERID>-<DATE>-<ROWID> (Example: "ELN-1005-20230518-30")
- ELN-<DATE>-<ROWID> (Example: "ELN-20230518-30")
- ELN-<ROWID> (Example: "ELN-30")
- Click Save.
| Token | Description | |
|---|---|---|
| <USERID> | ${CreatedBy} | User ID of the user creating the ELN |
| <DATE> | ${now:date('yyyyMMdd')} | The current date using the format yyyyMMdd |
| <ROWID> | ${RowId} | The unique site-wide row ID for the ELN |
Related Topics
Notebook Review
- Submit for Review (Authors)
- Recall Submission (Authors)
- Review Notebook (Reviewers)
- Manage Reviewers (Authors and Reviewers)
- Suggest Changes and Return/Reject this Version (Reviewers)
- Respond to Feedback (Authors)
- Approve Notebook (Reviewers)
- Customize Submission and Approval Text (Administrator)
- Review History
Submit for Review (Authors)
As a notebook author, when you decide your notebook is ready for review, click Submit for Review.- Select one or more reviewers from the Request review by dropdown. You'll see individual users as well as any project groups that have been assigned the "Editor" role.
- You can also provide a deadline, and comments if needed.
- Verify your identity by providing your LabKey login, i.e. email address and password. This verification positively identifies the submitter.
- Check the box to agree with the Notebook Submission Text that can be customized by an administrator.
Recall Submission (Authors)
After a notebook has been submitted, an author has the option to reopen it and click Recall Submission.Review Notebook (Reviewers)
When you are assigned as a reviewer for a notebook, you will receive a notification and see an alert in the For You section, i.e. # Awaiting your review. Click the link to see the notebook(s) awaiting your review.- Close and go home: Conclude this session of review without making any decision. This notebook will remain the same status awaiting your review.
- Suggest changes and return: Click to "reject" this notebook, returning it to the submitters with suggestions of changes.
- Go to signature: Click to "approve" this notebook and proceed to the signature page.
Manage Reviewers (Authors and Reviewers)
Any user or group assigned the Editor role or above can be assigned as the reviewer of a notebook when an author originally submits it for review. When a group is assigned as a reviewer, all group members will be notified and see the task on their dashboard. Any member of that group has the ability to complete the review.As the reviewer of a notebook, you also have the ability to add other reviewers, or remove yourself from the reviewer list (as long as at least one reviewer remains assigned). Open the notebook (but do not enter 'review' mode) and click the icon next to the reviewer username(s).Suggest Changes and Return/Reject this Version (Reviewers)
While in review mode, if you are not ready to approve the notebook, click Suggest changes and return. Provide comments and requests for changes in the popup, then click Yes, return notebook.Respond to Feedback (Authors)
When a notebook is returned for changes, the author(s) will see the Review Status and be able to Unlock and update the notebook to address them.Approve Notebook
As a reviewer in "review mode", when you are ready to approve the notebook, click Go to signature.Customize Submission and Approval Text (Administrator)
The default text displayed when a user either submits a notebook for review, or approves one as a reviewer is "I certify that the data contained in this notebook & all attachments are accurate." If your organization has different legal requirements or would otherwise like to customize this "signing text" phrase, you can do so. The text used when the document is signed will be included in exported PDFs.You must be an administrator to customize signing text. Select > Application Settings. Under Notebook Settings, provide the new desired text.- Notebook Submission Text: Text that appears next to the checkbox users select when submitting a Notebook.
- Notebook Approval Text: Text that appears next to the checkbox users select when approving a Notebook
Review History
At the bottom of the expanded details panel, you'll see the Review History showing actions like submission for review, return for comments, and eventual approval. On a second tab, you can access the Edit History for the notebook.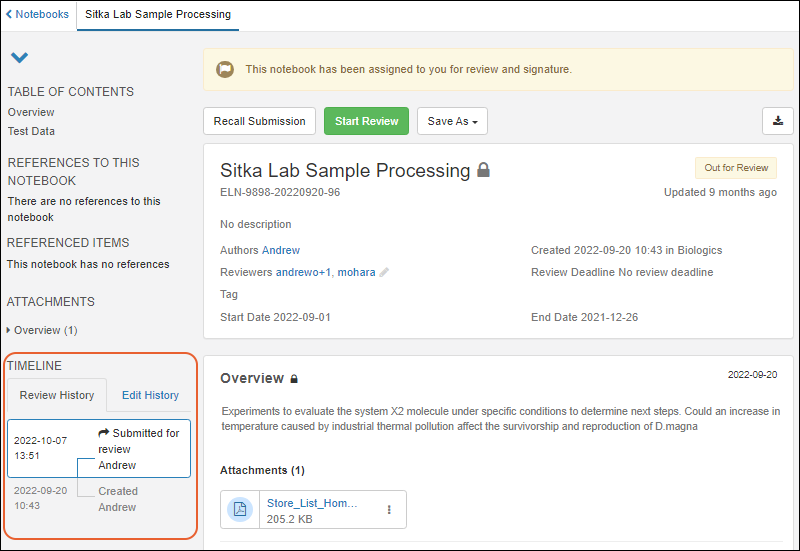 Review history events will include submission for review, return for comments, resubmission, and approval. When a notebook is amended, you'll also see events related to that update cycle.
Review history events will include submission for review, return for comments, resubmission, and approval. When a notebook is amended, you'll also see events related to that update cycle.Related Topics
Notebook Amendments
Amend a Notebook
To amend a signed notebook, open it and click Amend this Notebook.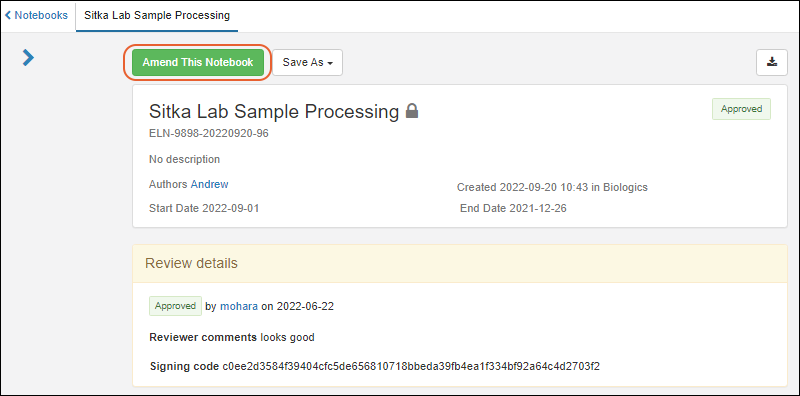 A new amendment revision will be created, giving you the option to edit, add, and correct what you need. While amending, you have the option to Restore Last Approved Version if you change your mind.
A new amendment revision will be created, giving you the option to edit, add, and correct what you need. While amending, you have the option to Restore Last Approved Version if you change your mind.Submit Amendment for Review
When finished making changes, you'll Submit for Review of the amended version, following the same signature process as when you originally submitted it. On the signature page, a Reason for amendment is always required. Click Submit Signed Notebook when ready.Timeline
All edit and review activities are tracked in a Timeline section of the notebook at the bottom of the expandable details panel. You'll see amendments on the Review History tab:Related Topics
Notebook Templates
- Create New Template
- Create Template from Notebook
- Create Template from Another Template
- Create Notebook from Template
- Create Notebook from Another Notebook
- Manage Templates
Create New Template
To create a new template from scratch, click Templates on the main menu under Notebooks. You can also select Manage > Templates from the Notebooks Dashboard.Click Create Template.Shared Templates
To share your template with your team, i.e. everyone with permission to create new notebooks, click Shared.Create Template from Notebook
You can also create a new template from an existing notebook. The template will include entry content, attachments, fields, and references for repeat use in other notebooks.Open the notebook, then select Save As > New Template.Create Template from Another Template
While editing a template, you can use the contents to create a new template by selecting Save As > New Template.Create Notebook from Template
To create a new notebook from a template, you have two options.Starting from the template, select Save As > New Notebook.Create a Notebook from Another Notebook
You can create a new notebook from an existing notebook by selecting Save As > Copy. Copying a notebook will include the value of any custom fields, where using a template does not preserve those values.Manage Templates
Open the Template Dashboard by selecting Templates from the main menu. You can also select Manage > Templates from the Notebook Dashboard.- Your Templates lists the templates you have created, whether they are shared with the team or not.
- Shared Templates lists templates created by anyone and shared with the team.
Archive Templates
Rather than fully delete a template, you have the option to archive it, meaning that it is no longer usable for new notebooks. You can select the rows for one or more templates on the Manage dashboard, then click Archive. You can also open any template and select Archive > Archive Template.Related Topics
Export Notebook
Export Notebook as PDF
To export a notebook as a PDF, you must have correctly configured the Puppeteer service.Once configured, you will see a button in your notebooks. Click and confirm that Notebook PDF is selected in the popup.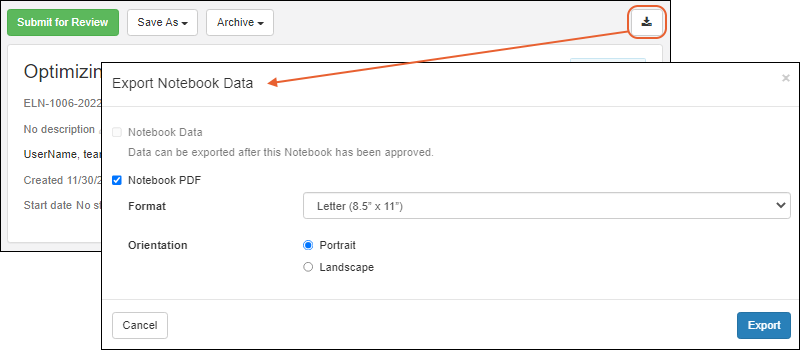 Adjust settings including:
Adjust settings including:
- Format: Letter or A4.
- Orientation: Portrait or Landscape.
- The exported document includes a panels of details about the notebook, including Title, Status, ID, Authors, Creation Date, and Project.
- The header on every page of the document includes the notebook title, ID, approval status, and author name(s).
Export Submitted/Signed Notebook
Once a notebook has been submitted for review, the exported PDF will include a full review and signing history as of the time of export.- Out for review: Shows who submitted it and when.
- Returned: Includes who reviewed it and when, as well as the return comments.
- Approved: The final review status panel will include a full history of submitting, reviewing, and end with who signed the Notebook and when these events occurred.
- A footer on every page includes when the document was printed, and once the notebook is approved, this footer repeats the details of when and by whom it was signed and witnessed.
Export Approved/Signed Notebook Archive
Once a notebook has been approved (i.e. signed by reviewers), you'll be able to export the data in an archive format so that you can store it outside the system and refer later to the contents.[notebook ID].export.zip
[notebook ID]_[approval date]_[approval time].notebook-snapshot.zip
[notebook ID]_[approval date]_[approval time].notebook-snapshot.zip
│
├───summary
│ └───[Notebook Title]([Notebook ID]).tsv Notebook properties and values
│
└───referenced data
├───assay
│ └───[Assay Name]([Assay ID]).tsv Details for referenced assay runs
│
├───sample
│ ├───[Sample Type1].tsv Details for any referenced samples
│ └───[Sample Type2].tsv of each type included
│
└───more folders as needed for other referenced items
Related Topics
Configure Puppeteer
Puppeteer Service Overview
The puppeteer-service is a standalone docker container web service that allows for generation of assets (e.g. PDFs, screenshots) via Puppeteer. Learn about deploying this service below. Note that the puppeteer-service can be stood up once and shared by several LabKey Server instances (e.g. staging, production, etc.).The puppeteer module is part of LabKey Server that communicates with this service to generate PDFs from Notebooks. It is run in the remote service mode to communicate with the service using the remote URL where it is deployed.Deploy the Puppeteer Service
Deployment
The puppeteer-service can be deployed in a Docker container on either a Linux or OSX host. Deploying this service on a Windows host is not supported nor tested.Docker is required to be installed on the host machine. Retrieve the latest Docker image from Docker Hub using the following command:docker pull labkeyteamcity/lk-puppeteer-service
docker run -i --init --rm --cap-add=SYS_ADMIN -p 3031:3031 --name puppeteer-service labkeyteamcity/lk-puppeteer-service
sudo docker run -di --init --cap-add=SYS_ADMIN -p--restart unless-stopped --name puppeteer-service labkeyteamcity/lk-puppeteer-service
Interface
POST to /pdf:- inputURL: The URL of the website to create a PDF or screenshot from.
- apiKey: (Optional) If visiting a LabKey site that requires credentials you can specify an apiKey.
- fileName: (Optional) Name of the file that is streamed back in the response. Defaults to generated.pdf.
- printFormat: (Optional) Currently supports Letter and A4 page sizes. Defaults to Letter.
- printOrientation: (Optional) Supports Portrait and Landscape orientations. Defaults to Portrait.
Configuration Properties
Docker Port The Docker Port property specifies what host port the service container should be bound to.Remote URL
The Remote URL where the web service can be found should be provided to whomever will now configure LabKey Server to connect to the puppeteer-service.Configure Puppeteer Module
- Select > Site > Admin Console.
- Under Configuration, click Puppeteer Service.
- Enable the service.
- Select Service Mode: Remote
- Provide the Remote URL, including the port.
- Click Save.
Startup Properties
All of the puppeteer module properties are also available as LabKey Server Startup Properties. An example properties file would look something like:puppeteer.enabled;startup=true
puppeteer.mode;startup=remote
puppeteer.docker.port;startup=3031
puppeteer.docker.remapLocalhost;startup=true
puppeteer.remote.url;startup=http://localhost:3031
Related Topics
Troubleshoot ELN Images (Guide)
You can add images to an Electronic Lab Notebook (ELN) in two main ways:
-
Attach as a file – Upload the image so it appears as an attachment.
-
Embed in the document – Paste the image so it appears directly in your ELN entry.
This guide focuses on embedding images.
Method 1: Copy and Paste from Your Computer (Recommended)
Best for: Preserving the exact file format and quality.
Steps:
-
Download the image to your computer.
-
Open your file browser (Finder on macOS, File Explorer on Windows).
-
Right-click the image file and select Copy
-
or use CTRL + C (Windows) / CMD + C (macOS).
-
-
In your ELN, click where you want the image to appear.
-
Right-click and select Paste
-
or use CTRL + V (Windows) / CMD + V (macOS).
-
Tip: This method avoids any format conversions that can happen when copying from a browser.
Method 2: Copy and Paste from a Browser
Best for: Quickly adding images from a webpage.
Steps:
-
Find the image in your browser.
-
Right-click the image and select Copy Image.
-
Paste into your ELN as described above.
Note: Most browsers (especially Chrome-based ones) will convert the image to PNG format before pasting. If you need the original format, use Method 1.
Method 3: Copy HTML Content (Websites or Other Apps)
Best for: Copying both text and images from a webpage, Word document, or other application.
Steps:
-
Highlight the content you want to copy (text and images).
-
Right-click and select Copy.
-
Paste into your ELN.
Common Errors and Fixes
When images from non-LabKey sites are blocked, you may see an error like this:
<insert example image of an error after pasting an image>
Case 1 — Image blocked by CSP
If only the image-src directive is enabled, images may display temporarily but remain linked to the external URL. If the source image is deleted, it will disappear from your ELN. In this case, you may also see errors such as:
<insert example image of an error after an image is pasted with image-src directive but no connect-src directive>
For administrators:
-
Allow images from the source domain by adding both:
-
image-srcdirective -
connect-srcdirective
-
-
Go to Admin Console → Allowed External Hosts to configure.
Case 2 — External site blocks image download
Even with the correct CSP settings, some sources use CORS restrictions to prevent downloading.
Best workaround for users: Download the image to your computer and paste it using Method 1.
Best Practice Summary
-
For reliable results – Always use Method 1 (from your computer).
-
To avoid broken links – Ensure images are embedded, not just linked.
-
Admins – Configure both
image-srcandconnect-srcdirectives when allowing external images.
Troubleshooting ELN Images
Copying and Pasting Images from the Filesystem
The most reliable method is to download the image to your computer, locate it in your file browser, and copy it (Right-click → Copy or use Ctrl+C on Windows / Cmd+C on macOS).Next, place your cursor in the ELN entry where you want the image, and paste it (Right-click → Paste, or Ctrl+V / Cmd+V).This approach inserts an exact copy of the image into the ELN.Copying and Pasting Images from a Browser
You can also right-click an image in your browser and select Copy Image, then paste it into your ELN entry as described above.This method is reliable, but note that most browsers will convert the image to PNG format before pasting. If preserving the exact file format is important, use the filesystem method instead.Copying and Pasting HTML Content from External Sources
When copying content from a website or another application (such as Microsoft Word), images may be included along with the HTML content. These pasted images often reference an external URL (for example, a CDN).Because LabKey Server enforces a strict Content Security Policy (CSP) by default, images from third-party sources are usually blocked. Pasting such content may produce errors, for example:
Recommendation
For the most consistent results, especially when working with external content, download images to your computer and paste them into the ELN from the filesystem. This ensures the image is embedded and preserved within the ELN entry.ELN: Frequently Asked Questions
How can I use Notebooks for my work?
Notebooks are available in the Biologics and Sample Manager applications.- Already using Biologics? You'll already have Notebooks, provided you are on a current version.
- Already using Sample Manager? You'll need to be using the Professional Edition of Sample Manager, or the Enterprise Edition of LabKey Server to access Notebooks.
- Not using either of the above? Please contact us to learn more.
Who can see my Notebooks?
Every user with "Read" access to your folder can also see Notebooks created there.What can I reference from a Notebook?
You can reference anything in your application, including data, experiments, specific samples, and other notebooks. Use a direct reference selector to place a color coded reference directly in your Notebook text. Add a single reference at a time, or add multiple references in bulk.The combined list of all elements referenced from a notebook is maintained in an Overview panel.Learn more in this topic: Add a ReferenceHow are Notebooks locked and protected once signed?
After the notebook has been approved we create a signed snapshot. The snapshot will include all notebook data including:- Notebook metadata and text
- Any data referenced in the notebook (samples, entities, experiments, assay runs, etc.)
- Attached files
What happens to my Notebooks when I upgrade?
- Once you create a Notebook, it will be preserved (and unaltered) by upgrades of LabKey.
What are team templates?
"Team templates" is a term that was in use in earlier versions for templates that are now called Shared Templates.When you view the Notebook Templates Dashboard, all the templates you created are listed on the Your Templates tab. Templates created by yourself or other users and shared (i.e. team templates) are listed on the Shared Templates tab.Related Topics
Data Resources
Topics
Use Sample Manager with LabKey Server
- Sample Manager in a Project
- Using Sample Manager in a Folder
- Export and Import of Sample Manager
- Sample Manager User Interface
- Shared Sample Types and Source Types
- Sample Management Users and Permissions
- Use Naming Prefixes
- Configure Storage Location Options
- Storage Management API
Sample Manager in a Project
Sample Manager is designed to be contained in a project on LabKey Server. This provides advantages in scoping permissions specific to sample management. For instance, many studies in other projects may need to be linked to sample data, but a single group doing the actual sample administration and storage management for all studies would have elevated permissions on the Sample Manager folder instead of needing them on all studies.Resources including Sample Types and Assay Designs that should be usable in both the Sample Manager project and other containers on the server should be placed in the Shared project.- Create a new project, selecting the folder type Sample Manager.
- Configure permissions now, or leave it visible to only your own user account until you have set things up, then open permissions to your users.
Using Sample Manager in a Folder
Sample Manager is designed to be rooted at the top-level project level of LabKey Server. In some limited use cases, it may work to use Sample Manager in a subfolder, however not all features are supported. If you do not expect to share any Sample Types, Assays, or other data with other containers, and will only use a single set of container-based permissions, using Sample Manager in a subfolder may support integration with LabKey Server features and reports. Note that you cannot use Sample Manager Folders unless it is rooted at the project level.If you choose either of the following subfolder options, you will be able to use the product selection menu to switch between Sample Manager and the traditional LabKey Server interface.Folder of "Sample Manager" Type
You can create a new folder and select the "Sample Manager" folder type. The application will launch every time you navigate to this folder, as if it were at the project level.Enable "SampleManagement" Module
If you want to use Sample Manager resources in a folder, but do not want to auto-navigate to the application, you can create a folder of another type (such as Study) and then enable the "SampleManagement" module via > Folder > Management > Folder Type.You will not be navigated directly to the application in this folder, but can still reach it by editing the URL to replace "project-begin.view?" with "sampleManager-app.view?".Export and Import of Sample Manager
Sample Manager projects and folders do not support full fidelity migration to other containers using folder export and reimport available in the LabKey Server interface. Some of the contents of Sample Manager can be migrated in this way, in some cases requiring manually migrating some data in a specific order. Others, including but not limited to Workflow and Notebook contents, cannot be moved in this way.If you need to migrate any Sample Manager data or structures between containers, please work with your Account Manager to identify whether this is feasible in your scenario. Note that it is never possible to "promote" a folder to the project level within LabKey Server.Sample Manager User Interface
When using Sample Manager with a Premium Edition of LabKey Server, you may see different features and options when viewing the "same" data in the two different interfaces. A few examples of differences are listed here, but this is not a comprehensive list.Learn about switching between the interfaces in this topic:Grid Charts are Available
When using Sample Manager with a Premium Edition of LabKey Server, you can define and use chart visualizations within the application. Learn more about charts in this topic for Biologics LIMS:Conditional Formatting is not Available
Conditional formatting of values is not supported in Sample Manager, though you will see the controls for adding such formats in the field editor.In order to use conditional formatting, you will need to define the formats and view your data in the LabKey Server Interface.Shared Sample Types and Source Types
Any Sample Types and Sources defined in the /Shared project will also be available for use in LabKey Sample Manager. You will see them listed on the menu and dashboards alongside local definitions.From within the Sample Manager application, users editing (or deleting) a shared Sample Type or Source Type will see a banner indicating that changes could affect other folders.Sample Management Users and Permission Roles
An administrator in the Sample Manager or LabKey Biologics applications can access a grid of active users using the Administration option on the user avatar menu. You'll see all the users with any access to the container in which the application is enabled.Sample Manager and Biologics both use a subset of LabKey's role-based permissions. The "Reader", "Editor", "Editor without Delete", "Folder Administrator", and "Project Administrator" roles all map to the corresponding container-scoped roles. Learn more about those permissions in this topic: Permission Roles. In addition, "Storage Editor" and "Storage Designer" roles are added to the Sample Manager and Biologics applications and described below.Reader, Editor, and Editor without Delete
- When a user is granted the "Reader", "Editor", or "Editor without Delete" role, either in the LabKey Server folder permissions interface or the Sample Manager > Administration > Permissions tab, they will have that role in both interfaces.
- Assigning a user to a role in either place, or revoking that role in either place will apply to both the Sample Manager and LabKey Server resources in that container.
- If Sample Manager is defined in a folder, and that folder inherits permissions from a parent container, you will not be able to change role assignments within the application.
Administrator Roles
In the stand-alone Sample Manager or application, a single "Administrator" role is provided that maps to the site role "Application Admin". This also means that any "Application Admin" on the site will appear in Sample Manager as an Administrator. The Sample Manager Documentation means these site-level roles when it identifies tasks available to an "Administrator".When using Sample Manager within LabKey Server, administrator permissions work differently. Users with the "Folder Administrator" or "Project Administrator" role on the project or folder containing Sample Manager have those roles carry into the application. These roles may also be assigned from within the application (by users with sufficient permission). The difference between these two roles in Sample Manager is that a Project Administrator can add new user accounts, but a Folder Administrator cannot. Both admin roles can assign the various roles to existing users and perform other administrative tasks, with some exceptions listed below.As in any LabKey Server Installation, "Site Admin" and "Application Admin" roles are assigned at the site level and grant administrative permission to Sample Manager, but are not shown in the Administrator listing of the application.Most actions are available to any administrator role in Sample Manager with some exceptions including:- Only "Site Admin" and "Application Admin" users can manage sample statuses.
Specific Roles for Storage Management
Two roles specific to managing storage of physical samples let Sample Manager and LabKey Biologics administrators independently grant users control over the physical storage details for samples.- Storage Editor: confers the ability to read, add, edit, and delete data related to items in storage, picklists, and jobs.
- Storage Designer: confers the ability to read, add, edit, and delete data related to storage locations and storage units.
Workflow Editor Role
The Workflow Editor role lets a user create and edit sample picklists as well as workflow jobs and tasks, though not workflow templates. Workflow editors also require the "Reader" role or higher in order to accomplish these tasks.Administrators can also perform the tasks available to users with the Workflow Editor role.Use Naming Prefixes
You can apply a container-specific naming prefix that will be added to naming patterns for all Sample Types and Source Types to assist integration of data from multiple locations while maintaining a clear association with the original source of that data.When you have more than one Sample Manager project or folder, or are using both Sample Manager and LabKey Biologics on the same LabKey Server, it can be helpful to assign unique prefixes to each container.Learn more about using prefixes here:Configure Storage Location Options
Freezers and other storage systems can be assigned physical locations to make it easier for users to find them in a larger institution or campus. Learn more about defining and using storage locations in this topic: Note that this is different from configuring location hierarchies within freezers or other storage systems.Storage Management API
In situations where you have a large number of storage systems to define, it may be more convenient to define them programmatically instead of using the UI. Learn more in the API documentation:- Java Storage API
- JavaScript Storage API
- Python Storage API
- Rlabkey Package API: See labkey.storage.create/delete/update.
Related Topics
Use Sample Manager with Studies
Include Participant and Visit Information in Sample Types
Fields of specific types can be included in a Sample Type, providing participant (Subject/Participant type) and visit information (either VisitDate or VisitID/VisitLabel depending on the timepoint type of the target study). These fields can be defined either:- On the type of the samples you want to link
- On the type of a parent sample of the samples you want to link.
Link Samples to Studies from Sample Manager
When the Sample Manager application is used on a Premium Edition of LabKey Server, you gain the ability to link samples to studies directly from within the application.From a Samples grid, select the samples of interest using the checkboxes, then select Edit > Link to LabKey Study.- You must have the necessary alignment fields defined.
- The name of your Sample Type cannot be the same as the name of any table or dataset in the study you are linking to.
Related Topics
Data Grid Basics
- Grid Menus
- Work with Data Grids
- Data Entry and Edit Grids
- Export
Column Header Menus
Click the column header in a grid to access the column header menu:- Filter: Click to add or edit filters; by default the current column is selected in the popup.
- Remove filter: Remove the filter on this column.
- Sort ascending
- Sort descending
- Clear sort: Remove the sort on this column.
- Edit Label: Edit the display label for this column.
- Insert Column: Select a new column to add to the grid.
- Hide Column: Hide this column from the grid.
Column Headers: Field Name and Tooltip
Hover over the column header in a grid to see the full field name. In the case of different labels, or ancestry lookups (or both shown here), it can be helpful to see the actual underlying name of the field:Scroll Large Grids
When scrolling a grid horizontally, the leftmost column, typically the SampleID, will remain 'locked' or visible on the left, making it easy to understand which sample the visible fields are referring to.Sample Grid Menus
Sample grid menus highlight the most common sample actions and group them by category:- Add:
- Add Manually
- Import from File: New samples only.
- Edit:
- Edit in Grid
- Edit in Bulk
- Update from File: Update existing samples or merge both new and existing samples from one spreadsheet.
- Edit Lineage
- Move to Folder (Only available when Folders are in use.)
- Delete
- Link to LabKey Study (Only available when Sample Manager is used as part of a Premium Edition of LabKey Server)
- Derive:
- Assay:
- Picklists:
- Jobs:
- Storage:
- Reports:
- Find Derivatives in Sample Finder: Open the Sample Finder using any filters applied to the current grid as search criteria for finding derived/child samples.
- View Assay Results for Selected: Create a comprehensive report of all assay results for a set of samples.
Select Rows
In LabKey Sample Manager, data is shown in grids, with a column of checkboxes on the left for selecting each individual row. Check the box in the header row to select all rows on the current page.Once you've selected a page of rows, you will see buttons to select all the rows, or clear those already checked.Page Through Data
Large sets of data are displayed on a paged grid. In the upper right, you see which rows you are viewing (here 1-20 of 824). Buttons give you the following control:- and : Step one page forward or back.
- The page number you are on is shown with a dropdown menu.
- You can jump to the first page or last page and see a count for the total number of pages.
- You can also change the pagination. Options for number of rows per page: 20, 40, 100, 250, 400.
Filter
Use the Filter option on any column header menu, or click the button above the grid to open the filter panel.- Filter settings for a grid will be maintained if you click away and later return to the same grid.
- When you hover over a lozenge, the filter icon will become an X you can click to delete that filter.
- Click Remove all to remove all the filters.
Sort by Column Value
In each header, click the to sort the grid by the values in that column. Select either ascending or descending sort.- Sort settings for a grid will be maintained if you click away and later return.
Search Grid
Enter your search terms in the Search box above the grid to search the text fields in the grid. Click the X to clear the search terms.Multi-Tabbed Sample Grids
When a grid could contain samples of multiple types, such as on a picklist or the View All Samples grid, you'll see a separate tab for each Sample Type as well as an All Samples tab showing only properties common to all types, including the Sample ID, Status, and Storage information. There is a limited set of actions available on this tab that can be used for samples of multiple types, as shown in the first image below. Each individual sample type tab has the full set of grid actions, as shown in the second image.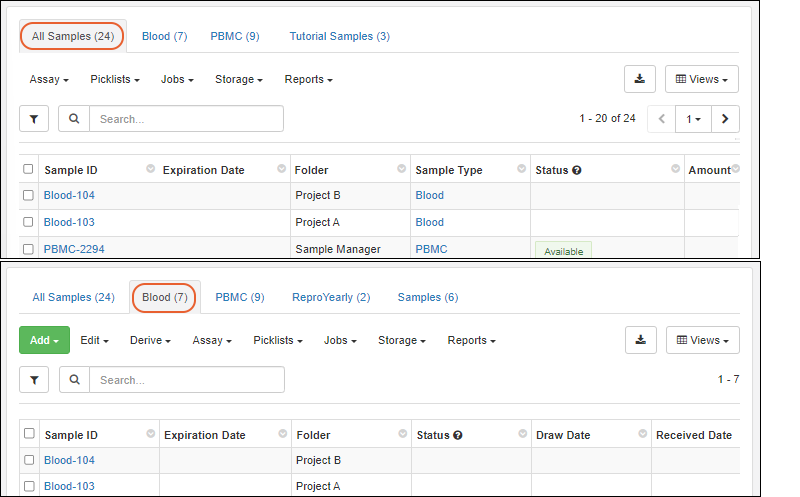 When there is only one type of sample in the grid, it will open on the specific Sample Type tab. When there are several, it will open on the All Samples tab, as shown above.
When there is only one type of sample in the grid, it will open on the specific Sample Type tab. When there are several, it will open on the All Samples tab, as shown above.
Editable Grids
Editable grids can be found throughout the application for entering and editing data for Samples, Sources, Assays, etc. See an example of using editable grids here.When using an editable grid, you can make use of the following options:- When a field cannot be edited, such as the Folder field shown below, it will be shown grayed-out.
- For a Sample or Source grid, if user-defined names are allowed, the Sample ID (or Source ID) values are editable in a grid.
- All rows in the grid must be from folders where user-defined names are allowed, otherwise IDs will be read-only and grayed-out.
- Note that you cannot "swap" the names of two samples in the grid, as only new unique names are allowed. To make a swap, use a temporary intermediate name.
- Any field that offers a selection menu (Sample Status, Text Choice, list lookups), will be shown with a .
- Type ahead to narrow the choices and click or tab to select the highlighted value.
- For example, the Status field in the image below shows an open menu.
- Hover over the for the Status column to see a legend of available statuses.
- Some validation, such as confirming data is of the expected type, will be performed as you enter values, giving you an immediate indication of errors.
- When a Sample or Source field has identifying fields set, they will be shown in the dropdowns. An exception is that when editing Assay Results, the additional fields for Samples will only be shown when the assay is set to map to a specific Sample Type.
- Fields that support multi-select, like sources and parents, will show any existing selection as well as allow you to select more from the dropdown.
- The Tutorial Labs field below already contains one value, and you can add an additional value as appropriate.
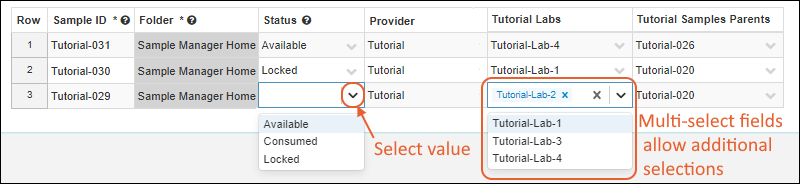 Editable grids provide "locking" of the sample ID column and column headers, so that it's easier to tell which cell is currently being edited.
Editable grids provide "locking" of the sample ID column and column headers, so that it's easier to tell which cell is currently being edited.
Drag to Populate Columns
Entering a first value (or row of values), then using the 'drag handle' to apply them to multiple rows is a convenient way to populate a grid.For a Text field, if you type some text in one field, then select it and grab the "cell-drag handle" in the lower right of the cell, you can drag to repeat it in the remaining cells.Cut/Paste to Duplicate Cells
Pasting from a grid of cells, such as 2x2, into a 2x2 area will copy the grid as expected. Pasting into a larger area, such as 4x8, will replicate the pasted grid in the other cells as shown below. The target area may (but does not need to) include the originally selected cells.Export Data
To export the data in a grid, click the (Export) button and select the format for export:- CSV
- Excel: Learn about exporting a multi-tabbed grid below.
- TSV
- If BarTender label printing is configured, you can also export and print labels and templates from this menu.
Multi-Tab Excel Exports
When you export from a grid that contains multiple tabs, such as one containing samples of different types as shown below, the exported Excel file can also include multiple tabs (sheets). Select > Excel, then in the popup, you will see the per-tab Count and which View will be used for each tab. Check the boxes for the tabs you want included in the export and click Export.Storage Map Exports
When viewing samples in a storage location, you can also export a Storage Map for sharing or offline use. Learn more here:Related Topics
Data Import Guidelines
- Use Import Templates
- Background Import
- Import Performance Considerations
- Batch Delete Limitations
- Data Structure Names
- Column/Field Names
- Data Preview Considerations
- Reserved Fields
- Import to Unrecognized Fields
- Migration of Inventory/Storage Amount Fields
- Amount/Units Display Details
- Import to Samples and Sources via API
Use Import Templates
For the most reliable method of importing data, first obtain a template for the data you are importing. You can then ensure that your data conforms to expectations before using either Add > Import from File or Edit > Update from File.For Source Types, Sample Types, and Assay Results, click the category from the main menu. You'll see a Template button for each data structure defined.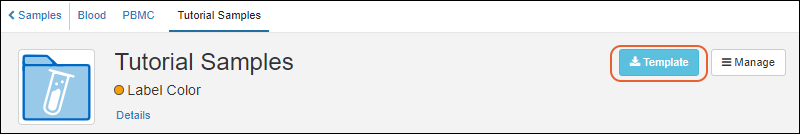 In case you did not already obtain a template, you can also download one from within the file import interface itself:
In case you did not already obtain a template, you can also download one from within the file import interface itself:- For a Sample Type, if you have defined Parent or Source aliases, all the possible columns will be included in the template, but only the ones you are using need to be included.
- In cases of columns that cannot be edited directly (such as the Storage Status of a sample, which is defined by a sample having a location and not being checked out), these columns will be omitted from the template.
- Note that the template for assay designs includes the results columns, but not the run or batch ones.
Learn more about LIMS features here.
Background Import (Asynchronous Import)
When import by file is large enough that it will take considerable time to complete, the import will automatically be done in the background. Files larger than 100kb will be imported asynchronously. This allows users to continue working within the app while the import completes.Import larger files as usual. You will see a banner message indicating the background import in progress, and a icon alongside that sample type until it completes: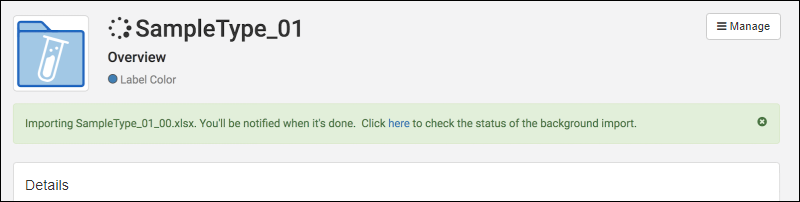 Any user in the application will see the spinner in the header bar. To see the status of all asynchronous imports in progress, select > View all activity (this menu may be a spinner when imports are in progress).
Any user in the application will see the spinner in the header bar. To see the status of all asynchronous imports in progress, select > View all activity (this menu may be a spinner when imports are in progress).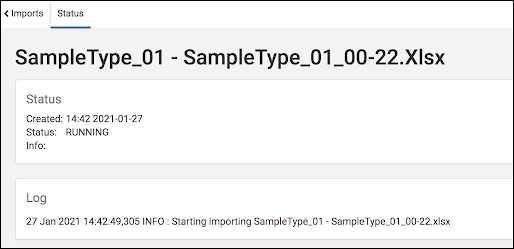 When the import is complete, you will receive an in-app notification via the menu.
When the import is complete, you will receive an in-app notification via the menu.
Import Performance Considerations
Excel files containing formulas will take longer to upload than files without formulas.The performance of importing data into any structure is related to the number of columns. If your sample type or assay design has more than 30 columns, you may encounter performance issues.Batch Delete Limitations
You can only delete 10,000 rows at a time. To delete larger sets of sample or assay data, select batches of rows to delete.Data Structure Names
Data structures (domains) like Sample Types, Source Types, Assay Designs, etc. must have unique names and avoid specific special characters, particularly if they are to be used in naming patterns or API calls. Names must follow these rules:- Must not be blank
- Must start with a letter or a number character.
- Must contain only valid unicode characters. (no control characters)
- May not contain any of these characters:
<>[]{};,`"~!@#$%^*=|?\ - May not contain 'tab', 'new line', or 'return' characters.
- May not contain space followed by dash followed by a character.
- i.e. these are allowed: "a - b" or "a-b" or "a–-b"
- these are not allowed: "a -b", "a –-b"
AliquotedFrom
~DataInputs
DataInputs
Inputs
~MaterialInputs
MaterialInputs
batchRandomId
containerPath
contextPath
sampleCount
rootSampleCount
dailySampleCount
dataRegionName
genId
monthlySampleCount
now
queryName
randomId
schemaName
schemaPath
selectionKey
weeklySampleCount
withCounter
yearlySampleCount
folderPrefix
:passThrough
:htmlEncode
:jsString
:urlEncode
:encodeURIComponent
:encodeURI
:first
:rest
:last
:trim
:date
:dailySampleCount
:weeklySampleCount
:yearlySampleCount
:monthlySampleCount
:defaultValue
:minValue
:number
:prefix
:suffix
:join
:withCounter
- Must not contain any of the following characters:
/:<>$[]{};,`"~!@#$%^*=|?\
Column/Field Names
When you create a column (field) with a special character like a space, slash, or other characters, you will see a warning in the UI. These warnings do not prevent you from saving, but instead of spaces or special characters, try renaming data columns to use CamelCasing or '_' underscores as word separators instead of special characters. Displayed column headers will parse the internal caps and underscores to show spaces in the column names.For any field name, you can also change the Label for the field (under Name and Linking Options in the field editor)) if desired to provide a longer name or name with special characters in it. For example, if you want to display a column with units included, you could import the data with a field name of platelets and then set the label to show "Platelets (per uL)" to the user.You can also use Import Aliases to map a column name that contains spaces to a sample type or assay field that does not. Remember to use "double quotes" around names that include spaces. For example, for "Platelets (per uL)", you would define your assay with a field named "platelets" and include "Platelets (per uL)" (including the quotes) in the Import Aliases box of the assay design definition (in addition to the label, if desired).Data Preview Considerations
Previewing data stored as a TSV or CSV file may be faster than previewing data imported as an Excel file, particularly when file sizes are large.Previewing Excel files that include formulas will take longer to preview than similar Excel files without formulas.Reserved Fields
There are a number of reserved field names used within LabKey for every data structure that will be populated internally when data is created or modified, or are otherwise reserved and cannot be redefined by the user:- Created
- CreatedBy
- Modified
- ModifiedBy
- RowId
- LSID
- Folder
- Properties
| Sample Type | Source Type |
|---|---|
| Name | Name |
| SampleId | SourceId |
| Description | Description |
| SampleState ("Status") | |
| MaterialExpDate ("Expiration Date") | |
| Flag | Flag |
| SourceProtocolApplication | |
| SourceApplicationInput | |
| RunApplication | |
| RunApplicationOutput | |
| Protocol | Protocol |
| Alias | Alias |
| SampleSet | DataClass |
| ClassId | |
| Run | |
| genId | genId |
| Inputs | Inputs |
| Outputs | Outputs |
| DataFileUrl | |
| QueryableInputs | |
| SampleCount | |
| StoredAmount ("Amount") | StoredAmount ("Amount") |
| Units (units associated with the StoredAmount field) | |
| SampleTypeUnits (units associated with the Sample Type) | |
| RawAmount: See below | |
| RawUnits: See below | |
| FreezeThawCount | |
| StorageStatus | |
| StorageLocation | |
| StorageRow | |
| StorageCol | |
| CheckedOutBy | |
| CheckedOut (Date) | |
| IsAliquot | |
| AliquotCount ("Aliquots Created Count") | |
| AliquotTotalVolume |
Inferral of Reserved Fields
If you infer a data structure from a file, and it contains any reserved fields, they will not be shown in the inferral but will be created for you. You will see a banner informing you that this has occurred:Import to Unrecognized Fields
If you import data that contains fields unrecognized by the system for that data structure (sample type, source type, or assay design), you will see a banner warning you that the field will be ignored:Migration of Inventory Fields
In version 23.4, some fields from the inventory schema have been migrated and renamed. If you happen to be using the new names in your system as well, this migration can cause conflicts. It is recommended that you keep these field name changes in mind. If you are using any fields listed below for your own purposes, you should rename them prior to upgrading:| Old Field | Action Taken | New Field |
|---|---|---|
| inventory.item.volume | migrated (with existing data) | exp.materials.StoredAmount |
| inventory.item.volumeUnits | migrated (with existing data) | exp.material.Units |
| inventory.item.initialVolume | removed |
Amount/Units Display Details
The StoredAmount column is labeled "Amount". When importing data via a file, the column header in the file may be named either "Amount" or "StoredAmount". Either will be mapped to the Amount/StoredAmount field.On import, the Amount and Unit values are stored in the database as follows:- The "base unit" of the sample type is determined:
- If the Display unit is set to a mass unit, then the base unit is "g".
- If the Display unit is set to a volume unit, then the base unit is "mL".
- If the Display unit is set to unit, then the base unit is "unit".
- The amount and unit values supplied by the user are converted to this "base unit" for storage in the database, either g, mL, or unit. For example, if a user enters 1 L, and the Display Unit is configured as mL, then the system will save the unit in the database as mL.
- When displayed, the values are converted from the database base unit to the display unit configured in the sample type.
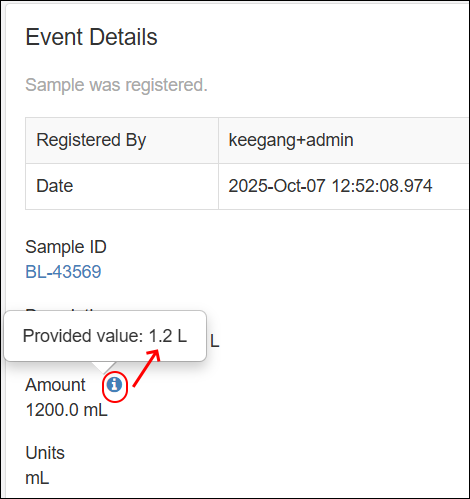
Import of Samples and Sources via API
Sample Types and Sources are similar, with a few key differences. Sources are "data classes". Upload source data to the "exp.data" schema.Sample Types are defined in the "exp" experiment schema, and some access to data will be through the "exp.materials" schema. However, all sample data should be uploaded to the "samples" schema.Related Topics
Custom Grid Views
- Customize Grid View
- Include Sorts and Filters in Custom Views
- Include Ancestor Information
- Save Grid View
- Use Saved Views
- Edit Saved Views
- Manage Saved Views
Customize Grid View
Quickly see data that pertains to you with custom saved grid views. Create a custom view of your data by rearranging, hiding or showing columns, adding filters or sorting data. With saved custom views, you can view your data in multiple ways depending on what’s useful to you.You can quickly customize a grid view directly from the column header menus, or open the grid view customizer by selecting Grid Views > Customize Grid View to make many changes at once.Change Column Order
Drag and drop column headers directly in the grid view, or open Views > Customize Grid View and using the sixblock handles to change the column order in the Shown in Grid panel.Insert Column
Select Insert Column from any column header, or open Views > Customize Grid View. Click the in the Available Fields panel to add a new column to the grid.Drag and drop the columns to the desired order.Hide Column
Select Hide Column from any column header to hide that column, or open Views > Customize Grid View and click one or more 'X's for fields you want to delete (from the Shown in grid panel.Edit Label
Edit the display label for a column. You can either select Edit Label from the column header and type the new label directly:Or click the for the field within the view customizer and type the new label.Include Sorts and Filters in Custom Views
When you save a grid view, any filters or sorts currently applied will be saved. Make the filtering and sorting adjustments you prefer prior to saving.Include Ancestor Information
To include information about ancestors, use the Ancestor node in the grid view customizer to find specific types and properties of sample ancestors. Learn more in this topic:Save Grid View
Once you've made changes, either directly using column headers or using the grid view customizer, you'll see a header banner on your grid view indicating that it has been editing and inviting you to:- Undo: revert to the default.
- Save your changes.
- Enter a Grid View Name in the box.
- If you have the Editor role (or higher) you can share your grid with other users. Check the box to Make this grid view available to all users. Otherwise your named grid views will be visible only to you.
- Select whether to Make this grid view available in all Folders.
- Click Save again.
Administrator Options for Saving Grid Views
For administrators, the options available are:- Save as default view for all users: Only available to administrators and their default.
- Save as a custom view: As for any user, select this option to save a named custom grid view. When selected, you'll see the same options to give the grid a name, and select whether to share with other users and/or make it available in all Folders.
- Click Save.
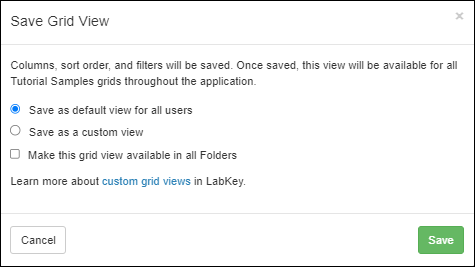
Use Saved Views
By saving a set of customized named views, you can create your own menu giving you quick access to whatever standardized groupings and details are relevant to your specific role. You might have different views filtering samples by attributes or storage location, and another showing only the shipping details needed for sets of samples.In addition to Your Saved Views, you'll see Shared Saved Views that were created by other users listed separately.Access both types of saved grids from the Views menu:Edit Saved Views
To edit a saved view, you simply select it from the menu, then make changes you want. The name of the grid you are viewing is shown in the header, and once you've made changes you'll see an "Edited" indicator and Undo and Save buttons. Editing a named view, you can also use the > Save as... option to save as a new grid view.Manage Saved Views
Select Views > Manage Saved Views for a popup listing them. You can:- (Edit)
- (Delete)
- Administrators will be also see options to:
- Revert an edited Default view to the system default.
- Make default: Make an existing named view the Default view for all users.
Related Topics
Identifying Fields
- Add Identifying Fields
- Use Identifying Fields
- Sample/Source Selection
- Storage
- Editable Grids
- Grids for Creating Derivatives or Aliquots
- Assay Results (Professional Edition Feature)
- Notebook References (Professional Edition Feature)
- Edit Identifying Fields
Add Identifying Fields
- Click the name of the Sample Type or Source Type on the main menu.
- Select Manage > Edit Identifying Fields.
- The Sample ID (or Source ID) is always selected and cannot be deleted.
- Using an interface very similar to the grid view customizer, add up to three more identifying fields by clicking the respective . Note that fields of type 'Calculation' cannot be added as identifying fields.
- Click Update to save.
Use Identifying Fields
Sample/Source Selection
When Identifying Fields are set, you will see these values in the selection boxes when choosing from dropdowns. In this example, the user can avoid choosing a sample with a similar name that is already "Consumed".Storage
You will see the values of identifying fields when you add samples to storage or move them, provided the samples are all of the same type.Editable Grids
When editing samples or sources in grids, including when editing lineage, you'll see any identifying fields displayed in non-editable columns alongside the ID.Grids for Creating Derivatives, or Aliquots
When deriving new samples or creating aliquots, the values for any identifying fields will be included in tooltips for selected parent/source sample in the creation grid, as for sample creation.If any identifying fields are from the Ancestors node in the grid customizer, those ancestor details will be shown in the grid as non-editable columns, giving detail to the user about the sample being derived from or aliquoted. For example, in the screenshot below DNA samples are being derived from parent Blood samples. The fields circled in red show data about the parent blood samples, because the DNA sample type has identifying fields that reference Ancestor fields in the Blood table.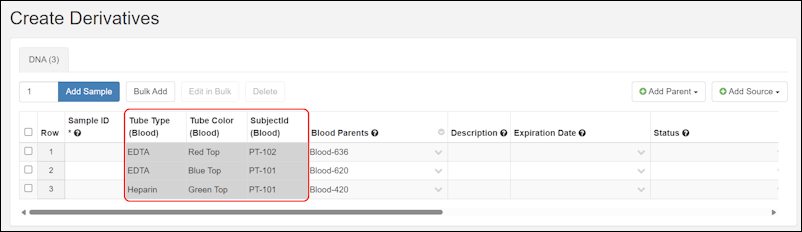 Note that non-ancestor identifying fields are not shown in this way as it would create ambiguity in the editable grid.
Note that non-ancestor identifying fields are not shown in this way as it would create ambiguity in the editable grid.
Assay Results (Professional Edition Feature)
When adding (or editing) assay results in a grid, identifying fields are shown when the assay design looks up to an specific Sample Type. As each row is populated with a SampleID, the values of the identifying fields will be shown for reference.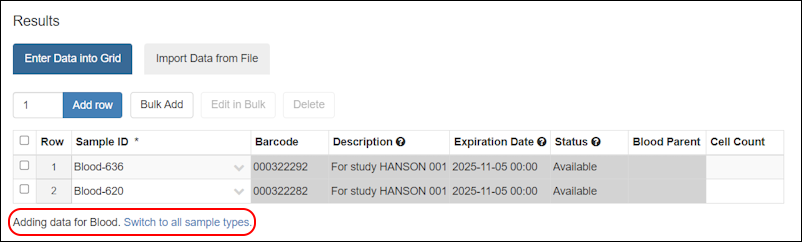
Notebook References (Professional Edition Feature)
When a Notebook contains a reference to a sample or source with identifying fields, these values will be shown first when you hover over the reference lozenge.Edit Identifying Fields
To change or remove identifying fields, reopen the same Edit Identifying Fields UI where they were set and click the to delete them from the Identifying Fields side of the panel. Click Update to save.Related Topics
Audit History
- View Audit History
- Available Audit Logs
- Customize Audit Views
- Require Users Provide Reasons for Certain Actions (Professional Feature)
View Audit History
Select > Audit Logs.Available Audit Logs
Use the selection menu near the top of the page to see a full listing of other logs available to administrators, including:- Assay Events: Run import, deletion, and reimport.
- Comments entered when assay data is deleted will be shown in the User Comment column.
- Note that for reimport, two assay events are created: one for the 'old' run being replaced and one for the 'new' run representing the new import.
- Attachment Events
- Data Update Events: When a row is updated, the log will show the details of what changed in that row.
- Domain Events: Tracks changes to columns in definitions (domains) of sample types, sources, and assays.
- Domain Property Events: Changes to the properties of a column in a domain.
- File Events: Records changes related to fields of type File, including uploading, updating, and moving files. When the server renames uploaded files to avoid duplicate file names, the original user-provided file name appears in the "Provided" column and the renamed file name appears in the "File" column.
- Folder Events: Folder editing events, including selective data exclusion for folders.
- List Events
- Notebook Events
- Notebook Review Events
- Query Update Events
- Roles and Assignment Events
- Sample Timeline Events: Records events for all samples.
- Individual sample timelines show all events for each sample.
- Detailed events for merge updates to sample data will show only the fields that were updated.
- Sample registration, update, storage changes, and check in/out actions are all logged.
- Comments entered when Samples are deleted will be shown in the User Comment column.
- Sample Type Events: Creation and modification of Sample Types.
- Sample Workflow Events: Job and task completion are recorded.
- Sources Events: Including creation of Source Types and registration of Sources.
- Comments entered when Sources are deleted will be shown in the User Comment column.
- Storage Management Events: See View Storage Activity.
- User Events: Creation of users; logging in and out.
Customize Audit Views
Use the grid view customizer to change the columns shown, labels, and order, as well as apply filters and sorts to give you the specific view of each type of audit log that you need. You can either save as a personal named view, share the view with other administrators, or change the default view all administrators will see.View Transactions
Add the column Transaction ID to show links to transactional, multi-part events. These links navigate to a tabbed view of the transactional audit events.View Method Used to Insert, Update, or Delete Records
The transaction details page provides details on the method used to modify records.For in-app data changes, see the Edit Method property.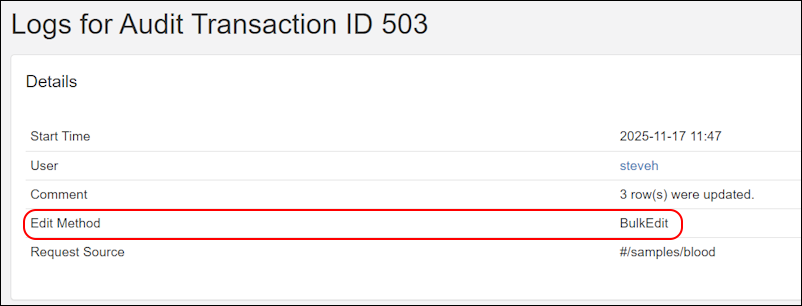 Possible values are:
Possible values are:
- GridInsert - Indicates that the record was inserted using the application grid.
- FormInsert - Indicates that the record was inserted using the application form.
- GridEdit - Indicates that the record was updated using the application grid.
- BulkEdit - Indicates that the record was updated using the bulk edit wizard.
- DetailEdit - Indicates that the record was updated using the details form.
- BulkEditLineage - Indicates that the lineage was updated using the bulk edit wizard.
- DetailEditLineage - Indicates that the lineage was updated using the details form.
- StorageViewAction - Indicates that the record was updated using the storage view.
- RequestSource: The URL where the request was made. For delete actions, URLs including a sample ID indicate deletion from a details page, otherwise, the delete occurred from a grid or API action.
- ImportFileName: File name used to import or update data.
- ImportOptions: Options that were selected during import or update:
- Cross Folder Import: a container field was provided.
- Cross Type Import: "Multiple sample types" was selected.
- IMPORT/UPDATE/MERGE: Whether it was an import, update, or merge operation.
- Background import: Whether the import was handled using a background job.
- Allow Create Storage: Creation of storage during sample import.
- Client Library: Indicates the client library used to perform the action(s).
- ETL: Includes the name of the ETL used for import.
- FileWatcher: Description of the file watcher, consisting of the name of the trigger and the filename that triggered the job.
Require Users Provide Reasons for Certain Actions (Professional Feature)
The following actions allow users to enter a reason (aka a "comment") for the action, which will be recorded in the audit log:- Editing samples individually from a sample details page, in a grid, in bulk, or via update from file
- Editing sources individually from a source details page, in a grid, in bulk, or via update from file
- Editing lineage
- Editing assay run or result data in a grid, in bulk, or via re-import
- Editing sample types, source types, and assay designs.
- Deleting samples and sample types
- Deleting sources and source types
- Deleting assay runs and assay designs
- Deleting jobs
- Deleting storage
- Updating sample amount or freeze/thaw count from a sample details page or on the sample details panel from the storage view
- Checking samples in or out of storage
- Removing samples from storage
- Moving data between folders
- Recalling a notebook submitted for review
- Select > Application Settings.
- Scroll down to the Audit Logging section.
- Select Yes, reasons are required.
Related Topics
Search
Search Basics
To search for samples, assays, and more, type the search terms in the box in the header of the application, or when the browser is narrow, the option will be on the search dropdown menu.Content Searched
- Sample Types (Name and description)
- Samples (SampleID and description)
- Storage Units, such as boxes (Name and Label)
- Source Types (Name and description)
- Sources (SourceID and description)
- Assay Designs (Name and description)
- Jobs (Name and description)
- Notebooks (Name, number, and contents)
Search for Storage Units
The Name and Label field for each storage unit are indexed so that you can easily search later for any particular storage. Instead of using the default/generic naming, customize the names of your storage or include any helpful text in the label (such as a barcode) that helps identify storage.When you are searching later for a specific storage unit, you can find it by terms in the name or label.Related Topics
Field Editor
- Create New Fields
- Edit, Rearrange, and Delete Fields
- Add/Edit Field Properties
- Sample Fields: Editable for Samples, Aliquots or Both
- Name and Linking Options
- Conditional Formatting and Validation Options
- Advanced Settings
- View Fields in Summary Mode
- Export Sets of Fields (Domains)
Create New Fields
Import or Infer Fields from File
When creating a sample type, source type, or assay design, you'll have the option to:- import fields from a specially prepared JSON file, OR
- infer them from an example data spreadsheet matching the structure of your data.
Manually Define Fields
To use the Field Editor to create a new set of fields and their properties manually, click Manually Define Fields.- Open the field editor.
- Click Manually Define Fields (or get started by importing or inferring fields, which will prepopulate the manual editor).
- To define a new field, click Add Field.
- Give the field a Name. Field names can contain a combination of letters, numbers, and underscores, should not contain spaces (or other special characters), and should start with a letter or underscore.
- Use the menu to select the Data Type. The set of data types available may vary with your configuration and each has a different set of properties you can set. Once you have saved fields, you can only make limited changes to the type.
- You can use the checkbox if you want to make it required that that field have a value in every row.
- Continue to add any new fields you need - one for each column of your data.
- Click the Finish Creating.../Save... button to save and exit the editor.
Edit Fields
To edit fields, reopen the editor and make the changes you need. If you attempt to navigate away with unsaved changes you will have the opportunity to save or discard them. When you are finished making changes, click Save.Once you have saved a field or set of fields, you can change the name and most options and other settings. However, you can only make limited changes to the type of a field. For example, you can change among text types, but cannot change a text field into a number or a boolean.Rearrange Fields
To change field order, drag and drop the rows using the six-block handle on the left.Delete Fields
To one or many fields, select them using the checkboxes and click Delete. You can use the checkbox at the top of the column to select all fields in the section.Save Fields
Click Save when finished.Add/Edit Field Properties
Each field can have additional properties defined. The properties available vary based on the field type. To open the properties for a field, click the icon on the right (it will become a handle for closing the panel).Fields of different types include some or all of these sections:- Sample Fields Only: Editable for Samples, Aliquots or Both
- Type-specific options for formatting values: See types in the field properties reference.
- Text and integer fields can be marked as barcode fields.
- Name and Linking Options
- Validation Options
Sample Fields: Editable for Samples, Aliquots or Both
Fields in Sample Type definitions can specify whether they should be settable/editable for Samples, Aliquots, or both.- Under Sample/Aliquot Options, select one:
- Editable for samples only (default): Aliquots will inherit the value of the field from the sample.
- Editable for aliquots only: Samples will not display this field, but it will be included for aliquots.
- Separately editable for samples and aliquots: Both samples and aliquots can set a different value for this property. Note that if you change an existing Sample Type field from "Editable for samples only" to this "Separately editable for samples and aliquots" option, any stored values for aliquots will be dropped.
Name and Linking Options
All types of fields allow you to set the following properties:- Description: An optional text description. This will appear in the hover text for the field you define.
- Label: Different text to display in column headers for the field. This label may contain spaces. The default label is the Field Name with camelCasing indicating separate words. For example, the field "firstName" would by default be labelled "First Name".
- Import Aliases: Define alternate field names to be used when importing from a file to this field. Multiple aliases may be separated by spaces or commas. To define an alias that contains spaces, use double-quotes (") around it.
- URL: Use this property to change the display of the field value within a data grid into a link. Multiple formats are supported, which allow ways to easily substitute and link to other locations in LabKey. Learn more about using URL Formatting Options.
- Open links in a new tab: When clicked, links will open a new browser tab.
Conditional Formatting and Validation Options
Conditional Formatting is available for most fields when using the LabKey LIMS and Biologics LIMS applications, and when Sample Manager is used with a Premium Edition of LabKey Server. Conditional formatting will display values with color or text highlighting in grids and detail views when those values meet certain criteria. Learn more in this topic: Validation Options are offered on most fields. String-based fields offer regular expression validation. Numeric, date, and user fields offer range expression validation.Create Regular Expression Validator
- Click Add Regex to open the popup.
- If you don't see this option, it is not supported for your field type.
- If any regex validators are defined, you'll also see a link showing the number active and the button will read Edit Regex.
- Enter the Regular Expression that this field's value will be evaluated against. All regular expressions must be compatible with Java regular expressions as implemented in the Pattern class.
- Description: Optional description.
- Error Message: Enter the error message to be shown to the user when the value fails this validation.
- Check the box for Fail validation when pattern matches field value in order to reverse the validation: With this box unchecked (the default) the pattern must match the expression. With this box checked, the pattern may not match.
- Name: Enter a name to identify this validator.
- You can use Add Validator to add a second condition. The first panel will close and show the validator name you gave. You can reopen that panel using the (pencil) icon.
- Click Apply when your regex validators for this field are complete.
- Click Save or Finish in the editor.
Create Range Expression Validator
- Click Add Range to open the popup.
- If you don't see this option, it is not supported for your field type.
- If any range validators are defined, you'll also see a link showing the number active and the button will read Edit Ranges.
- Enter the First Condition that this field's value will be evaluated against. Select a comparison operator and enter a value.
- Optionally enter a Second Condition.
- Description: Optional description.
- Error Message: Enter the error message to be shown to the user when the value fails this validation.
- Name: Enter a name to identify this validator.
- You can use Add Validator to add a second condition. The first panel will close and show the validator name you gave. You can reopen that panel using the (pencil) icon.
- Click Apply when your range validators for this field are complete.
- Click Save or Finish in the editor.
Advanced Settings
When using Sample Manager with a Premium Edition of LabKey Server, you may see additional options in the field editor that are not covered in this topic. For more information about these options, please see the companion topics in the LabKey Server documentation:View Fields in Summary Mode
In the upper right, choose the Mode:- Detail: Fields can be expanded to edit properties.
- Summary: Only a summary of fields are shown; limited editing is available.
Export Sets of Fields (Domains)
Once you have defined a set of fields (domain) that you want to be able to save or reuse, you can export it by clicking (Export).If you want to only export a subset of the fields included, use the selection checkboxes to select the fields to export.- When any (or all) boxes are checked, only the checked fields are exported.
- If no boxes are checked, all fields will be exported.
 A Fields_*.fields.json file describing your fields as a set of key/value pairs will be downloaded. All properties that can be set for a field in the user interface will be included in the exported file contents.You can use this file as a template to generate a set of field definitions for import elsewhere.
A Fields_*.fields.json file describing your fields as a set of key/value pairs will be downloaded. All properties that can be set for a field in the user interface will be included in the exported file contents.You can use this file as a template to generate a set of field definitions for import elsewhere.Related Topics
- Field Properties Reference: Type specific formatting and advanced options.
- Barcode Fields
Field Properties Reference
Type Specific Formatting Options
Text Options
- Text Options:
- Maximum Text Length. Sets the maximum character count for the field. Choose either "Unlimited" or "No longer than X characters", providing a value in the box. The default is 4000.
- Barcode Field: Check the box to include this field when searching samples by barcode.
- Name and Linking Options
- Validation Options: Regular expression validators.
Text Choice Options
A Text Choice field lets you define a set of values that will be presented to the user as a dropdown list. For example, you might offer a "Tube Type" field and let the user choose Heparin, EDTA, or Unknown.- Text Choice Options:
- Add and manage the set of drop-down values offered for this field, as shown below.
- Name and Linking Options
- Up to 200 values can be included in the drop-down options. Values can be single- or multi-word.
- You can change an existing Text field to the Text Choice type; the list of drop down values will be populated for you.
- Values that are in use cannot be deleted. If they are in use by read-only data, they can neither be edited nor deleted.
- All changes to the set of values for a text choice field are audited.
Multi-Line Text and Flag Options
- Multi-line Text Field Options (or Flag Options):
- Maximum Text Length. Sets the maximum character count for the field. Choose either "Unlimited" or "No longer than X characters", providing a value in the box. The default is 4000.
- Name and Linking Options
- Validation Options: Regular expression validators.
Boolean Options
- Boolean Field Options: Format for Boolean Values: Use boolean formatting to specify the text to show when a value is true and false. Text can optionally be shown for null values. For example, "Yes;No;Blank" would output "Yes" if the value is true, "No" if false, and "Blank" for a null value.
- Name and Linking Options
Integer/Decimal Options
- Integer Options/Decimal Options:
- Format for Numbers: To control how a number value is displayed, provide a string format compatible with the Java class DecimalFormat. Learn more about using Number formats in LabKey.
- Default Scale Type: Choose Linear or Log.
- Name and Linking Options
- Validation Options: Range validators.
Date, Time, and Date Time Options
For samples, sources, assay results and assay runs, three different field types are available, letting you choose how best to represent the data needed. Other types of fields in Sample Manager only support the DateTime combined field.- Date Time: Both date and time are included in the field. Fields of this type can be changed to either "Date-only" or "Time-only" fields, though this change will drop the data in the other part of the stored value.
- Date: Only the date is included. Fields of this type can be changed to be "Date Time" fields.
- Time: Only the time portion is represented. Fields of this type cannot be changed to be either "Date" or "Date Time" fields.
- Date and Time Options:
- Format for Dates: To control how a date, time or date/time value is displayed, select one of the available formats.
- Name and Linking Options
- Validation Options: Range validators are available for "Date Time" and "Date" fields, but not for "Time" fields.
Calculation Options (Available in the Professional Edition)
A calculation field lets you include SQL expressions using values in other fields in the same row to provide calculated values. The Expression provided must be valid LabKey SQL and can use the default system fields, custom fields, constants, and operators. To use field names containing special characters in a calculated field, surround the name with double quotes. String constants use single quotes. Examples:| Operation | Example |
|---|---|
| Addition | numericField1 + numericField2 |
| Subtraction | numericField1 - numericField2 |
| Multiplication | numericField1 * numericField2 |
| Division by value known never to be zero | numericField1 / nonZeroField1 |
| Division by value that might be zero | CASE WHEN numericField2 <> 0 THEN (numericField1 / numericField2 * 100) ELSE NULL END |
| Subtraction of dates/datetimes (ex: difference in days) | TIMESTAMPDIFF('SQL_TSI_DAY', CURDATE(), MaterialExpDate) |
| Relative date (ex: 1 week later) | TIMESTAMPADD('SQL_TSI_WEEK', 1, dateField) |
| Conditional calculation based on another field | CASE WHEN FreezeThawCount < 2 THEN 'Viable' ELSE 'Questionable' END |
| Conditional calculation based on a text match | CASE WHEN ColorField = 'Blue' THEN 'Abnormal' ELSE 'Normal' END |
| Text value for every row (ex: to use with a URL property) | 'clickMe' |
| Text concatenation (use fields and/or strings) | City || ', ' || State |
| Addition when field name includes special characters | "Numeric Field Name" + "Field/Name & More" |
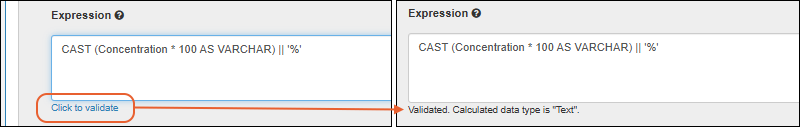
File/Attachment Options
Sources, samples, and assay designs (run and batch fields) support including file attachments, known in different structures as either File or Attachment fields.- Source Types use Attachment
- Sample Types use File
- Assay Designs allow File fields in batch and run field sections
- "File" values are stored in the filesystem and "Attachment" values are stored as blobs in the database.
- Learn more about the differences in the main LabKey Server documentation here.
User Options
- Name and Linking Options
- Validation Options: Range validators.
Sample Options
- Sample Options: Select where to look up samples for this field.
- You can choose All Samples to reference any sample in the container, or select a specific sample type to filter by.
- This selection will be used to validate and link incoming data, populate lists for data entry, etc.
- Lookup Validator: Ensure Value Exists in Lookup Target. Check the box to require that any value is present in the target sample type (or in "all samples" if selected).
- Name and Linking Options
Unique ID Options
A field of type "Unique ID" is read-only and used to house barcode values generated by LabKey. Learn more in this topic: Barcode FieldsRelated Topics
- Field Editor
- Conditional Formatting: Available in LabKey LIMS, Biologics LIMS, and Sample Manager when used with a Premium Edition of LabKey Server.
- Fields for Integrating with LabKey Studies: Available when using Sample Manager on a Premium Edition of LabKey Server
Attach Images and Other Files
- Add File Field (Administrator)
- Upload File (Editor)
- View Thumbnails and Expand Images (Reader)
- Download File (Reader)
- Remove or Change the File (Editor)
- Missing Files
Add File Field (Administrator)
An administrator must add the field to the data structure. In Sample Types, the field is of type "File" and in Source Types, the field is of type "Attachment". In this example, the field is included in a Sample Type and named "Image".Upload File (Editor)
Now by editing the Details for any sample in the system, you will be able to upload a file by either clicking in the selection window or dragging and dropping from your desktop. Click the sample name on any grid to open the Overview tab. When you are editing sample details, you can click to select or drag and drop a file to upload it.View Thumbnails and Expand Images (Reader)
A small thumbnail and the filename will be shown in the details panel and in the column of sample grids. Click the thumbnail or filename to open the image in a larger window.Download File (Reader)
To download the file, select Download from the menu.Remove or Change the File (Editor)
To change the attached file, reopen the sample details, click to open the Details panel editor and select Remove file from the menu. This option is only available in edit mode.Missing Files
If a file is missing, whether because of an upload problem or later deletion, you will see a red warning symbol and hovertext will tell you the file is unavailable.Related Topics
URL Field Property
Substitution Syntax
You can use one or more field values as parameters when creating the URL link, using the ${ } substitution syntax. Put the name of the column whose value you want to use inside the braces. The field with the URL property defined on it is available, but so are any other fields in the data structure.For example, if your data includes a "GeneSymbol" field displaying values like "BRCA", you could link to related information in The Gene Ontology by using a search URL. When the user clicked the value "BRCA", they would go to:http://amigo.geneontology.org/amigo/search/ontology?q=BRCAIn this case the field value is passed as a search parameter, so to create the URL property on the GeneSymbol field, you would include the GeneSymbol field in the URL property definition:http://amigo.geneontology.org/amigo/search/ontology?q=${GeneSymbol} You could also define this URL property on a different field, letting you display the gene symbol unlinked and link a value in another field (like "Click to Search") to perform the search.Substitutions are allowed in any part of the URL, either in the main path, in the query string, or both. For example, here are two different formats for creating links to an article in wikipedia, here using a "CompanyName" field value:- as part of the path:
- as a parameter value:
Local Links
To link to content in the current LabKey folder, use the controller and action name. You can optionally prepend a / (slash) or ./ (dot-slash), but they are not necessary.<controller>-<action>
list-details.view?listId=5&pk=${Building}External Links
To link to a resource on an external server or any website, include the full URL link.http://server/path/page.html?id=${Param}URL Encoding Options
You can specify the type of URL encoding for a substitution marker, in case the default behavior doesn't work for the URLs needed. This flexibility makes it possible to have one column display the text and a second column can contain the entire href value, or only a part of the href.The fields referenced by the ${ } substitution markers might contain any sort of text, including special characters such as question marks, equal signs, and ampersands. If these values are copied straight into the link address, the resulting address would be interpreted incorrectly. To avoid this problem, LabKey Server encodes text values before copying them into the URL. In encoding, characters such as ? are replaced by their character code %3F. By default, LabKey encodes all special character values except '/' from substitution markers. If you know that a field referenced by a substitution marker needs no encoding (because it has already been encoded, perhaps) or needs different encoding rules, inside the ${ } syntax, you can specify encoding options as described in the topic String Expression Format Functions.Related Topics
Date, Time, and Number Formats
- Set DateTime, Date, and Time Display Formats
- DateTime, Date, and Time Display Options
- Number Format Strings
Set DateTime, Date, and Time Display Formats
An administrator can configure the way DateTime, Date, and Time field values are displayed in the application. This does not affect how values may be imported, but can be used to standardize or simplify the display to users.There are three separate format selectors available for dates, date-times, and time-only values, as shown in the image below. For each, you can check the box to Use Default or select the pattern you prefer.- Select > Application Settings.
- When the Use Default box is checked, you will be inheriting a default setting.
- Uncheck the box and choose a different Display format for date/date-times/time-only values if desired.
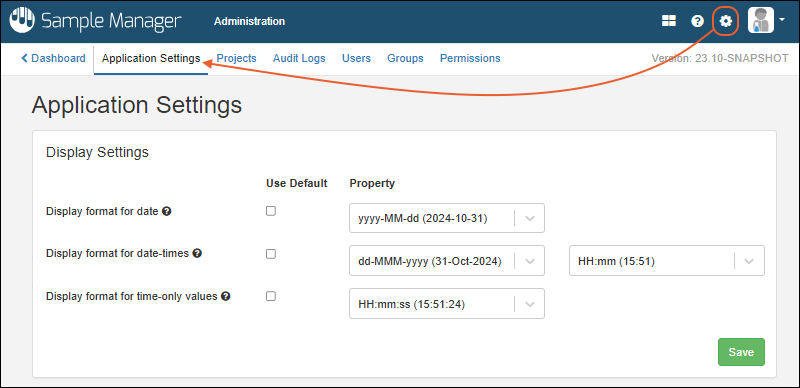
- Click Save.
DateTime, Date, and Time Display Options
Date, Time, and DateTime display formats are selected from a set of standard options, giving you flexibility for how users will see these values. DateTime fields combine one of each format, with the option of choosing "<none>" as the Time portion.Date formats available:| Format Selected | Display Example |
|---|---|
| yyyy-MM-dd | 2024-08-14 |
| yyyy-MMM-dd | 2024-Aug-14 |
| yyyy-MM | 2024-08 |
| dd-MM-yyyy | 14-08-2024 |
| dd-MMM-yyyy | 14-Aug-2024 |
| dd-MMM-yy | 14-Aug-24 |
| ddMMMyyyy | 14Aug2024 |
| ddMMMyy | 14Aug24 |
| MM/dd/yyyy | 08/14/2024 |
| MM-dd-yyyy | 08-14-2024 |
| MMMM dd yyyy | August 14 2024 |
| Format Selected | Display Example |
|---|---|
| HH:mm:ss | 13:45:15 |
| HH:mm | 13:45 |
| HH:mm:ss.SSS | 13:45:15.000 |
| hh:mm a | 01:45 PM |
Number Format Strings
Format strings for Integer and Decimal fields must be compatible with the format that the java class DecimalFormat accepts. A valid DecimalFormat is a pattern specifying a prefix, numeric part, and suffix. For more information see the Java documentation. The following table has an abbreviated guide to pattern symbols:| Symbol | Location | Localized? | Meaning |
|---|---|---|---|
| 0 | Number | Yes | Digit |
| # | Number | Yes | Digit, zero shows as absent |
| . | Number | Yes | Decimal separator or monetary decimal separator |
| - | Number | Yes | Minus sign |
| , | Number | Yes | Grouping separator |
Examples
The following examples apply to Decimal fields.| Format String | Display Result |
|---|---|
| <no string> | 85.0 |
| 0 | 85 |
| 000 | 085 |
| .00 | 85.00 |
| 000.000 | 085.000 |
| 000,000 | 085,000 |
| -000,000 | -085,000 |
Related Topics
String Expression Format Functions
Reference
The following string formatters can be used when building URLs, or creating unique Sample IDs/names.
| Name | Synonym | Input Type | Description | Example | |
|---|---|---|---|---|---|
| General | |||||
| defaultValue(string) | any | Use the string argument value as the replacement value if the token is not present or is the empty string. | ${field:defaultValue('missing')} | ||
| passThrough | none | any | Don't perform any formatting. | ${field:passThrough} | |
| URL Encoding | |||||
| encodeURI | uri | string | URL encode all special characters except ',/?:@&=+$#' like JavaScript encodeURI() | ${field:encodeURI} | |
| encodeURIComponent | uricomponent | string | URL uncode all special characters like JavaScript encodeURIComponent() | ${field:encodeURIComponent} | |
| htmlEncode | html | string | HTML encode | ${field:htmlEncode} | |
| jsString | string | Escape carrage return, linefeed, and <>"' characters and surround with a single quotes | ${field:jsString} | ||
| urlEncode | path | string | URL encode each path part preserving path separator | ${field:urlEncode} | |
| String | |||||
| join(string) | collection | Combine a collection of values together separated by the string argument | ${field:join('/'):encodeURI} | ||
| prefix(string) | string, collection | Prepend a string argument if the value is non-null and non-empty | ${field:prefix('-')} | ||
| suffix(string) | string, collection | Append a string argument if the value is non-null and non-empty | ${field:suffix('-')} | ||
| trim | string | Remove any leading or trailing whitespace | ${field:trim} | ||
| Date | |||||
| date(string) | date | Format a date using a format string or one of the constants from Java's DateTimeFormatter. If no format value is provided, the default format is 'BASIC_ISO_DATE' | ${field:date}, ${field:date('yyyy-MM-dd')} | ||
| Number | |||||
| number | format | Format a number using Java's DecimalFormat | ${field:number('0000')} | ||
| Array | |||||
| first | collection | Take the first value from a collection | ${field:first:defaultValue('X')} | ||
| rest | collection | Drop the first item from a collection | ${field:rest:join('_')} | ||
| last | collection | Drop all items from the collection except the last | ${field:last:suffix('!')} |
Examples
| Function | Applied to... | Result |
| ${Column1:defaultValue('MissingValue')} | null | MissingValue |
| ${Array1:join('/')} | [apple, orange, pear] | apple/orange/pear |
| ${Array1:first} | [apple, orange, pear] | apple |
| ${Array1:first:defaultValue('X')} | [(null), orange, pear] | X |
LabKey SQL Syntax
LabKey SQL
LabKey SQL is a SQL dialect that supports (1) most standard SQL functionality and (2) provides extended functionality that is unique to LabKey, including:
- Security. Before execution, all SQL queries are checked against the user's security roles/permissions.
- Lookup columns. Lookup columns use an intuitive syntax to access data in other tables to achieve what would normally require a JOIN statement. For example: "SomeTable.ForeignKey.FieldFromForeignTable" The special lookup column "Datasets" is injected into each study dataset and provides a syntax shortcut when joining the current dataset to another dataset that has compatible join keys. See example usage.
- Cross-folder querying. Queries can be scoped to folders broader than the current folder and can draw from tables in folders other than the current folder. See Cross-Folder Queries.
- Parameterized SQL statements. The PARAMETERS keyword lets you define parameters for a query. An associated API gives you control over the parameterized query from JavaScript code. See Parameterized SQL Queries.
- Pivot tables. The PIVOT...BY and PIVOT...IN expressions provide a syntax for creating pivot tables. See Pivot Queries.
- User-related functions. USERID() and ISMEMBEROF(groupid) let you control query visibility based on the user's group membership.
- Ontology-related functions. (Premium Feature) Access preferred terms and ontology concepts from SQL queries. See Ontology SQL.
- Lineage-related functions. (Premium Feature) Access ancestors and descendants of samples and data class entities. See Lineage SQL Queries.
- Annotations. Override some column metadata using SQL annotations. See Use SQL Annotations.
Reference Tables:
- Keywords
- Constants
- Operators
- Operator Order of Precedence
- Aggregate Functions - General
- SQL Functions - General
- General Syntax
Keywords
| Keyword | Description |
| AS |
Aliases can be explicitly named using the AS keyword. Note that the AS
keyword is optional: the following select clauses both create an alias
called "Name":
SELECT LCASE(FirstName) AS Name Implicit aliases are automatically generated for expressions in the SELECT list. In the query below, an output column named "Expression1" is automatically created for the expression "LCASE(FirstName)": SELECT LCASE(FirstName) |
| ASCENDING, ASC | Return ORDER BY results in ascending value order. See the ORDER BY section for troubleshooting notes. ORDER BY Weight ASC |
| CAST(AS) |
CAST(R.d AS VARCHAR)
Defined valid datatype keywords which can be used as cast/convert targets, and to what java.sql.Types name each keyword maps. Keywords are case-insensitive. BIGINT Examples: CAST(TimeCreated AS DATE) CAST(WEEK(i.date) as INTEGER) as WeekOfYear, Precision and scale are supported when casting to NUMERIC. Example: CAST($Num AS NUMERIC(10,2)) |
| DESCENDING, DESC | Return ORDER BY results in descending value order. See the ORDER BY section for troubleshooting notes. ORDER BY Weight DESC |
| DISTINCT | Return distinct, non duplicate, values. SELECT DISTINCT Country FROM Demographics |
| EXISTS | Returns a Boolean value based on a subquery. Returns TRUE if at least one
row is returned from the subquery. The following example returns any plasma samples which have been assayed with a score greater than 80%. Assume that ImmuneScores.Data.SpecimenId is a lookup field (aka foreign key) to Plasma.Name.; SELECT Plasma.Name FROM Plasma WHERE EXISTS (SELECT * FROM assay.General.ImmuneScores.Data WHERE SpecimenId = Plasma.Name AND ScorePercent > .8) |
| FALSE | |
| FROM |
The FROM clause in LabKey SQL must contain at least one table. It can also
contain JOINs to other tables. Commas are supported in the FROM clause: FROM TableA, TableB WHERE TableA.x = TableB.x Nested joins are supported in the FROM clause: FROM TableA LEFT JOIN (TableB INNER JOIN TableC ON ...) ON... To refer to tables in LabKey folders other than the current folder, see Cross-Folder Queries. |
| GROUP BY |
Used with aggregate functions to group the results. Defines the "for
each" or "per". The example below returns the number of records "for
each" participant:
SELECT ParticipantId, COUNT(Created) "Number of
Records" |
| HAVING | Used with aggregate functions to limit the results. The following
example returns participants with 10 or more records in the Physical Exam
table: SELECT ParticipantId, COUNT(Created) "Number of Records" FROM "Physical Exam" GROUP BY ParticipantId HAVING COUNT(Created) > 10 HAVING can be used without a GROUP BY clause, in which case all selected rows are treated as a single group for aggregation purposes. |
| JOIN, RIGHT JOIN, LEFT JOIN, FULL JOIN, CROSS JOIN |
Example: SELECT * FROM "Physical Exam" FULL JOIN "Lab Results" ON "Physical Exam".ParticipantId = "Lab Results".ParticipantId |
| LIMIT |
Limits the number or records returned by the query. The following
example returns the 10 most recent records:
SELECT * |
| NULLIF(A,B) | Returns NULL if A=B, otherwise returns A. |
| ORDER BY |
One option for sorting query results. It may produce unexpected results when dataregions or views also have sorting applied, or when using an expression in the ORDER BY clause, including an expression like table.columnName. If you can instead use a sort on the custom view or via, the API, those methods are preferred (see Troubleshooting note below).
For best ORDER BY results, be sure to a) SELECT the columns on which you are sorting, b) sort on the SELECT column, not on an expression. To sort on an expression, include the expression in the SELECT (hidden if desired) and sort by the alias of the expression. For example: SELECT A, B, A+B AS C @hidden ... ORDER BY C ...is preferable to: SELECT A, B ... ORDER BY A+B Use ORDER BY with LIMIT to improve performance: SELECT ParticipantID, Height_cm AS Height FROM "Physical Exam" ORDER BY Height DESC LIMIT 5 Troubleshooting: "Why is the ORDER BY clause not working as expected?" 1. Check to ensure you are sorting by a SELECT column (preferred) or an alias of an expression. Syntax like including the table name (i.e. ...ORDER BY table.columnName ASC) is an expression and should be aliased in the SELECT statement instead (i.e. SELECT table.columnName AS C ... ORDER BY C 2. When authoring queries in LabKey SQL, the query is typically processed as
a subquery within a parent query. This parent query may apply it's own
sorting overriding the ORDER BY clause in the subquery. This parent "view layer" provides
default behavior like pagination, lookups, etc. but may also unexpectedly apply an additional sort. |
| PARAMETERS |
Queries can declare parameters using the PARAMETERS keyword. Default values
data types are supported as shown below:
PARAMETERS (X INTEGER DEFAULT 37) Parameter names will override any unqualified table column with the same name. Use a table qualification to disambiguate. In the example below, R.X refers to the column while X refers to the parameter: PARAMETERS(X INTEGER DEFAULT 5) Supported data types for parameters are: BIGINT, BIT, CHAR, DECIMAL, DOUBLE, FLOAT, INTEGER, LONGVARCHAR, NUMERIC, REAL, SMALLINT, TIMESTAMP, TINYINT, VARCHAR Numeric parameters can include precision and scale: PARAMETERS($NUM NUMERIC(10,2)) Parameter values can be passed via JavaScript API calls to the query. For details see Parameterized SQL Queries. |
| PIVOT/PIVOT...BY/PIVOT...IN | Re-visualize a table by rotating or "pivoting" a portion of it, essentially promoting cell data to column headers. See Pivot Queries for details and examples. |
| SELECT |
SELECT queries are the only type of query that can currently be written in
LabKey SQL. Sub-selects are allowed both as an expression, and in the
FROM clause.
Aliases are automatically generated for expressions after SELECT. In the query below, an output column named "Expression1" is automatically generated for the expression "LCASE(FirstName)": SELECT LCASE(FirstName) FROM... |
| TRUE | |
| UNION, UNION ALL | The UNION clause is the same as standard SQL. LabKey SQL supports UNION in subqueries. |
| VALUES ... AS | A subset of VALUES syntax is supported. Generate a "constant table" by
providing a parenthesized list of expressions for each row in the table.
The lists must all have the same number of elements and corresponding
entries must have compatible data types. For example:
VALUES (1, 'one'), (2, 'two'), (3, 'three') AS t; You must provide the alias for the result ("AS t" in the above), aliasing column names is not supported. The column names will be 'column1', 'column2', etc. |
| WHERE | Filter the results for certain values. Example: SELECT * FROM "Physical Exam" WHERE YEAR(Date) = 2010 |
| WITH |
Define a "common table expression" which functions like a subquery or inline view table. Especially useful for recursive queries. Usage Notes: If there are UNION clauses that do not reference the common table expression (CTE) itself, the server interprets them as normal UNIONs. The first subclause of a UNION may not reference the CTE. The CTE may only be referenced once in a FROM clause or JOIN clauses within the UNION. There may be multiple CTEs defined in the WITH. Each may reference the previous CTEs in the WITH. No column specifications are allowed in the WITH (as some SQL versions allow). Exception Behavior: Testing indicates that PostgreSQL does not provide an exception to LabKey Server for a non-ending, recursive CTE query. This can cause the LabKey Server to wait indefinitely for the query to complete. A non-recursive example: WITH AllDemo AS
(
SELECT *
FROM "/Studies/Study A/".study.Demographics
UNION
SELECT *
FROM "/Studies/Study B/".study.Demographics
)
SELECT ParticipantId from AllDemo
A recursive example: In a table that holds parent/child information, this query returns all of the children and grandchildren (recursively down the generations), for a given "Source" parent. PARAMETERS
(
Source VARCHAR DEFAULT NULL
)
WITH Derivations AS
(
-- Anchor Query. User enters a 'Source' parent
SELECT Item, Parent
FROM Items
WHERE Parent = Source
UNION ALL
-- Recursive Query. Get the children, grandchildren, ... of the source parent
SELECT i.Item, i.Parent
FROM Items i INNER JOIN Derivations p
ON i.Parent = p.Item
)
SELECT * FROM Derivations
|
Constants
The following constant values can be used in LabKey SQL queries.
| Constant | Description |
| CAST('Infinity' AS DOUBLE) | Represents positive infinity. |
| CAST('-Infinity' AS DOUBLE) | Represents negative infinity. |
| CAST('NaN' AS DOUBLE) | Represents "Not a number". |
| TRUE | Boolean value. |
| FALSE | Boolean value. |
Operators
| Operator | Description |
| String Operators | Note that strings are delimited with single quotes. Double quotes are used for column and table names containing spaces. |
| || | String concatenation. For example: SELECT ParticipantId, City || ', ' || State AS CityOfOrigin FROM Demographics If any argument is null, the || operator will return a null string. To handle this, use COALESCE with an empty string as it's second argument, so that the other || arguments will be returned: City || ', ' || COALESCE (State, '') |
| LIKE | Pattern matching. The entire string must match the given pattern. Ex: LIKE 'W%'. |
| NOT LIKE | Negative pattern matching. Will return values that do not match a given pattern. Ex: NOT LIKE 'W%' |
| Arithmetic Operators | |
| + | Add |
| - | Subtract |
| * | Multiply |
| / | Divide |
| Comparison operators | |
| = | Equals |
| != | Does not equal |
| <> | Does not equal |
| > | Is greater than |
| < | Is less than |
| >= | Is greater than or equal to |
| <= | Is less than or equal to |
| IS NULL | Is NULL |
| IS NOT NULL | Is NOT NULL |
| BETWEEN | Between two values, inclusive. Values can be numbers, strings or dates. |
| IN | Example: WHERE City IN ('Seattle', 'Portland') |
| NOT IN | Example: WHERE City NOT IN ('Seattle', 'Portland') |
| Bitwise Operators | |
| & | Bitwise AND |
| | | Bitwise OR |
| ^ | Bitwise exclusive OR |
| Logical Operators | |
| AND | Logical AND |
| OR | Logical OR |
| NOT | Example: WHERE NOT Country='USA' |
Operator Order of Precedence
| Order of Precedence | Operators |
| 1 | - (unary) , + (unary), CASE |
| 2 | *, / (multiplication, division) |
| 3 | +, -, & (binary plus, binary
minus) |
| 4 | & (bitwise and) |
| 5 | ^ (bitwise xor) |
| 6 | | (bitwise or) |
| 7 | || (concatenation) |
| 8 | <, >, <=, >=, IN, NOT IN, BETWEEN, NOT BETWEEN, LIKE, NOT LIKE |
| 9 | =, IS, IS NOT, <>, != |
| 10 | NOT |
| 11 | AND |
| 12 | OR |
Aggregate Functions - General
| Function | Description |
| COUNT | The special syntax COUNT(*) is supported as of LabKey v9.2. |
| MIN | Minimum |
| MAX | Maximum |
| AVG | Average |
| SUM | Sum |
| GROUP_CONCAT | An aggregate function, much like MAX, MIN, AVG, COUNT, etc. It can be used
wherever the standard aggregate functions can be used, and is subject to the
same grouping rules. It will return a string value which is comma-separated list
of all of the values for that grouping. A custom separator, instead of the
default comma, can be specified. Learn more here.
The example below specifies a semi-colon as the separator: SELECT Participant, GROUP_CONCAT(DISTINCT Category, ';') AS CATEGORIES FROM SomeSchema.SomeTable To use a line-break as the separator, use the following: SELECT Participant, GROUP_CONCAT(DISTINCT Category, chr(10)) AS CATEGORIES FROM SomeSchema.SomeTable |
| stddev(expression) | Standard deviation |
| stddev_pop(expression) | Population standard deviation of the input values. |
| variance(expression) | Historical alias for var_samp. |
| var_pop(expression) | Population variance of the input values (square of the population standard deviation). |
| median(expression) | The 50th percentile of the values submitted. |
Aggregate Functions - PostgreSQL Only
| Function | Description |
| bool_and(expression) | Aggregates boolean values. Returns true if all values are true and false if any are false. |
| bool_or(expression) | Aggregates boolean values. Returns true if any values are true and false if all are false. |
| bit_and(expression) | Returns the bitwise AND of all non-null input values, or null if none. |
| bit_or(expression) | Returns the bitwise OR of all non-null input values, or null if none. |
| every(expression) | Equivalent to bool_and(). Returns true if all values are true and false if any are false. |
| corr(Y,X) | Correlation coefficient. |
| covar_pop(Y,X) | Population covariance. |
| covar_samp(Y,X) | Sample covariance. |
| regr_avgx(Y,X) | Average of the independent variable: (SUM(X)/N). |
| regr_avgy(Y,X) | Average of the dependent variable: (SUM(Y)/N). |
| regr_count(Y,X) | Number of non-null input rows. |
| regr_intercept(Y,X) | Y-intercept of the least-squares-fit linear equation determined by the (X,Y) pairs. |
| regr_r2(Y,X) | Square of the correlation coefficient. |
| regr_slope(Y,X) | Slope of the least-squares-fit linear equation determined by the (X,Y) pairs. |
| regr_sxx(Y,X) | Sum of squares of the independent variable. |
| regr_sxy(Y,X) | Sum of products of independent times dependent variable. |
| regr_syy(Y,X) | Sum of squares of the dependent variable. |
| stddev_samp(expression) | Sample standard deviation of the input values. |
| var_samp(expression) | Sample variance of the input values (square of the sample standard deviation). |
SQL Functions - General
Many of these functions are similar to standard SQL functions -- see the JBDC escape syntax documentation.
| Function | Description |
| abs(value) | Returns the absolute value. |
| acos(value) | Returns the arc cosine. |
| age(date1, date2) |
Supplies the difference in age between the two dates, calculated in years. |
| age(date1, date2, interval) |
The interval indicates the unit of age measurement, either SQL_TSI_MONTH or SQL_TSI_YEAR. |
| age_in_months(date1, date2) | Behavior is undefined if date2 is before date1. |
| age_in_years(date1, date2) | Behavior is undefined if date2 is before date1. |
| asin(value) | Returns the arc sine. |
| atan(value) | Returns the arc tangent. |
| atan2(value1, value2) | Returns the arctangent of the quotient of two values. |
| case |
CASE can be used to test various conditions and return various results based on the test. You can use either simple CASE or searched CASE syntax. In the following examples "value#" indicates a value to match against, where "test#" indicates a boolean expression to evaluate. In the "searched" syntax, the first test expression that evaluates to true will determine which result is returned. Note that the LabKey SQL parser sometimes requires the use of additional parentheses within the statement. Example: SELECT "StudentName", |
| ceiling(value) | Rounds the value up. |
| coalesce(value1,...,valueN) | Returns the first non-null value in the argument list. Use to set default values for display. |
| concat(value1,value2) | Concatenates two values. |
| contextPath() | Returns the context path starting with “/” (e.g. “/labkey”). Returns the empty string if there is no current context path. (Returns VARCHAR.) |
| cos(radians) | Returns the cosine. |
| cot(radians) | Returns the cotangent. |
| curdate() | Returns the current date. |
| curtime() | Returns the current time |
| dayofmonth(date) | Returns the day of the month (1-31) for a given date. |
| dayofweek(date) | Returns the day of the week (1-7) for a given date. (Sun=1 and Sat=7) |
| dayofyear(date) | Returns the day of the year (1-365) for a given date. |
| degrees(radians) | Returns degrees based on the given radians. |
| exp(n) | Returns Euler's number e raised to the nth power. e = 2.71828183 |
| floor(value) | Rounds down to the nearest integer. |
| folderName() | LabKey SQL extension function. Returns the name of the current folder, without beginning or trailing "/". (Returns VARCHAR.) |
| folderPath() | LabKey SQL extension function. Returns the current folder path (starts with “/”, but does not end with “/”). The root returns “/”. (Returns VARCHAR.) |
| greatest(a, b, c, ...) |
Returns the greatest value from the list expressions provided. Any number
of expressions may be used. The expressions must have the same data type,
which will also be the type of the result. The LEAST() function is similar,
but returns the smallest value from the list of expressions. GREATEST() and
LEAST() are not implemented for SAS databases.
When NULL values appear in the list of expressions, different database
implementations as follows: Example: SELECT greatest(score_1, score_2, score_3) As HIGH_SCORE |
| hour(time) | Returns the hour for a given date/time. |
| ifdefined(column_name) | IFDEFINED(NAME) allows queries to reference columns that may not be present on a table. Without using IFDEFINED(), LabKey will raise a SQL parse error if the column cannot be resolved. Using IFDEFINED(), a column that cannot be resolved is treated as a NULL value. The IFDEFINED() syntax is useful for writing queries over PIVOT queries or assay tables where columns may be added or removed by an administrator. |
| ifnull(testValue, defaultValue) | If testValue is null, returns the defaultValue. Example: IFNULL(Units,0) |
| isequal | LabKey SQL extension function. ISEQUAL(a,b) is equivalent to (a=b OR (a IS NULL AND b IS NULL)) |
| ismemberof(groupid) | LabKey SQL extension function. Returns true if the current user is a member of the specified group. |
| javaConstant(fieldName) | LabKey SQL extension function. Provides access to public static final variable values. For details see LabKey SQL Utility Functions. |
| lcase(string) | Convert all characters of a string to lower case. |
| least(a, b, c, ...) | Returns the smallest value from the list expressions provided. For more details, see greatest() above. |
| left(string, integer) | Returns the left side of the string, to the given number of characters. Example: SELECT LEFT('STRINGVALUE',3) returns 'STR' |
| length(string) | Returns the length of the given string. |
| locate(substring, string) locate(substring, string, startIndex) | Returns the location of the first occurrence of substring within string. startIndex provides a starting position to begin the search. |
| log(n) | Returns the natural logarithm of n. |
| log10(n) | Base base 10 logarithm on n. |
| lower(string) | Convert all characters of a string to lower case. |
| ltrim(string) | Trims white space characters from the left side of the string. For example: LTRIM(' Trim String') |
| minute(time) | Returns the minute value for the given time. |
| mod(dividend, divider) | Returns the remainder of the division of dividend by divider. |
| moduleProperty(module name, property name) |
LabKey SQL extension function. Returns a module property, based on the module and property names. For details see LabKey SQL Utility Functions. |
| month(date) | Returns the month value (1-12) of the given date. |
| monthname(date) | Return the month name of the given date. |
| now() | Returns the system date and time. |
| overlaps |
LabKey SQL extension
function. Supported only when PostgreSQL is installed as the
primary database. SELECT OVERLAPS (START1, END1, START2, END2) AS COLUMN1 FROM MYTABLE The LabKey SQL syntax above is translated into the following PostgreSQL syntax: SELECT (START1, END1) OVERLAPS (START2, END2) AS COLUMN1 FROM MYTABLE |
| pi() | Returns the value of pi;. |
| power(base, exponent) | Returns the base raised to the power of the exponent. For example, power(10,2) returns 100. |
| quarter(date) | Returns the yearly quarter for the given date where the 1st quarter = Jan 1-Mar 31, 2nd quarter = Apr 1-Jun 30, 3rd quarter = Jul 1-Sep 30, 4th quarter = Oct 1-Dec 31 |
| radians(degrees) | Returns the radians for the given degrees. |
| rand(), rand(seed) | Returns a random number between 0 and 1. |
| repeat(string, count) | Returns the string repeated the given number of times. SELECT REPEAT('Hello',2) returns 'HelloHello'. |
| round(value, precision) | Rounds the value to the specified number of decimal places. ROUND(43.3432,2) returns 43.34 |
| rtrim(string) | Trims white space characters from the right side of the string. For example: RTRIM('Trim String ') |
| second(time) | Returns the second value for the given time. |
| sign(value) | Returns the sign, positive or negative, for the given value. |
| sin(value) | Returns the sine for the given value. |
| startswith(string, prefix) | Tests to see if the string starts with the specified prefix. For example, STARTSWITH('12345','2') returns FALSE. |
| sqrt(value) | Returns the square root of the value. |
| substring(string, start, length) | Returns a portion of the string as specified by the start location (1-based) and length (number of characters). For example, substring('SomeString', 1,2) returns the string 'So'. |
| tan(value) |
Returns the tangent of the value. |
| timestampadd(interval, number_to_add, timestamp) |
Adds an interval to the given timestamp value. The interval value must be surrounded by quotes. Possible values for interval: SQL_TSI_FRAC_SECOND Example: TIMESTAMPADD('SQL_TSI_QUARTER', 1, "Physical Exam".date) AS NextExam |
| timestampdiff(interval, timestamp1, timestamp2) |
Finds the difference between two timestamp values at a specified interval. The interval must be surrounded by quotes. Example: TIMESTAMPDIFF('SQL_TSI_DAY', SpecimenEvent.StorageDate, SpecimenEvent.ShipDate) Note that PostgreSQL does not support the following intervals: SQL_TSI_FRAC_SECOND As a workaround, use the 'age' functions defined above. |
| truncate(numeric value, precision) | Truncates the numeric value to the precision
specified. This is an arithmetic truncation, not a string truncation. TRUNCATE(123.4567,1) returns 123.4 TRUNCATE(123.4567,2) returns 123.45 TRUNCATE(123.4567,-1) returns 120.0 May require an explict CAST into NUMERIC, as LabKey SQL does not check data types for function arguments. SELECT PhysicalExam.Temperature, TRUNCATE(CAST(Temperature AS NUMERIC),1) as truncTemperature FROM PhysicalExam |
| ucase(string), upper(string) | Converts all characters to upper case. |
| userid() | LabKey SQL extension function. Returns the userid, an integer, of the logged in user. |
| username() | LabKey SQL extension function. Returns the current user display name. VARCHAR |
| version() | LabKey SQL extension function. Returns the current schema version of the core module as a NUMERIC with four decimal places. For example: 20.0070 |
| week(date) | Returns the week value (1-52) of the given date. |
| year(date) | Return the year of the given date. Assuming the system date is March 4 2023, then YEAR(NOW()) return 2023. |
SQL Functions - PostgreSQL Specific
LabKey SQL supports the following PostgreSQL functions.
See the PostgreSQL
docs for usage details.
| PostgreSQL Function | Docs |
| ascii(value) | Returns the ASCII code of the first character of value. |
| btrim(value, trimchars) |
Removes characters in trimchars from the start and end of
string. trimchars defaults to white space. BTRIM(' trim ') returns TRIM BTRIM('abbatrimabtrimabba', 'ab') returns trimabtrim |
|
character_length(value), char_length(value) |
Returns the number of characters in value. |
| chr(integer_code) | Returns the character with the given integer_code. CHR(70) returns F |
| concat_ws(sep text, val1 "any" [, val2 "any" [,...]]) -> text |
Concatenates all but the first argument, with separators. The first argument is used as the separator string, and should not be NULL. Other NULL arguments are ignored. See the PostgreSQL docs. concat_ws(',', 'abcde', 2, NULL, 22) → abcde,2,22 |
| decode(text, format) |
See the PostgreSQL docs. |
| encode(binary, format) |
See the PostgreSQL docs. |
| is_distinct_from(a, b) OR is_not_distinct_from(a, b) |
Not equal (or equal), treating null like an ordinary value. |
| initcap(string) | Converts the first character of each separate word in string to uppercase and the rest to lowercase. |
| lpad(string, int, fillchars) |
Pads string to length int by prepending characters fillchars. |
| md5(text) | Returns the hex MD5 value of text. |
| octet_length(string) | Returns the number of bytes in string. |
| overlaps | See above for syntax details. |
| quote_ident(string) | Returns string quoted for use as an identifier in an SQL statement. |
| quote_literal(string) | Returns string quoted for use as a string literal in an SQL statement. |
| regexp_replace | See PostgreSQL docs for details: reference doc, example doc |
| repeat(string, int) | Repeats string the specified number of times. |
| replace(string, matchString, replaceString) |
Searches string for matchString and replaces occurrences with replaceString. |
| rpad(string, int, fillchars) |
Pads string to length int by postpending characters fillchars. |
| similar_to(A,B,C) | String pattern matching using SQL regular expressions. 'A' similar to 'B' escape 'C'. See the PostgreSQL docs. |
| split_part(string, delimiter, int) |
Splits string on
delimiter and returns fragment number int
(starting the count from 1). SPLIT_PART('mississippi', 'i', 4) returns 'pp'. |
| string_to_array | See Array Functions in the PostgreSQL docs. |
| strpos(string, substring) |
Returns the position of substring in string. (Count starts from 1.) |
| substr(string, fromPosition, charCount) |
Extracts the number of characters specified by charCount from string starting at position fromPosition. SUBSTR('char_sequence', 5, 2) returns '_s' |
| to_ascii(string, encoding) |
Convert string to ASCII from another encoding. |
| to_hex(int) | Converts int to its hex representation. |
| translate(text, fromText, toText) |
Characters in string matching a character in the fromString set are replaced by the corresponding character in toString. |
| to_char | See Data Type Formatting Functions in the PostgreSQL docs. |
| to_date(textdate, format) |
See Data Type Formatting Functions in the PostgreSQL docs. |
| to_timestamp | See Data Type Formatting Functions in the PostgreSQL docs. |
| to_number | See Data Type Formatting Functions in the PostgreSQL docs. |
| unnest | See Array Functions in the PostgreSQL docs. |
JSON and JSONB Operators and Functions
LabKey SQL supports the following PostgreSQL JSON and JSONB operators and functions. Note that LabKey SQL does not natively understand arrays and some other features, but it may still be possible to use the functions that expect them.
See the PostgreSQL docs for usage details.
| PostgreSQL Operators and Functions | Docs |
| ->, ->>, #>, #>>, @>, <@, ?, ?|, ?&, ||, -, #- | LabKey SQL supports these operators via a pass-through function, json_op. The function's first argument is the operator's first operand. The first second is the operator, passed as a string constant. The function's third argument is the second operand. For example, this Postgres SQL expression:
a_jsonb_column --> 2 can be represented in LabKey SQL as:
json_op(a_jsonb_column, '-->', 2) |
| parse_json, parse_jsonb | Casts a text value to a parsed JSON or JSONB data type. For example,
'{"a":1, "b":null}'::jsonbCAST('{"a":1, "b":null}' AS JSONB)parse_jsonb('{"a":1, "b":null}') |
| to_json, to_jsonb | Converts a value to the JSON or JSONB data type. Will treat a text value as a single JSON string value |
| array_to_json | Converts an array value to the JSON data type. |
| row_to_json | Converts a scalar (simple value) row to JSON. Note that LabKey SQL does not support the version of this function that will convert an entire table to JSON. Consider using "to_jsonb()" instead. |
| json_build_array, jsonb_build_array | Build a JSON array from the arguments |
| json_build_object, jsonb_build_object | Build a JSON object from the arguments |
| json_object, jsonb_object | Build a JSON object from a text array |
| json_array_length, jsonb_array_length | Return the length of the outermost JSON array |
| json_each, jsonb_each | Expand the outermost JSON object into key/value pairs. Note that LabKey SQL does not support the table version of this function. Usage as a scalar function like this is supported: SELECT json_each('{"a":"foo", "b":"bar"}') AS Value
|
| json_each_text, jsonb_each_text | Expand the outermost JSON object into key/value pairs into text. Note that LabKey SQL does not support the table version of this function. Usage as a scalar function (similar to json_each) is supported. |
| json_extract_path, jsonb_extract_path | Return the JSON value referenced by the path |
| json_extract_path_text, jsonb_extract_path_text | Return the JSON value referenced by the path as text |
| json_object_keys, jsonb_object_keys | Return the keys of the outermost JSON object |
| json_array_elements, jsonb_array_elements | Expand a JSON array into a set of values |
| json_array_elements_text, jsonb_array_elements_text | Expand a JSON array into a set of text values |
| json_typeof, jsonb_typeof | Return the type of the outermost JSON object |
| json_strip_nulls, jsonb_strip_nulls | Remove all null values from a JSON object |
| jsonb_insert | Insert a value within a JSON object at a given path |
| jsonb_pretty | Format a JSON object as indented text |
| jsonb_set | Set the value within a JSON object for a given path. Strict, i.e. returns NULL on NULL input. |
| jsonb_set_lax | Set the value within a JSON object for a given path. Not strict; expects third argument to specify how to treat NULL input (one of 'raise_exception', 'use_json_null', 'delete_key', or 'return_target'). |
| jsonb_path_exists, jsonb_path_exists_tz | Checks whether the JSON path returns any item for the specified JSON value. The "_tz" variant is timezone aware. |
| jsonb_path_match, jsonb_path_match_tz | Returns the result of a JSON path predicate check for the specified JSON value. The "_tz" variant is timezone aware. |
| jsonb_path_query, jsonb_path_query_tz | Returns all JSON items returned by the JSON path for the specified JSON value. The "_tz" variant is timezone aware. |
| jsonb_path_query_array, jsonb_path_query_array_tz | Returns as an array, all JSON items returned by the JSON path for the specified JSON value. The "_tz" variant is timezone aware. |
| jsonb_path_query_first, jsonb_path_query_first_tz | Returns the first JSON item returned by the JSON path for the specified JSON value. The "_tz" variant is timezone aware. |
General Syntax
| Syntax Item | Description |
| Case Sensitivity | Schema names, table names, column names, SQL keywords, function names are case-insensitive in LabKey SQL. |
| Comments |
Comments that use the standard SQL syntax can be included in queries. '--'
starts a line comment. Also, '/* */' can surround a comment block:
-- line comment 1 |
| Identifiers | Identifiers in LabKey SQL may be quoted using double quotes. (Double quotes
within an identifier are escaped with a second double quote.) SELECT "Physical Exam".* ... |
| Lookups |
Lookups columns reference data in other tables. In SQL terms, they
are foreign key columns. See Lookups for details on creating lookup columns. Lookups
use a convenient syntax of the form
"Table.ForeignKey.FieldFromForeignTable" to achieve what would normally
require a JOIN in SQL. Example:
Issues.AssignedTo.DisplayName |
| String Literals |
String literals are quoted with single quotes ('). Within a single quoted
string, a single quote is escaped with another single quote.
|
| Date/Time Literals |
Date and Timestamp (Date&Time) literals can be specified using the JDBC escape syntax {ts '2001-02-03 04:05:06'} {d '2001-02-03'} |
Related Topics
LabKey LIMS
- Generate customizable reports with ease.
- Add automation with transformation scripts for assay data.
- Customize the downloadable templates for data structures like samples, sources, and assays.
LIMS: Downloadable Templates
- Add Custom Templates (Administrator)
- Download Templates
Add Custom Templates (Administrator)
- Open the Sample Type, Source Type, or Assay Design for which you want to include customized templates.
- Select Manage > Manage Templates.
- You'll see the Default Template and can click to download it as a starting place if desired.
- Click Add a Template to add your own additional ones.
- Give the new template a name that will help your users know when to use it.
- Select or drag and drop the new template file into the target area.
- Click Save.
Download Templates
When a data structure has more than the default template defined, users will be able to select a template by name.Related Topics
LIMS: Samples
- Sample Types:
- Samples:
- Add Samples
- Aliquots, Derivatives, and Sample Pooling
- Sample Details: Lineage, Aliquots, Assays, Jobs, Timeline
- Sample Status
- Sample Picklists
- Sample Grids Containing Samples of Several Types
- Multi-tab Excel Exports
- Sample Finder
Related Topics
- Entity Lineage
- Samples: General information about Sample Types
- Sample ID Naming
- Sample Lineage / Parentage
- Barcode Fields
- Print Labels with BarTender
Print Labels with BarTender
This topic describes how to use LabKey applications, including Sample Manager and Biologics LIMS, with BarTender for printing labels for your samples. Note that an administrator must first complete the one-time steps to configure BarTender Automation. Once configured any user may send labels to the web service for printing.
Configuration steps and management of label templates:Print BarTender Labels for Samples
Before you can print labels, an administrator must have completed the one-time setup steps in this topic, and configured LabKey to print labels. The admin can also specify a folder-specific default label file. When printing, the user can specify a different variant to use.
After configuring BarTender, all users will see the options to print labels in the user interface.
Print Single Sample Label
Open the Sample Type from the main menu, then open details for a sample by clicking the SampleID.
Select Print Labels from the Manage menu.
In the popup, specify (or accept the defaults):
- Number of copies: Default is 1.
- Label template: Select the template to use among those configured by an admin. If the admin has set a default template, it will be preselected here, but you can use the menu or type ahead to search for another.
- Click Yes, Print to send the print request to BarTender.
Print Multiple Sample Labels
From the Sample Type listing, use checkboxes to select the desired samples, then select Print Label from the (Export) menu.
In the popup, you have the following options:
- Number of copies: Specify the number of labels you want for each sample. Default is 1.
- Selected samples to print: Review the samples you selected; you can use the Xs to delete one or more of your selections or open the dropdown menu to add more samples to your selection here.
- Label template: Select the template to use among those configured by an admin. You can type ahead to search. The default label template file can be configured by an admin.
- Click Yes, Print to send the print request to BarTender.
Download BarTender Template
To obtain a BarTender template in CSV format, select > Download Template.

Troubleshooting
If you have trouble printing to BarTender from Chrome (or other Chromium-based browser), try again using Firefox.
If you are not seeing values populate in your labels, confirm that you have configured BarTender to have the necessary data source connection.
Error Reporting
If there is a problem with your configuration or template, you will see a message in the popup interface. You can try again using a different browser, such as Firefox, or contact an administrator to resolve the configuration of BarTender printing.
Related Topics
LIMS: Assay Data
| SampleID | Expression Run | SampleDate | InjVol | Vial | Cal CurveID | Dilution | ResultID | MAb |
|---|---|---|---|---|---|---|---|---|
| 2016_01_22-C3-12-B9-01 | ER-005 | 2016-01-23 | 30 | 2:A,19 | 46188 | 1.0000 | 47836 | 0.2000 |
| 2016_04_21-C7-73-B6-02 | ER-005 | 2016-04-22 | 30 | 2:B,19 | 46188 | 1.0000 | 47835 | 0.2300 |
| 2015_07_12-C5-39-B4-02 | ER-004 | 2015-07-13 | 30 | 2:C,19 | 46188 | 1.0000 | 47834 | 0.3000 |
Documentation
Learn more about using the general assay framework in the Sample Manager documentation here:Additional Features with LabKey LIMS
Additional Features with Biologics LIMS
LIMS: Charts
Add a Chart
To add a chart:- On a data grid, click the Charts menu and choose Create Chart.
- In the pop up dialog, enter the following fields and options:
- Enter a Name and select whether to:
- Make this chart available to all users
- Make this chart available in child folders
- Select a Chart Type:
- Bar
- Box
- Line
- Pie
- Scatter
- Depending on the chart type, you will see selectors for X-axis, Y-axis, and various other settings needed for the chart. Required fields are marked with an asterisk.
- Once you have made selections, you will see a preview of the chart.
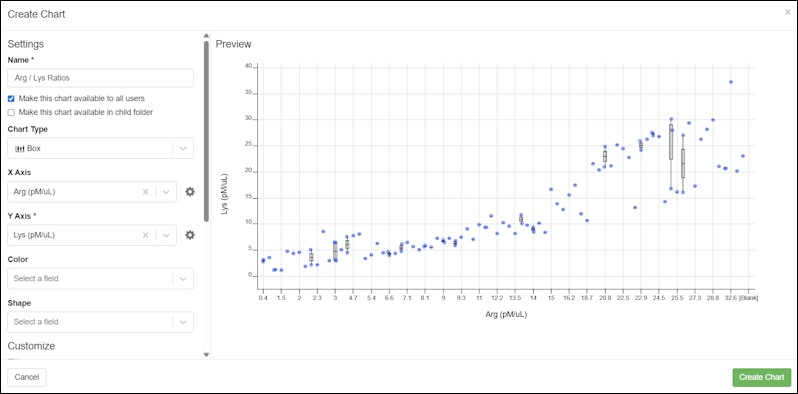
- Click Create Chart to save it.
Trendline Options
View a Chart
Any charts available on a data grid will be listed on the Charts menu above the grid.Selecting a chart will render it above the grid as shown here.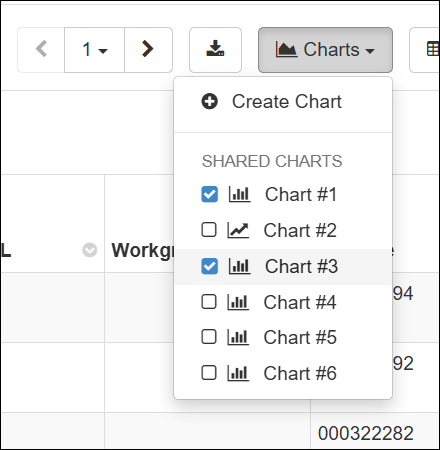
Edit a Chart
To edit a chart, open it from the menu and click the .Error Bars and Aggregation Methods
For Bar and Line charts, aggregation and error bar options are available by clicking the Y-axis gear icon.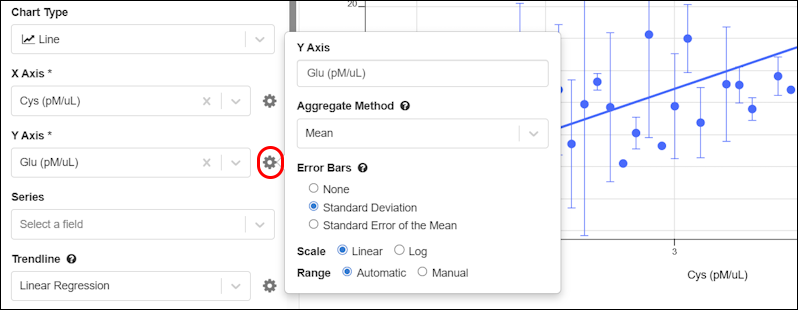 When the aggregation method is set to Mean, then the options for error bars are shown: None, Standard Deviation, Standard Error of the Mean.
When the aggregation method is set to Mean, then the options for error bars are shown: None, Standard Deviation, Standard Error of the Mean.
Export a Chart as PDF or PNG
Click the (Download) button to choose either PNG or PDF format for your export.Related Topics
LIMS: Storage Management
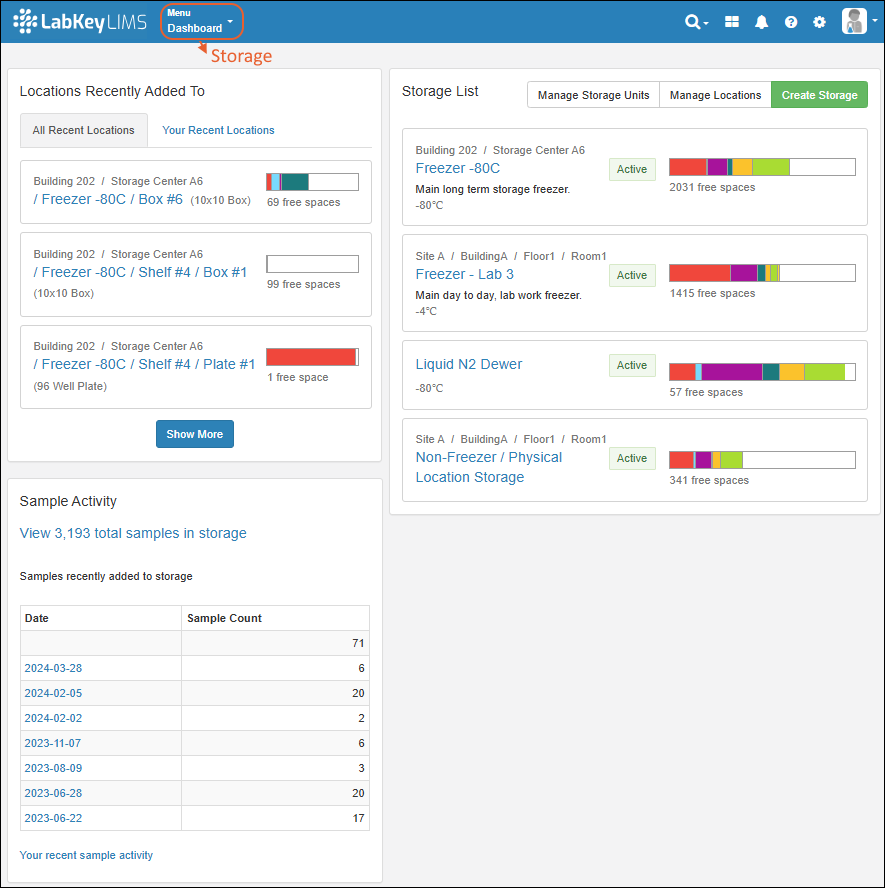
Topics
Learn more in the Sample Manager documentation here:- Storage Management
- Create Storage
- Store Samples
- Manage Storage
- Manage Storage Unit Types
- View Storage Details
- Edit Storage Definition
- Manage Storage Locations
- Move Stored Samples
- Manage Stored Samples
- Check Out and Check In
- View Storage Activity
- Migrate Storage Data into LabKey
- Storage Roles
- Storage Management API
- Shared Storage Across Folders
LIMS: Workflow
- Use workflow templates to standardize common task sequences
- Create jobs to track and prioritize sequential tasks
- Assign work to the right users
- Track progress toward completion
Biologics LIMS
- Record and document your ongoing work in a data-aware Electronic Lab Notebook.
- Easily and uniquely register all biological entities individually or in bulk.
- Track and query the lineage of samples throughout many generations of derivation.
- Integrate biological entity and sample information with downstream assay data used to evaluate therapeutic properties and provide a holistic view of experiment results.
- Manage and monitor the execution of laboratory tasks and requests, supporting efficient collaboration across teams.
- Manage the contents of your freezers and other storage systems using a digital match for your physical storage.
Biologics Release Notes
Biologics Overview
The Biologics LIMS product builds upon the same application core as Sample Manager. As you get started with Biologics, you may find the introductory Sample Manager documentation helpful in learning the basics of using the application.Basic Navigation and Features
Bioregistry
- Biologics: Bioregistry
- Create Registry Sources
- Register Nucleotide Sequences
- Register Protein Sequences
- Register Leaders, Linkers, and Tags
- Register Molecules
- Protein Sequence Annotations
- CoreAb Sequence Classification
- Biologics: Chain and Structure Formats
- Molecules, Sets, and Molecular Species
- Vectors, Constructs, Cell Lines, and Expression Systems
- Entity Lineage
- Bulk Registration of Entities
- Registry API
- Configuring Columns Seen in Grids and Detail Views
- Biologics: Protect Sequence Fields
Samples
Plates
Assay Management
- Biologics: Assay Data
- Biologics: Assay Batches and QC
- Biologics: Upload Assay Data
- Biologics: Specialty Assays
Media Registration
- Biologics: Media Registration
- Managing Ingredients and Raw Materials
- Registering Mixtures (Recipes)
- Registering Batches
Workflow
ELN
Storage Management
Related Topics
Introduction to LabKey Biologics
- A consolidated Bioregistry knowledge base to store and organize all your data and entity relationships
- Sample and Storage management tracking production batches, inventories, and all contents of your physical storage systems
- A data-integrated electronic lab notebook (ELN) for recording your research and coordinating review
- Tools promoting collaboration among teams for automated workflows, smooth handoffs, and reproducible procedures
- Media management for clear tracking of ingredients and mixtures in the Enterprise Edition
Knowledge Base
Bioregistry
The Bioregistry forms the team's shared knowledge base, containing detailed information on your sequences, cell lines, expression systems, and other research assets. The application is prepopulated with common Registry Source Types.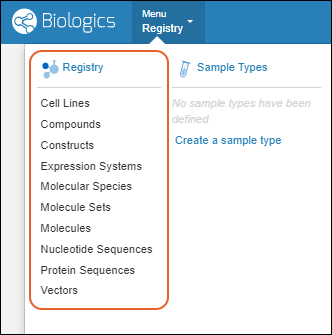
- Cell Lines
- Compounds
- Constructs
- Expression System
- Molecules
- Molecule Sets
- Molecule Species
- Nucleotide Sequences
- Protein Sequences
- Vectors
Entity Details
Click an entity in a grid to see its details page. Each details page shows the entity's properties and relationships to other entities.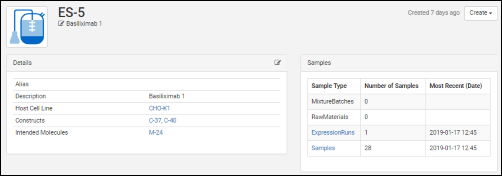 All details pages can be configured to show the most relevant data. For example, the details page for a protein sequence shows the chain format, average mass, extinction coefficient, the number of S-S bonds, etc., while the details page for a expression system shows the cell lines, constructs, and target molecule, as well as the samples drawn from it.
All details pages can be configured to show the most relevant data. For example, the details page for a protein sequence shows the chain format, average mass, extinction coefficient, the number of S-S bonds, etc., while the details page for a expression system shows the cell lines, constructs, and target molecule, as well as the samples drawn from it.
Entity Relationships
Each details page contains a panel of relationships to other entities. For example, for a given protein sequence, the relationship panel shows: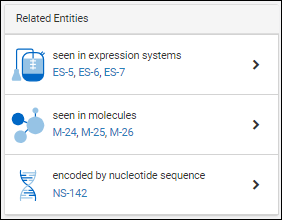
- which expression systems it is included in
- which molecules it is a part of
- which nucleotide sequence encodes for it
Sample Management and Data Integration
Data about your samples, experiments, and instrument assay results can be integrated to provide a full data landscape. Design customized methods for representing and linking your information. Using Biologics LIMS follows the same interface and has all the capabilities of the other Sample Management applications.Analytics and Visualizations
Analytics and visualizations can be included throughout LabKey Biologics to provide key insights into the large molecule research process.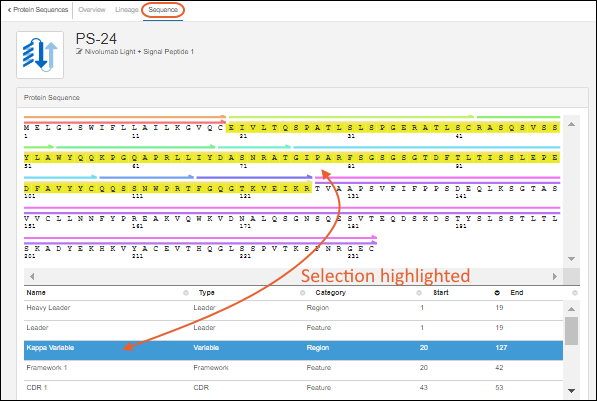
Protein Classification Engine
When new protein sequences are added to the Registry, they are passed to the classification and annotation engine, which calculates their properties and identifies key regions on the protein chain, such as leader sequences, variable regions, and constant regions. (Researchers can always override the results of the classification engine, if desired.) Results are displayed on the Sequence tab of the protein's detail page. Identified regions are displayed as multicolored bars, which, when clicked, highlight the annotation details in the scrollable section below the sequence display.Workflow Collaboration
A fully customizable task-based workflow system is included in LabKey Biologics. Individual users can all be working from the same knowledge base with personalized task queues and priorities. Several example jobs with representative tasks are included in your trial, you can also create your own.Media Management (Enterprise Edition feature)
In the Media section of the main menu, you will find sections for tracking:- Batches
- Ingredients
- Mixtures
- Raw Materials
Electronic Lab Notebook
Our data-integrated ELN (electronic lab notebook) helps you record results and support collaboration and publication of your research. Directly link to relevant registry entities, specific result runs, and other elements of your biologics research. Submit for review and track feedback and responses within the same application.Learn more in this section:Storage/Freezer Management
When you use Storage Management tools within LabKey Biologics, you can directly track the locations of samples and media in all the freezers and other storage systems your team uses. Create an exact digital match of your physical storage and record storage details and movements accurately. Sample data is stored independently of storage data, so that all data is retained even after a sample is consumed or removed from storage.Learn more in the Sample Manager documentation here:Release Notes: Biologics
Biologics LIMS offers strong support for antibody discovery operations. Learn more in our webinar:
Biologics LIMS includes all features available in LabKey LIMS, plus:
- Bioregistry and classification engine
- Media management
- Molecule property calculator
- Plate and plate set support
- Antibody screening and characterization
Each new release of Biologics LIMS includes all the feature updates covered in the Sample Manager and LIMS Release Notes, plus additional features listed on this page.
Release 26.2, February 2026
- GenBank import improved: nearly all information GenBank files is captured on import, including the original file.
- Improved Molecule creation: Select protein sequences to kick off the molecule creation process.
- Column widths now adjust dynamically, allowing more columns to be visible at once with less horizontal scrolling. (docs)
- Configured URL links can now be opened in a new browser tab for easier comparison and multitasking. (docs)
- Entities you don't have access to in lineage views are now shown as restricted rather than being omitted, preserving full context without exposing details. (docs)
- Sample Status is available as a filter for "All Sample Types" in Sample Finder. (docs)
Release 26.1, January 2026
- Support for multiple unit types provides improved inventory and material management. (docs)
- Move workflow jobs to different folders to better reflect changes in projects or organization. (docs)
- Workflow tasks now support sample filters, allowing you to control which samples are included at each step. (docs)
- Improved plot customization with new layout, axis, size, color, and per-series line controls. (docs)
- Client APIs can query and update samples using the RowId value; using the LSID value is no longer required.
- Sample names (SampleId) can be updated via a file, when RowId is provided.
Release 25.12, December 2025
- Amount and Units Fields - Improvements have been made to ensure that the Amounts & Units fields function as paired fields. (docs)
- Negative Amount Values Disallowed - Sample Manager now enforces that the Amounts field cannot have a negative value.(docs)
- Identifying Fields - Identifying fields are now shown in more assay import scenarios. (docs)
Release 25.11, November 2025
- Audit log captures the method used to insert, update, and delete records. (docs)
- When an ELN notebook is recalled by an administrator, the author will now receive an email notification, improving visibility and timely follow-up.
- The Customize Grid View and Filter pop-up dialogs now list fields alphabetically, making it faster and more intuitive to find and select fields.
- Error bars are available on Bar and Line charts. (docs)
- Multiple charts can be displayed above data grids. Select up to 5 charts to display. (docs)
Release 25.3, March 2025
- Rapidly find the plates and experiments in which samples have been used, and vice versa.
- Automatically generate analytics like regressions and statistics to accelerate your work
Release 25.2, February 2025
- Support for advanced plate layouts using using dilutions.
- Use the "Replicate Group" column to denote a plate well as a replicate instead of setting the well's type to "Replicate".
- Replicate wells have a type of "Sample" and the "Replicate Group" will need to be filled in.
- Add Samples to an existing Plate Set.
- Navigate from a plate set to any notebooks that reference it.
- Edits to outlier exclusions will result in the rerunning of any transform scripts that are configured to run on update.
Release 25.1, January 2025
- Users can now specify hit selection filter criteria on Assay fields. When a run is imported/edited the hit selections for the assay results will be recomputed and automatically applied based on these criteria.
- Navigate from a sample to the plate(s) it has appeared on.
- Perform many types of linear regression analysis and chart them.
- Exclude outlier plate-based assay data points and have that reflected in calculations and charts.
Release 24.12, December 2024
- Plate sets can be referenced from an Electronic Lab Notebook.
Release 24.11, November 2024
Major antibody discovery and characterization updates including:
- Campaign modeling with plate set hierarchy support.
- Plan plates easier with graphical plate design and templating.
- Automate routine analyses from raw data collected.
- Perform hit selection from multiple, integrated results across plates and data types.
- Generate instructions for liquid handlers and other instruments.
- Automatically integrate multi-plate results including interplate replicate aggregation.
- Dive deeper into plated materials to understand their characteristics and relationships from plates.
Release 24.10, October 2024
- Charts are added to LabKey LIMS, making them an "inherited" feature set from other product tiers. (docs)
Release 24.7, July 2024
- A new menu has been added for exporting a chart from a grid. (docs)
Release 23.12, December 2023
- The Molecule physical property calculator offers additional selection options and improved accuracy and ease of use. (docs)
Release 23.11, November 2023
Release 23.9, September 2023
- Charts, when available, are now rendered above grids instead of within a popup window. (docs)
Release 23.4, April 2023
- Molecular Physical Property Calculator is available for confirming and updating Molecule variations. (docs)
- Lineage relationships among custom registry sources can be represented. (docs)
- Users of the Enterprise Edition can track amounts and units for raw materials and mixture batches. (docs | docs)
Release 23.3, March 2023
- Potential Backwards Compatibility Issue: In 23.3, we added the materialExpDate field to support expiration dates for all samples. If you happen to have a custom field by that name, you should rename it prior to upgrading to avoid loss of data in that field.
- Note that the built in "expirationDate" field on Raw Materials and Batches will be renamed "MaterialExpDate". This change will be transparent to users as the new fields will still be labelled "Expiration Date".
Release 23.2, February 2023
- Protein Sequences can be reclassified and reannotated in cases where the original classification was incorrect or the system has evolved. (docs)
- Lookup views allow you to customize what users will see when selecting a value for a lookup field. (docs)
- Users of the Enterprise Edition may want to use this feature to enhance details shown to users in the "Raw Materials Used" dropdown for creating media batches. (docs)
Release 23.1, January 2023
- Heatmap and card views of the bioregistry, sample types, and assays have been removed.
- The term "Registry Source Types" is now used for categories of entity in the Bioregistry. (docs)
Release 22.12, December 2022
- Projects were added to the Professional Edition of Sample Manager, making this a common feature shared with other tiers.
Release 22.11, November 2022
- Improvements in the interface for managing Projects. (docs)
- New documentation:
Release 22.10, October 2022
- Improved interface for creating and managing Projects in Biologics. (docs)
Release 22.9, September 2022
- When exploring Media of interest, you can easily find and review any associated Notebooks from a panel on the Overview tab. (docs)
Release 22.8, August 2022
- Search for data across projects in Biologics. (docs)
Release 22.7, July 2022
- Biologics subfolders are now called 'Projects'; the ability to categorize notebooks now uses the term 'tags' instead of 'projects'. (docs | docs)
Release 22.6, June 2022
- New Compound Bioregistry type supports Simplified Molecular Input Line Entry System (SMILES) strings, their associated 2D structures, and calculated physical properties. (docs)
- Define and edit Bioregistry entity lineage. (docs)
- Bioregistry entities include a "Common Name" field. (docs)
Release 22.3, March 2022
- Mixture import improvement: choose between replacing or appending ingredients added in bulk. (docs)
Release 21.12, December 2021
Release 21.11, November 2021
- Apply a container specific prefix to all bioregistry entity and sample naming patterns. (docs)
Release 21.10, October 2021
- Customize the definitions of data classes (both bioregistry and media types) within the application. (docs)
Release 21.9, September 2021
- Customize the names of entities in the bioregistry (docs)
Release 21.7, July 2021
- Removal of the previous "Experiment" mechanism. Use workflow jobs instead.
Release 21.5, May 2021
- Nucleotide and Protein Sequence values can be hidden from users who have access to read other data in the system. (docs)
Release 21.4, April 2021
- Specialty Assays can now be defined and integrated, in addition to Standard Assays (docs)
- Creation of Raw Materials in the application uses a consistent interface with other sample type creation (docs)
Release 21.3, March 2021
- Biologics LIMS begins using the same user interface as Sample Manager.
- Release notes for this and other versions prior to this change can be found in the documentation archives.
Release 17.1, March 2017
Biologics: Navigate
- LabKey Biologics: Home Page
- Navigation
- Menu > Registry Sources - Browse all of the registry sources.
- Menu > Sample Types - A dashboard for tracking samples.
- Menu > Assays - Assay results for candidate molecules.
- Menu > Storage - Manage freezer contents and capacities.
- Menu > Workflow - Create and track custom workflow jobs and tasks.
- Menu > Media - A dashboard for ingredients, media, mixtures, and recipes. (Enterprise Edition Feature)
- Menu > Picklists - User-defined lists of samples.
- Menu > Notebooks - Electronic lab notebooks integrated directly with your data.
LabKey Biologics: Home Page
The main dashboard on the home page provides quick links into different aspects of the data.The top header bar is available throughout the application and includes:- The clickable logo, for returning to this dashboard at any time.
- A main Menu button, giving access to all of your data and activities.
- A set of common resources in the upper right:
- : Search options
- : A product selection menu for switching between Biologics and LabKey Server when needed
- : Notifications
- : Help Opens the documentation for LabKey Biologics LIMS.
- : Administration Settings
- A user menu, indicated by your avatar, or a default graphical one
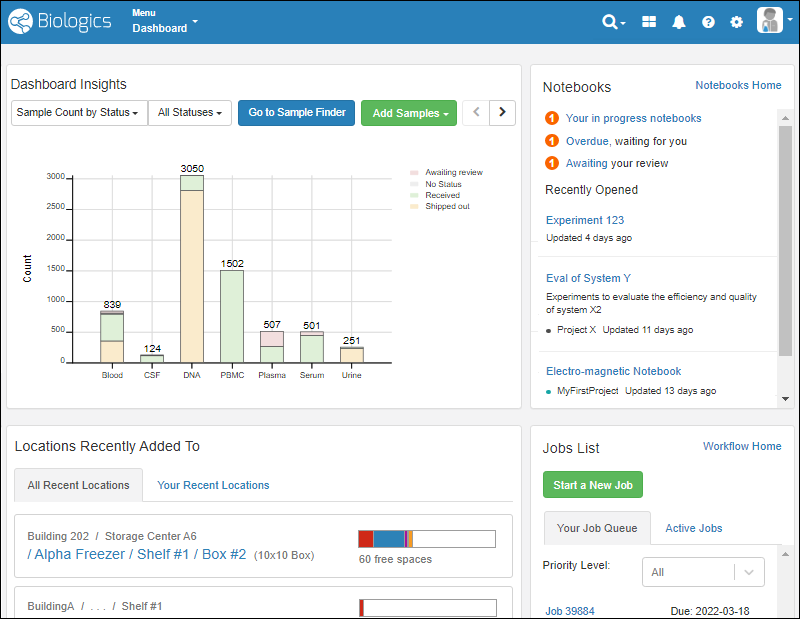 On the dashboard, you'll see panels displaying:
On the dashboard, you'll see panels displaying:
- Dashboard Insights: A selectable set of visualizations, defaulting to the Sample Count by Status.
- Sample Types can be selectively excluded from the graphs on this dashboard, as described here.
- Notebooks: See task status, recent notebooks, and link to your Electronic Lab Notebooks dashboard.
- Locations Recently Added To: Access to storage where samples have been added by you or others.
- Jobs List: The home for workflow jobs. If any tasks were assigned to you, they would be shown on Your Job Queue and can be filtered by priority.
Header Menus
The menus in the upper right offer these selections:- : Search options:
- Site-wide text search
- Find samples by barcode () or ID ()
- Sample Finder: Find samples by their properties or associated data
- : A product selection menu for switching between Biologics and LabKey Server when needed
- : Notifications
- Documentation:
- Help: Opens the documentation for LabKey Biologics LIMS.
- Release Notes: Biologics
- : Administration Settings:
- Application Settings: See below.
- Audit Logs
- Groups
- Permissions
- Folders
- Users
- User avatar menu
- Notification Settings
- Profile: Your user details, including the option to add an avatar image.
- Sign Out (or Sign In)
Application Settings
Select > Application Settings to manage:- Display Settings
- Notebook Settings
- BarTender Configuration
- ID/Name Settings
- Manage Sample Statuses
- Audit Logging
- Protected Data Settings
Navigation
The main Menu is available throughout the application and gives you quick access to all aspects of Biologics. It will include the name of the category you are in ("Dashboard" if not one of the specific categories).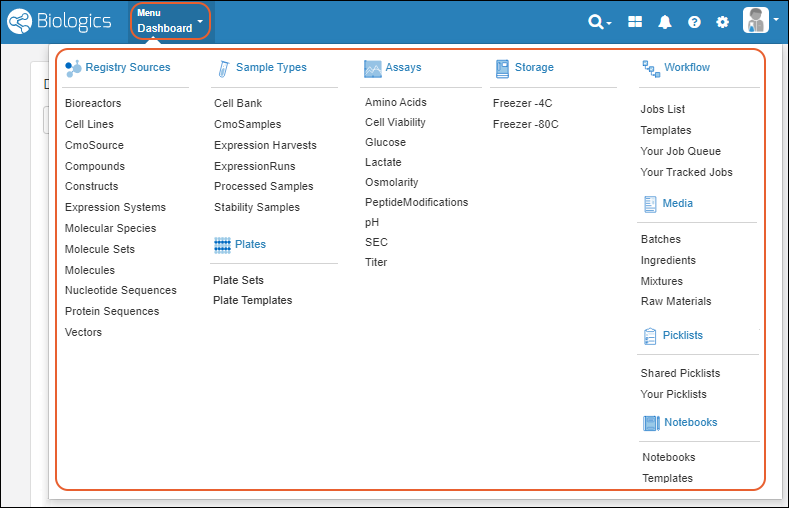 Click the menu categories for sub-dashboards, such as for Registry Sources, Notebooks, or Sample Types. Click individual menu items for that specific category.
Click the menu categories for sub-dashboards, such as for Registry Sources, Notebooks, or Sample Types. Click individual menu items for that specific category.
Biologics: Projects and Folders
- Options for Sharing Biologics Data
- Use Multiple Biologics Folders
- ID/Name Settings
- Shared Data Structures (Sample Types Registry Source Types, Assays, Storage Systems)
Options for Sharing Biologics Data
Biologics runs in the context of a LabKey project or folder container. Each container can have unique permissions assignments and data, supporting a secure and isolated workspace. If desired, many resources, including assay designs and sample type definitions, can also be shared across multiple containers. Further, within a LabKey project of type "Biologics", an administrator can add folders which allow data partitioning.The Biologics administrator should determine the configuration that best represents the organization.Example Scenarios:1. You might have two groups doing distinct work with no need to share data or compare results. These two groups could each use a separate Biologics container (LabKey project or folder) and keep all resources within it. Permissions are assigned uniquely within each container, with no built-in interaction between the data in each.2. If two groups needed to use the same assay designs or sample types, but would not otherwise share or compare the actual data, those definitions could be placed in the Shared project, with all collection and analysis in containers (LabKey projects or folders) for the two groups as in option 1. Note that in this scenario, samples created of those shared types would have the same naming pattern and if you used a container-specific prefix, it would not be applied to the samples of the shared type.3. If multiple groups need to share and compare data, and also keep clear which data came from which group originally, you could apply a container prefix in each group's folder so that all data would be consistently identified. In this scenario, each container would need its own sample type definitions so that the naming patterns would propagate the prefix into the created samples.4. For more integrated sharing, configure Biologics in a top level LabKey project, then add one or more subfolders, all also of folder type "Biologics". Bioregistry and Sample definitions can be shared among the Folders and permissions controlled independently. Learn about this option in the next section.5. Storage systems (freezers, etc.) can be shared among folders with different permissions. Users will only be able to see details for the stored samples to which they have been granted access. Learn more about shared storage in the Sample Manager documentation:Manage Multiple Biologics Folders
When using Biologics in a top-level LabKey Server container, administrators have the option to manage multiple Biologics Folders directly from within the admin interface. Note that this is not supported when the top level Biologics container is a folder or subfolder in LabKey Server.To manage Folders, select > Folders. You can customize types of data and specific storage systems available per Folder. Learn more in the Sample Manager documentation here: When in use, you'll see Folders listed on the left side of the main menu. Select the desired Folder, then click the item of interest on the menu or the links for its dashboard or settings. You will see the contents of the application 'scoped' to the selected Folder. Selecting the top-level container (aka the "home") will show all data the user can access across all Folders to which they have been granted "Read" permissions.Cross-Folder Actions
When multiple Biologics Folders are in use, the notes about cross-folder actions are the same as for Sample Manager, where Registry Sources are called Sources. Learn more in this topic:Restricted Visibility of Inaccessible Data
When a user has access to a subset of folders, there can be situations where this user will be restricted from seeing data from other folders, including identifiers such as Sample IDs that might contain restricted details. In detail pages, listing pages, and grids, entities a user does not have access to will show "unavailable" in the display instead of a name or rowid. Learn more in this topic:ID/Name Settings
Select > Application Settings and scroll down to the ID/Name Settings section to control several options related to naming of entities in the application.Force Usage of Naming Patterns for Consistency
To maintain consistent naming, particularly when using container-specific naming prefixes, you may want to restrict users from entering their own names for entities. Learn more here:Apply Naming Prefix
You can apply a container-specific naming prefix that will be added to naming patterns to assist integration of data from multiple locations while maintaining a clear association with the original source of that data.This prefix is typically short, 2-3 characters long, but will not be limited. Prefixes must be unique site-wide, and should be recognizable to your users. Before setting one, make sure you understand what will happen to the naming patterns and names of existing entities in your application.- The prefix will be applied to names created with naming patterns for all Sample Types and Registry Source Types in the container.
- New samples and entities created after the addition of the prefix will have names that include the prefix.
- Existing samples and entities created prior to the addition of the prefix will not be renamed and thus will not have the prefix (or might have a different previously-applied prefix).
- Sample aliquots are typically created and named including the name of the sample they are aliquoted from. This could mean that after the prefix is applied, new aliquots may or may not include the prefix, depending on whether the originating sample was created before or after the prefix was applied. Learn more about aliquot naming here: Aliquot Naming Patterns.
- Select > Application Settings.
- Scroll down to the ID/Name Settings section.
- Enter the prefix to use. You will see a preview of what a name generated with the Naming Pattern with the prefix applied might look like using a representative example, Blood-${GenId}:
- Click Apply Prefix to apply it.
- This action will change the Naming Pattern for all new and existing Sample Types and Registry Source Types. No existing IDs/Names will be affected. Are you sure you want to apply the prefix?
- Click Yes, Save and Apply Prefix to continue.
Naming Pattern Elements/Tokens
The sampleCount and rootSampleCount tokens are used in Naming Patterns across the application.Learn more about these naming pattern tokens in this topic:Shared Data Structures (Sample Types, Registry Source Types, Assays, and Storage)
Sample Types, Registry Source Types, Assays, and Storage Systems are defined in the home folder and available in subfolders, provided an administrator has not 'hidden' them. Administrators can edit these shared data structures from the home any folder, but the changes will be made at the home level, applying to all folders.In addition, any Sample Types, Registry Source Types, and Assay definitions defined in the /Shared project will be available for use in LabKey Biologics. You will see them listed on the menu and dashboards alongside local definitions.From within the Biologics application, users editing (or deleting) a shared definition will see a banner indicating that changes will affect other folders. The same message is applied when editing a data structure from a folder within the Biologics application.Related Topics
Biologics: Bioregistry
- Create Registry Sources
- Register Nucleotide Sequences
- Register Protein Sequences
- Register Leaders, Linkers, and Tags
- Register Molecules
- Registry Reclassification
- Biologics: Terminology
- Protein Sequence Annotations
- CoreAb Sequence Classification
- Biologics: Chain and Structure Formats
- Molecules, Sets, and Molecular Species
- Vectors, Constructs, Cell Lines, and Expression Systems
- Compounds and SMILES Lookups
- Entity Lineage
- Customize the Bioregistry
- Bulk Registration of Entities
- Use the Registry API
- Biologics: Detail Pages and Entry Forms
Create Registry Sources
- Create Registry Sources in the User Interface
- Create/Import Registry Sources from File
- Naming Patterns
- Hide Name Entry/Edit Options
- Review Entity Details
Create Registry Sources in the User Interface
In this example, we show creation of a new cell line. Other kinds of sources will have different fields that compose them, and may also have additional tabs in the creation wizard. See specific documentation listed at the end of this topic.- From the main menu, click the type of registry source to create. Then use the Add > menu:
- Add Manually: Use the data entry wizard.
- Add Manually from Grid: Create one or more using a grid interface.
- Import from File
Add Manually
- Provide the details in the registration wizard:
- Name: Provide a short unique name, or leave this field blank to have one generated using the naming pattern for this Registry Source Type.
- Hover over the to see an example generated name.
- Description: Optional, but will be shown in the grids and can be a helpful way to illustrate the entity.
- Common Name: Every entity includes a field for the common name.
- Remaining fields: Required fields are marked with an asterisk.
- When the fields are completed, click Finish to create the new entity.
Add Manually from Grid
Using the grid interface to create Registry Sources as described in the Sample Manager documentation for Sources.Note that this option not supported for all Registry Source Types. You will not see this option on the menu for Nucleotide Sequences, Protein Sequences, Molecules, Molecular Species, etc.Create/Import Entities from File
For bulk registration of many entities, including registry sources, samples, assay result data, ingredients, and raw materials, you can make use of importing new data from file. Templates are available to assist you in reliably uploading your data in the expected format.Learn more in the Sample Manager documentation for importing Samples from file.Update or Merge Registry Sources from File
To update data for existing Registry Sources from file, or to import a spreadsheet merging updates with creation of new Registry Sources, use Edit > Update from File.Learn more in the Sample Manager documentation for updating Samples from file.Entity Naming Patterns
If you do not provide a name, the naming pattern for the Registry Source Type will be used to generate one. Hover over the to see the naming pattern in a tooltip, as well as an 'example name' using that pattern.| Registry Source Type | Default Naming Pattern |
|---|---|
| Cell Lines | CL-${genId} |
| Compounds | CMP-${genId} |
| Constructs | C-${genId} |
| Expression Systems | ES-${genId} |
| Molecular Species | MSp-${genId} |
| Molecule Sets | MS-${genId} |
| Molecules | M-${genId} |
| Nucleotide Sequences | NS-${genId} |
| Protein Sequences | PS-${genId} |
| Vectors | V-${genId} |
Hide Name Entry/Edit Options
An administrator can hide the Name field for insert, update, or both. When insert of names is hidden, they will be generated using the naming pattern for the registry source type. When update of names is hidden, names remain static after entity creation.This can be done:- On an application-wide basis by disabling user-supplied names entirely.
- For some but not all Registry Source Types, by using Query Metadata overrides.
Review Registry Source Details
Once created, you'll see a grid of all entities of the given type when you select it from the main menu.To see details for a specific Registry Source, click the name.- Overview: Panels for details, samples, related entities (for built in Registry Source Types), notebooks, and parent sources.
- Lineage: See the lineage in graph or grid format.
- Samples: Contains a grid of all samples 'descended' from this source.
- Assays: See all assay data available for samples 'descended' from this source.
- Jobs: All jobs involving samples 'descended' from this source.
- Sequence (Available for Protein and Nucleotide Sequences).
- Molecular Property Calculator (Available for Molecules)
Related Topics
Register Nucleotide Sequences
Nucleotide Sequence Validation
For nucleotide sequences, we allow DNA and RNA bases (ACTGU) as well as the IUPAC notation degenerate bases (WSMKRYBDHVNZ). On import, whitespace will be removed from a nucleotide sequence. If the sequence contains other letters or symbols, an error will be raised.For protein sequences, we only allow standard amino acids letters and zero or more trailing stop codon '*'. On import, whitespace will be removed from a protein sequence. If the sequence contains stop codons in the middle of the sequence or a other letters or symbols, an error will be raised.When translating a nucleotide codon triple to a protein sequence, where the codon contains one or more of the degenerate bases, the system attempts to find a single amino acid that could be mapped to by all of the possible nucleotide combinations for that codon. If a single amino acid is found, it will be used in the translated protein. If not, the codon will be translated as an 'X'.For example, the nucleotide sequence 'AAW' is ambiguous since it could map to either 'AAA' or 'AAT' (representing Lysine and Asparagine respectively), so 'AAW' will be translated as an 'X' However, 'AAR' maps to either 'AAA" or 'AAG' which are both are translated to Lysine, so it will be translated as a 'K'.Create a Nucleotide Sequence
To add a new nucleotide sequence to the registry:- Open the Nucleotide Sequence page, then select Add > Add manually.
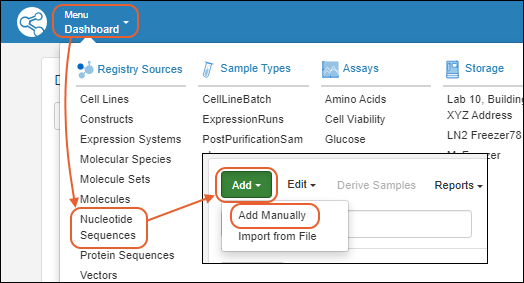 The creation wizard has two tabs:
The creation wizard has two tabs:
Details
On the Register a new Nucleotide Sequence page, in the Details panel, populate the fields: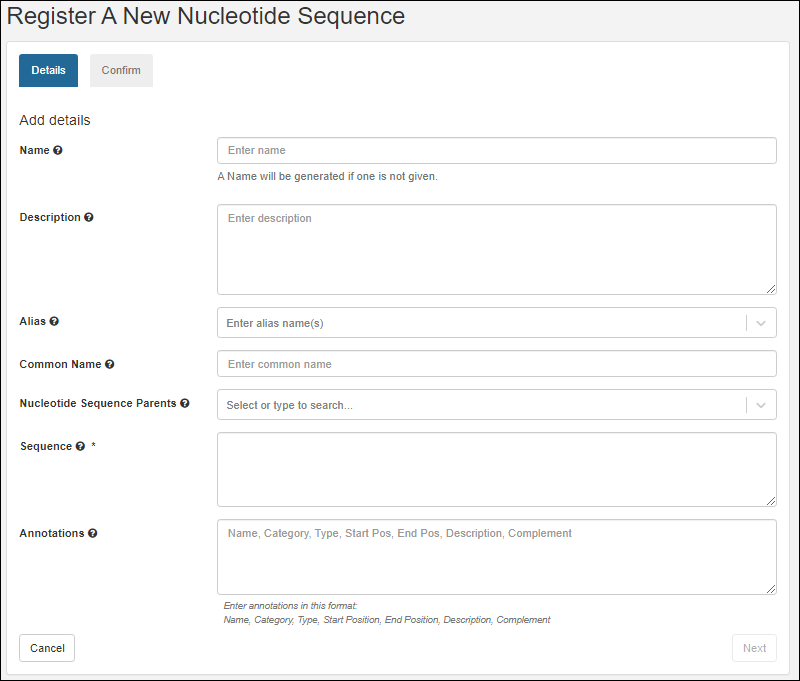
- Name: Provide a name, or one will be generated for you. Hover to see the naming pattern
- Description: (Optional) A text description of the sequence.
- Alias: (Optional) Alternative names for the sequence. Type a name, click enter when complete. Continue to add more as needed.
- Common Name: (Optional) The common name for this sequence, if any.
- Nucleotide Sequence Parents: (Optional) Parent components. A related sequence the new sequence is derived from, for example, related as a mutation. You can select more than one parent. Start typing to narrow the pulldown menu of options.
- Sequence: (Required) The nucleotide sequence
- Annotations: (Optional) A comma separated list of annotation information:
- Name - a freeform name
- Category - region or feature
- Type - for example, Leader, Variable, Tag, etc.
- Start and End Positions are 1-based offsets within the sequence.
- Description
- Complement
Confirm
Review the details on the Confirm tab.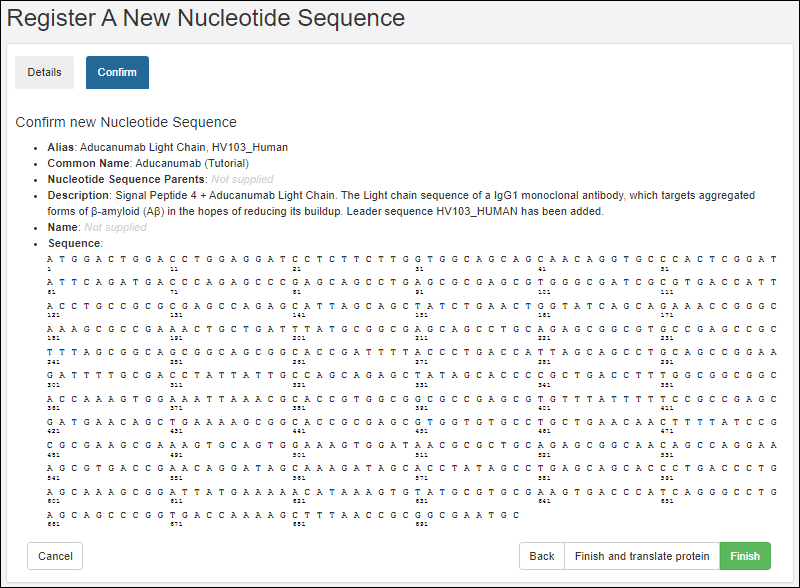 Options to complete registration:
Options to complete registration:
- Finish: Register this nucleotide sequence and exit.
- Finish and translate protein: Both register this nucleotide sequence and register the corresponding protein. This option will take you to the registry wizard for a new protein, prepopulating it with the protein sequence based on the nucleotide sequence you just defined.
Related Topics
Register Protein Sequences
- Via the nucleotide sequence wizard. When registering a nucleotide sequence, you have the option of continuing on to register the corresponding protein sequence.
- Via the header bar. Select Registry > Protein Sequences.
- Select Add > Add manually.
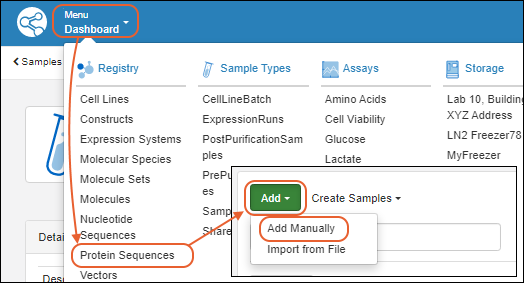
Protein Sequence Wizard
The wizard for registering a new protein sequence proceeds through five tabs:Details
- Name: Provide a name, or one will be generated for you. Hover to see the naming pattern
- Description: (Optional) A text description of the sequence
- Alias: (Optional) List one or more aliases. Type a name, click enter when complete. Continue to add more as needed.
- Organisms: (Optional) Start typing the organism name to narrow the pulldown menu of options. Multiple values are accepted.
- Protein Sequence Parents: (Optional) List parent component(s) for this sequence. Start typing to narrow the pulldown menu of options.
- Seq Part: (Optional) Indicates this sequence can be used as part of a larger sequence. Accepted values are 'Leader', 'Linker', and 'Tag'. When set, chain format must be set to 'SeqPart'.
Sequence
On the sequence tab, you can translate a protein sequence from a nucleotide sequence as outlined below. If you prefer to manually enter a protein sequence from scratch click Manually add a sequence at the bottom.- Nucleotide Sequence: (Optional) The selection made here will populate the left-hand text box with the nucleotide sequence.
- Translation Frame: (Required). The nucleotide sequence is translated into the protein sequence (which will be shown in the right-hand text box) by parsing it into groups of three. The selection of translation frame determines whether the first second or third nucleotide in the series 'heads' the first group of three. Options: 1,2,3.
- Sequence Length: This value is based on the selected nucleotide sequence.
- Nucleotide Start: This value is based on the nucleotide sequence and the translation frame.
- Nucleotide End: This value is based on the nucleotide sequence and the translation frame.
- Translated Sequence Length: This value is based on the nucleotide sequence and the translation frame.
- Protein Start: Specific the start location of the protein to be added to the registry.
- Protein End: Specific the end location of the protein to be added to the registry.
Annotations
The annotations tab displays any matching annotations found in the annotation library. You can also add annotations manually at this point in the registration wizard.- Name: a freeform name
- Type: for example, Leader, Variable, Tag, etc. Start typing to narrow the menu options.
- Category: 'Feature' or 'Region'
- Description: (Optional)
- Start and End Positions: 1-based offsets within the sequence
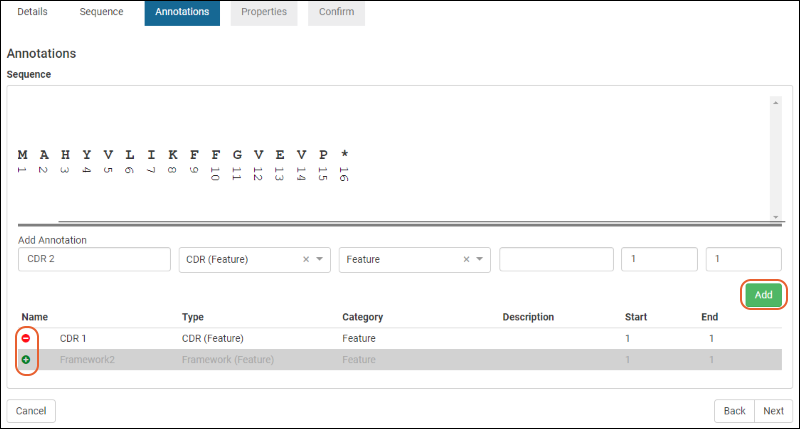 For complete details on using the annotation panel see Protein Sequence Annotations.Click Next to continue the wizard.
For complete details on using the annotation panel see Protein Sequence Annotations.Click Next to continue the wizard.
Properties
- Chain Format: select a chain format from the dropdown (start typing to filter the list of options). An administrator defines the set of options on the ChainFormats list. LabKey Biologics will attempt to classify the protein's chain format if possible.
- ε: the extinction coefficient
- Avg. Mass The average mass
- Num. S-S The number of disulfide bonds
- pI The isoelectric point
- Num Cys. The number of cysteine elements
Confirm
The Confirm panel provides a summary of the protein about to be added to the registry.Click Finish to add the protein to the registry.Editing Protein Sequence Fields
Once you have defined a protein sequence, you can locate it in a grid and click the name to reopen to see the details. Some fields are eligible for editing. Those that are "in use" by the system or other entities cannot be changed. All edits are logged.Related Topics
Register Leaders, Linkers, and Tags
- From the main menu, click Protein Sequences.
- Select Add > Add Manually.
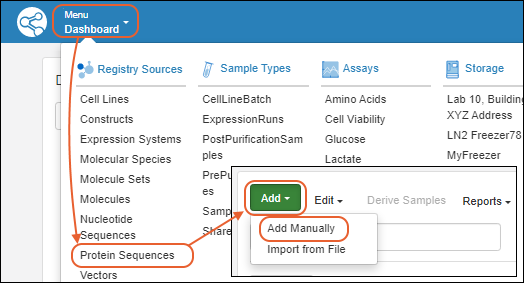
- This will start the wizard Register A New Protein Sequence.
- Leader
- Linker
- Tag
- Click Next.
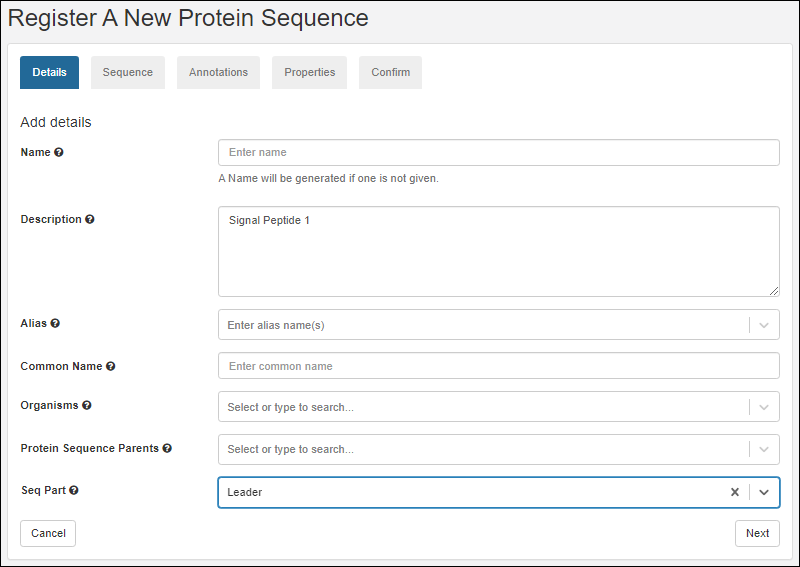
- On the Sequence tab, scroll down and click Manually add a sequence.
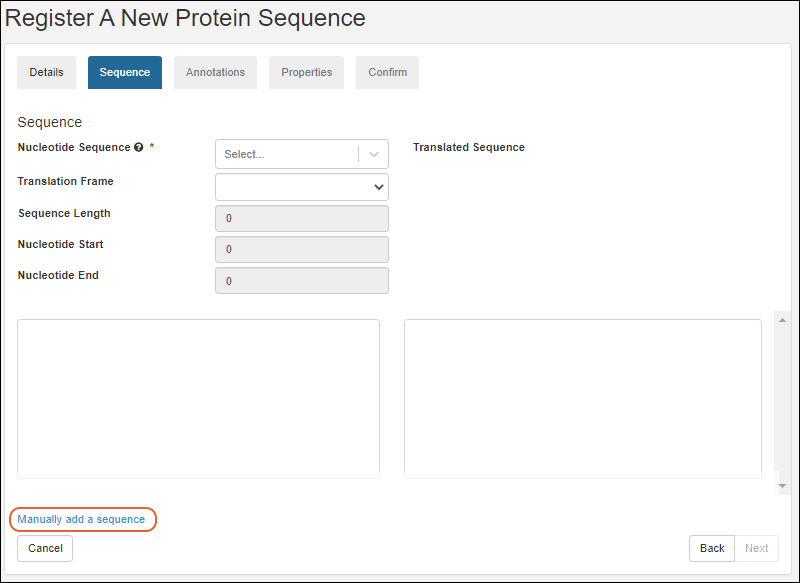
- Enter the sequence and click Next.
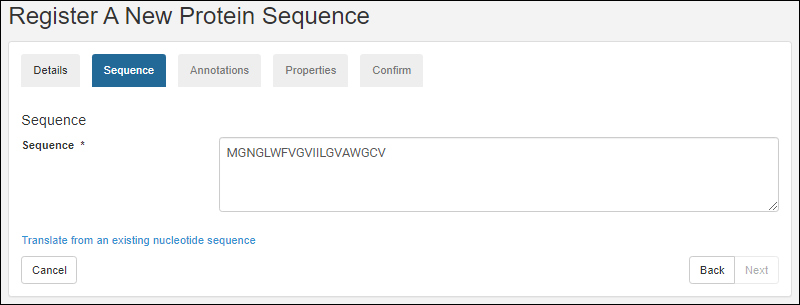
- Complete the rest of the wizard as described in the topic Register Protein Sequences.
Related Topics
Vectors, Constructs, Cell Lines, and Expression Systems
Definitions
- Vectors are typically plasmids that can inject genetic material into cells. Must have a specific nucleotide sequence.
- Constructs are Vectors which have been modified to include the genetic material intended for injection into the cell.
- Cell Lines are types of cells that can be grown in the lab.
- Expression Systems are cell lines that have been injected with a construct.
Add Manually within the User Interface
Select the desired entity from the main menu, then select Add > Add Manually.| Entity Type | Default Fields |
|---|---|
| Vector | Name Common Name Description Alias Sequence Selection Methods Vector Parents |
| Construct | Name Common Name Description Alias Vector Cloning Site Complete Sequence Insert Sequences Construct Parents |
| Cell Lines | Name Common Name Description Alias Expression System Stable Clonal Organisms Cell Line Parents |
| Expression Systems | Name Common Name Description Alias Host Cell Line Constructs Expression System Parents |
Add Manually from Grid
Select the desired entity from the main menu, then select Add > Add Manually from Grid. Learn about creating entities with this kind of grid in the topic:Import from File
Creation of many entities of a given type can also be done in bulk via file import using a template for assistance. Select Add > Import from File and upload the file. Learn more in this topic:Related Topics
Registry Reclassification
Reclassify Protein Sequence
From the Protein Sequences dashboard, select the protein sequence you wish to reclassify. Select Manage > Reclassify.Update Annotations
On the first page of the reclassification wizard, you will see the current Chain Format and Annotations for the sequence.- Review any updates to the Annotations that have been introduced since this protein sequence was last classified. These will be highlighted in green as shown below this list.
- Review the Chain Format assignment that may have been recalculated since the original classification. If needed, you can select another Chain Format.
- Add additional Annotations. Complete the required elements of the Add Annotation section, then click Add.
- Delete previous annotations using the in the "Delete" column. You'll see the deleted annotation struck out and have the option to re-add it by clicking the before continuing.
Adjust Molecules
After reviewing and adjusting the annotations, click Next to continue to the Molecules reclassification step.View Classification Status from Molecule
Starting from the overview page for a Molecule, you can see the Classification Status in the Details panel.Related Topics
Biologics: Terminology
- Definitions: "Registry Source" vs "Sample"
- Diagram of Registry Source Relationships
- Details of Each Registry Source Type
Definitions: "Registry Source" vs "Sample"
In LabKey Biologics LIMS, the terms "registry" and "samples" refer to different components or aspects of the system. The Registry is a database or collection of information about biologics entities, while Samples refer to the actual biological material or representations of it that are being managed within the system. The registry helps organize and provide context to the information about these entities, and samples are the tangible or digital representations linked to these entries in the registry.Registry Sources:- "Registry" typically refers to a comprehensive database or collection of information related to biologics entities. This can include details about biologics, such as antibodies, cell lines, or other biological molecules, that are being studied or managed within the system. The registry serves as a centralized repository for metadata and information about these biological entities, "Registry Sources", often including details like names, identifiers, descriptions, origin, properties, and associated data.
- "Samples" in LabKey Biologics usually refer to the physical or virtual representations of biological material that have been collected, stored, and are being managed within the system. Samples can include a variety of biological materials such as tissues, cell lines, fluids, or other substances. Each sample is typically associated with specific information, like collection date, source, location, processing history, and other relevant details. Samples are often linked to entries in the registry to provide a clear relationship between the biological entity and the actual physical or virtual sample associated with it.
Diagram of Registry Source Relationships
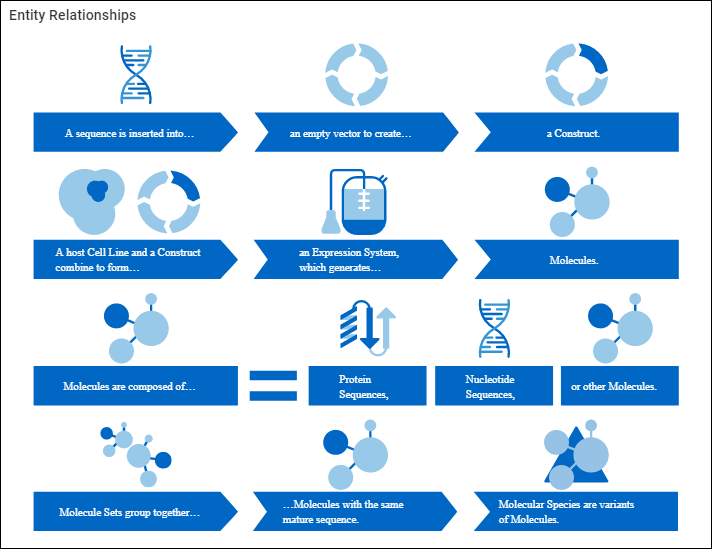
- A Sequence is inserted into an empty
- Vector to create a
- Construct.
- A host Cell Line and a Construct combine to form an
- Expression System, which generates
- Molecules.
- Molecules are composed of:
- Protein Sequences
- Nucleotide Sequences
- and/or other molecules.
- Molecule Sets group together molecules with the same mature sequence.
- Molecular Species are variants of molecules.
Molecule
Composed of a mixture of protein sequences, nucleotide sequences, chemistry elements, and other molecules. Generally, "molecule" refers to the target entity.| Example Molecules |
|---|
| Molecule W = 1 (protein sequence A) |
| Molecule X = 1 (protein sequence B) + 2 (protein sequence C) |
| Molecule Y = 1 (nucleotide sequence D) |
| Molecule Z = 2 (nucleotide sequence E) + 2 (protein sequence F) + 2 (molecule W) |
Molecule Set
The set of molecules that only differ from one another in terms of their leader sequences. See Vectors, Constructs, Cell Lines, and Expression Systems.Molecular Species
After protein expression and other processes, all the entities that are detected for a particular molecule (different cleavage sites, post-translational modifications, genomic drift). See Vectors, Constructs, Cell Lines, and Expression Systems.Protein Sequence
Single sequence comprised of amino acids (20 different amino acids).| Example Protein Sequences |
|---|
| protein sequence A = ACELKYHIKL CEEFGH |
| protein sequence B = HIKLMSTVWY EFGHILMNP |
Nucleotide Sequence
A single sequence comprised of nucleic acids, can be either DNA (A, T, G, C) or RNA (A, U, G, C).| Example Nucleotide Sequences |
|---|
| nucleotide sequence A (DNA) = AGCTGCGTGG GCACTCACGCT |
| nucleotide sequence B (RNA) = AGCUGUUGCA GCUUACAUCGU |
Vector
DNA molecule used as a vehicle to artificially carry foreign genetic material into another cell. The most common type of vector is a plasmid - a long double-stranded section of DNA that has been joined at the ends to circularize it. In molecular biology, generally, these plasmids have stretches of DNA somewhere in their makeup that allow for antibiotic resistance of some sort, providing a mechanism for selecting for cells that have been successfully transfected with the vector.Construct
A vector (generally a plasmid) that has had a stretch of DNA inserted into it. Generally, this DNA insertion encodes for a protein of interest that is to be expressed.Cell Line
A host cell line is transfected with a Construct, bringing the new DNA into the machinery of the cell. These transfected cells (called an Expression System) are then processed to select for cells that have been successfully transfected and grown up, allowing for the production of cells that have the construct and are manufacturing the protein of interest. At times, these transfections are transient and all of the cells are used in the process. Other times, a new stable cell line (still a Cell Line) is produced that can continue to be used in the future.Expression System
A combination of a Construct and a host cellRelated Topics
Protein Sequence Annotations
Annotation Detail Page
To view protein sequence annotations, go to Registry > Protein Sequences, and click a protein id in the Name column. Click the Sequence tab.- The upper panel represents the sequence graphically, with different annotations called out with colored bars.
- The panel below provides annotation details in a grid view.
- The bottom left panel provides an editor with two tabs for adding and editing annotations.
- The bottom right panel offers controls of the display area
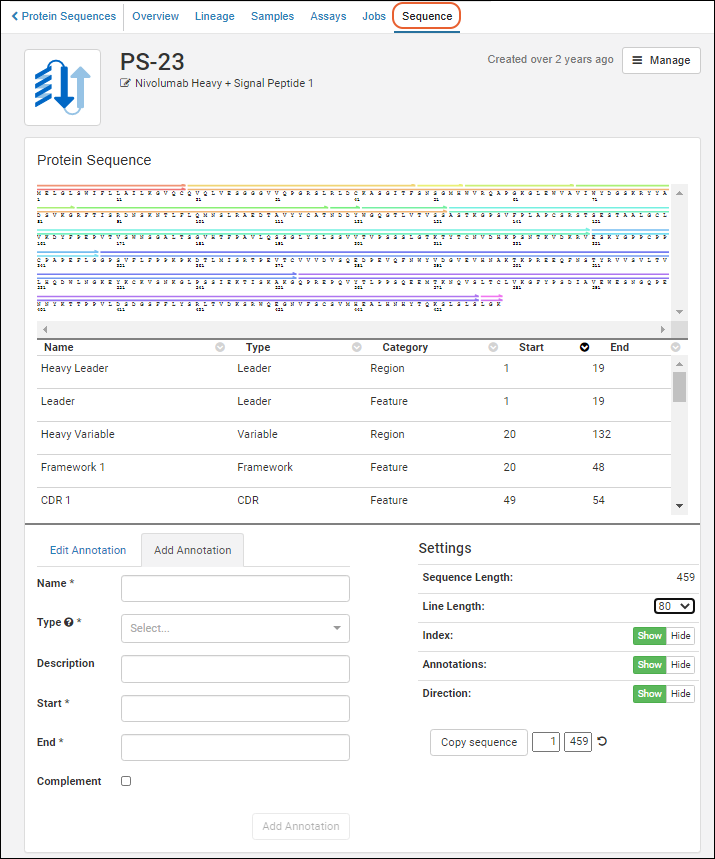 A somewhat simplified version of the annotations viewer is also used within the protein sequence registration wizard.
A somewhat simplified version of the annotations viewer is also used within the protein sequence registration wizard.
Graphical Representation
Clicking an annotation’s color bar highlights its description in the grid below, For example, if you click the green bar for sequence position 24-34, that annotation’s details are highlighted in the grid below, and vice versa.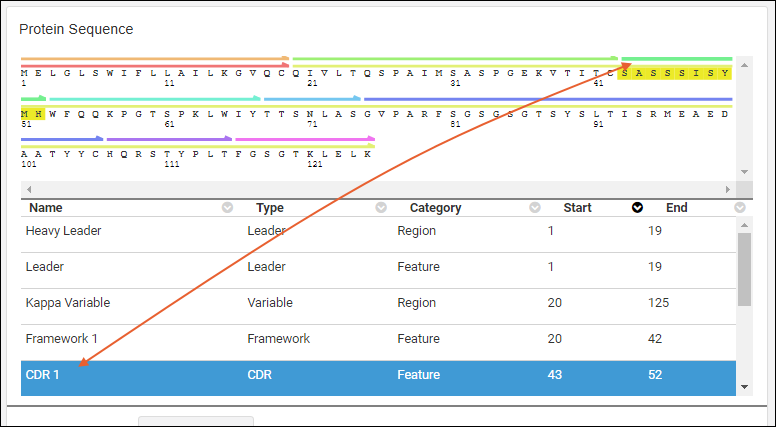
Details Grid
The annotations details grid can be sorted by any of the available columns. The grid will always default to sorting the "Start" column in ascending order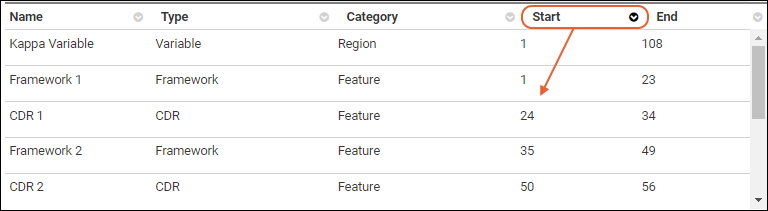
Annotation Editor
The annotation editor has two tabs: Edit Annotation and Add Annotation. Adding or editing an annotation will refresh the grid and viewer to include your changes.- Note the asterisks marking required fields on the Add Annotation tab. The button at the bottom of the form will remain grayed out until all fields have been filled out.
- The Type dropdown is pre-populated with the items in the list "AnnotationTypes".
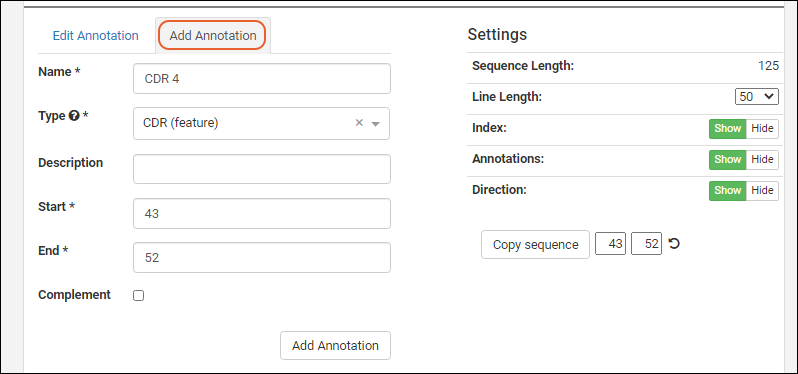 In order to edit an annotation, select one from the grid or viewer and click the Edit Annotation tab.
In order to edit an annotation, select one from the grid or viewer and click the Edit Annotation tab.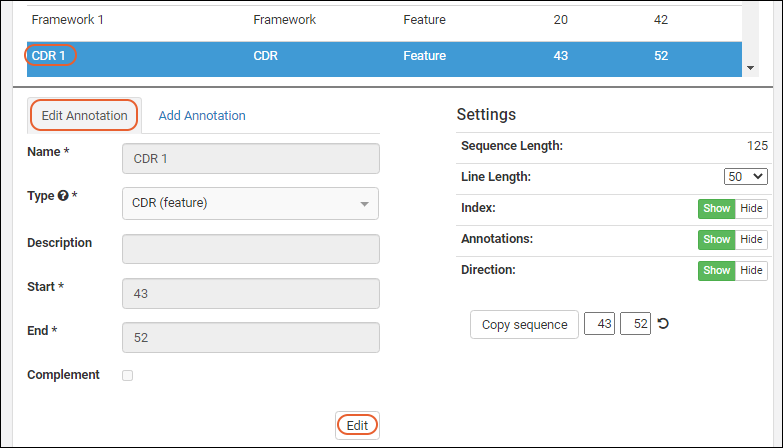 Click the Edit button to enable the field inputs and action buttons:
Click the Edit button to enable the field inputs and action buttons:
- Remove Annotation: Removes the annotation and deletes the details row from the grid.
- Cancel: Cancels the edit.
- Save: Save any changes to the selected annotation.
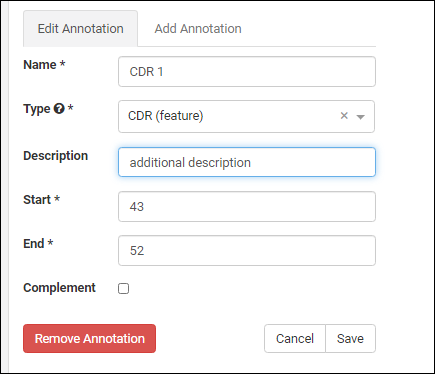
Add AnnotationType
To add a new option to the Annotation Type dropdown, such as "Protease Cleavage Site", switch to the LabKey Server interface and edit the "AnnotationTypes" list. Insert a new row, specifying the name, abbreviation and category you want to use.Settings/Controls
The controls in the bottom right panel let you customize the display region.- Sequence Length: The length of the entire sequence.
- Line Length: Select to control the display scale. Options: 50, 60, 80, 100.
- Index: Show or hide or show the position index numbers.
- Annotations: Show or hide the annotation color bars.
- Direction: Show or hide the direction indicators on the bars.
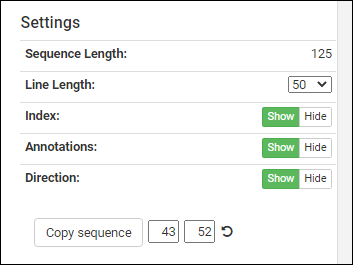 Below these controls, you can see a sequence selection widget next to the Copy sequence button. The position boxes for start and end update to the currently selected annotation. These can be manually changed to any index values in the sequence, or can be reset to the full sequence by pressing the refresh icon.Clicking Copy sequence will copy the indicated segment to your clipboard for pasting (useful when registering a new protein sequence into the Registry).
Below these controls, you can see a sequence selection widget next to the Copy sequence button. The position boxes for start and end update to the currently selected annotation. These can be manually changed to any index values in the sequence, or can be reset to the full sequence by pressing the refresh icon.Clicking Copy sequence will copy the indicated segment to your clipboard for pasting (useful when registering a new protein sequence into the Registry).
Related Topics
CoreAb Sequence Classification
- CoreAb Sequence Classification Introduction
- Classification Process
- Reference Germline Data Extraction Process
CoreAb Sequence Classification: White Paper Introduction
This methodology was published in March 2020, and is also available here:- https://just-evotecbiologics.com/white-papers-application-notes/coreab-sequence-classification/
- Author: J. Alex Taylor
- Publication: March 2020
Keywords: antibody; antibody numbering; structural numbering; antibody engineering; antibody analysis
Classification Process
The CoreAb Java library (developed at Just - Evotec Biologics) contains algorithms for the classification and alignment of antibodies and antibody-like sequences. A high-level summary of the classification process is presented in Figure 1. The first step in the classification process is the detection of antibody variable and constant regions specified in the detection settings. The default regions for detection are kappa variable, lambda variable, heavy variable, light constant, heavy constant Ig (CH1), heavy constant Fc-N (CH2), and heavy constant Fc-C (CH3). A Position-Specific Sequence Matrix (PSSM) has been pre-built for each type and is used as a low threshold first pass filter for region detection using the Smith-Waterman algorithm to find local alignments. Each local alignment is then refined by a more careful alignment comparison to the germline gene segments from species specified in the detection settings. If germline data for the query’s species of origin does not exist or is incomplete in the resource files contained in CoreAb, other, more complete, germline gene data sets from other species can be used to identify homologous regions. The germline sequences are stored as ASN-aligned so that the resulting region alignments are also ASN-aligned.To generate an alignment for variable regions, the PSSM-matched sub-sequence is aligned to both germline V-segments and J-segments and these results are combined to synthesize an alignment for the entire variable region. For heavy variable regions, the germine D-segments are aligned to the residues between the V-segment match and the J-segment match. As a final step in the variable region alignment refinement process, CDR regions are center-gapped to match the AHo/ASN numbering system. Fig. 1 High-level antibody classification pseudocode1. Identify antibody variable and constant regions (domains)
a. Loop over the region types that were specified in settings
i. Use a PSSM for the region type to find local alignments in the query
ii. Loop over each local alignment from the query
1. Loop over the germline sets that were specified in settings
a. Generate a refined region alignment for the PSSM alignment
i. Only keep alignments that meet the minimum number of identities and
region percent identity specified in settings
ii. Assign ASN numbering
iii. If variable region, refine the alignment and adjust CDR gapping
iv. If constant region and alignment is < 10 aa, toss it unless it is at
the start of the region
2. Identify potential leader region matches (can use SeqParts)
3. Resolve overlapping regions giving priority to the higher scoring region
4. Assign gaps between identified regions (can use SeqParts)
5. Cleanup constant regions
6. Assign chain and structure format (based on the arrangement of regions)
<AbFormatCache version='2'>
<ChainFormats>
...
<ChainFormat id='55' name='DVD-IgG1 Heavy Chain' abbrev='DVD-IgG HC' description='DVD-IgG1 Heavy Chain'>
{Ldr} ; HV ; Lnk ; HV ; IgG1:HCnst-Ig ; IgG1:Hinge ; IgG1:Fc-N ; IgG1:Fc-C ; IgG1:HCnst-Po
</ChainFormat>
<ChainFormat id='56' name='IgG1 Fc-Fusion' abbrev='IgG1 Fc-Fusion' description='IgG1 Fc-Fusion'>
{Ldr} ; Unk ; {Lnk}; IgG1:Hinge ; IgG1:Fc-N ; IgG1:Fc-C ; IgG1:HCnst-Po
</ChainFormat>
<ChainFormat id='57' name='IgG1 Heavy Chain F(ab')2' abbrev='IgG1 HC F(ab')2' description='IgG1 Heavy Chain F(ab')2'>
{Ldr} ; HV ; IgG1:HCnst-Ig ; IgG1:Hinge<C113,C116>
</ChainFormat>
...
</ChainFormats>
<StructureFormats>
...
<StructureFormat id='28' name='DVD-IgG1' abbrev='DVD-IgG1' description='Dual Variable Domain (DVD) Bispecific IgG1 Antibody'>
<ChainCombination>
<ChainFormat id='53' stoichiometry='2' />
<ChainFormat id='55' stoichiometry='2' />
</ChainCombination>
<ChainCombination>
<ChainFormat id='54' stoichiometry='2' />
<ChainFormat id='55' stoichiometry='2' />
</ChainCombination>
</StructureFormat>
<StructureFormat id='29' name='IgG1 Fc-Fusion' abbrev='IgG1 Fc-Fusion' description='IgG1 Fc-Fusion'>
<ChainCombination>
<ChainFormat id='56' stoichiometry='2' />
</ChainCombination>
</StructureFormat>
<StructureFormat id='30' name='IgG1 F(ab')2' abbrev='IgG1 F(ab')2' description='IgG1 F(ab')2 Fragment'>
<ChainCombination>
<ChainFormat id='5' stoichiometry='2' />
<ChainFormat id='57' stoichiometry='2' />
</ChainCombination>
<ChainCombination>
<ChainFormat id='8' stoichiometry='2' />
<ChainFormat id='57' stoichiometry='2' />
</ChainCombination>
</StructureFormat>
...
</StructureFormats>
</AbFormatsCache>
Reference Germline Data Extraction Process
The extraction and compilation of antibody germline gene data can be a difficult and time consuming process. In cases where gene annotation is provided by the NCBI, a CoreAb tool is used to extract and align the gene information. Incomplete or unannotated genomes require a more de novo approach. CoreAb also contains a tool that can scan for potential V-segments, J-segments, and D-segments using PSSMs designed to locate the Recombination Signal Sequence (RSS) sequences used to join the variable region segments. Manual curation is still required to filter and adjust the results but this automation can alleviate most of the tedious work. When possible, names for extracted genes are set to those from IMGT since that is the source of official naming. Figure 3 displays a section of an XML-formatted germline data resource file. Default XML-formatted germline data is included in CoreAb and loaded at runtime. Additional or alternate germline data can be provided by the user. Full or partial antibody gene data is currently included in CoreAb for the following organisms: Bos taurus, Camelus bactrianus, Camelus dromedarius, Canis familiaris, Cavia porcellus, Gallus gallus, Homo sapiens, Macaca mulatta, Mus musculus, Oryctolagus cuniculus, Ovis aries, Protopterus dolloi, Rattus norvegicus, Struthio camelus, and Vicugna pacos. Fig. 3 Snippet of the Bos taurus HV.xml germline data file of heavy variable genes extracted from genomic sequences.<?xml version='1.0' encoding='UTF-8' standalone='yes' ?>
<GermlineGeneSetRsrc igGeneGroup='IGHV' taxonId='9913'>
...
<GermlineGene id='IGHV1-20*01'>
<Note>Num RSS mismatches: 0</Note>
<GenomicLoc build='ARS-UCD1.2' chromosome='21' contig='NW_020190105.1' contigLength='69862954'
strand='Forward'>join(278220..278274,278353..278658)</GenomicLoc>
<DB_Xref db='Just-TempId'>IGHV1-4S*01</DB_Xref>
<DB_Xref db='IMGT/GENE-DB'>IGHV1-20*01</DB_Xref>
<Intron donorSite='a gtgtc' acceptorSite='acag g' seqLength='78'>
<GenomicLoc build='ARS-UCD1.2' chromosome='21' contig='NW_020190105.1' contigLength='69862954'
strand='Forward'>278275..278352</GenomicLoc>
<DNA>
gtgtctctgggtcagacatgggcacgtggggaagctgcctctgagcccacgggtcaccgtgcttctctctctccacag
</DNA>
</Intron>
<AlignedRegions>
<AlignedRegion region='HLdr'>
<Protein>
MNPLWEPPLVLSSPQSGVRVLS
</Protein>
<DNA>
atgaacccactgtgggaacctcctcttgtgctctcaagcccccagagcggagtgagggtcctgtcc
</DNA>
</AlignedRegion>
<AlignedRegion region='HV'>
<Protein>
QVQLRES-GPSLVKPSQTLSLTCTVSG-FSLSS-----YAVGWVRQAPGKALEWLGGISS----GGSTYYNPALKSRLSITKDNSKSQVSLSVSSVTPEDTATYYCAK-----------------------------------------
</Protein>
<DNA RSS_3prime='cacagtg aggggaaatcagtgtgagcccag acaaaaacc' splitCodon3prime='ga'>
caggtgcagctgcgggagtcgggccccagcctggtgaagccctcacagaccctctccctcacctgcacggtctctggattctcactgagcagctatgctgtaggctgggtccgccaggctccagggaaggcgctggagtggctcggtggtataagcagtggtggaagcacatactataacccagccctgaaatcccggctcagcatcaccaaggacaactccaagagccaagtctctctgtcagtgagcagcgtgacacctgaggacacggccacatactactgtgcgaagga
</DNA>
</AlignedRegion>
</AlignedRegions>
</GermlineGene>
...
</GermlineGeneSetRsrc>
Related Topics
Biologics: Chain and Structure Formats
Overview
After the classification engine has determined the pattern of regions within the sequence, the chain and structure format information that was provided is used to first match the regions to a chain pattern and then to match the assigned chain formats to a unique structure format.If antibody regions are present in a ProteinSeq, but it does not match a chain format, it will be assigned a chain format of "Unrecognized Antibody Chain" and the Molecule is assigned a structure format of "Unrecognized Antibody Format".If no antibody regions are present, the ProteinSeq is assigned a chain format of "Non-Antibody Chain" and the Molecule structure format is set to "Non-Antibody".In order for changes to the chain and structure formats to take effect, an administrator needs to clear caches from the Administration Console.Chain Formats
Chain formats are stored in the ChainFormats List.A chain format specification is specified at the region level using region abbreviations. Recognized regions are listed in the following table.| Region | Abbreviation |
|---|---|
| Leader | Ldr |
| Light Variable | KV/LmdV |
| Light Constant | KCnst/LmdCnst |
| Kappa Leader | KLdr |
| Kappa Variable | KV |
| Kappa Constant Ig Domain | KCnst-Ig |
| Post Kappa Constant Ig Present in some species as a short C-terminal tail | KCnst-Po |
| Lambda Leader | LmdLdr |
| Lambda Variable | LmdV |
| Lambda Constant Ig Domain | LmdCnst-Ig |
| Post Lambda Constant Ig Present in some species as a short C-terminal tail | LmdCnst-Po |
| Heavy Leader | HLdr |
| Heavy Variable | HV |
| Heavy Constant Ig Domain | HCnst-Ig |
| Heavy Constant Hinge | Hinge |
| Heavy Constant Fc N-terminal Domain | Fc-N |
| Heavy Constant Fc C-terminal Domain | Fc-C |
| Post Heavy Constant Ig | HCnst-Po |
| Linker | Lnk |
| Tag | Tag |
| Protease Cleavage Site | Cut |
| Unrecognized | Unk |
| TCR-alpha Leader | TRALdr |
| TCR-alpha Variable | TRAV |
| TCR-alpha Constant Ig Domain | TRACnst-Ig |
| TCR-alpha Constant Connector | TRACnst-Connector |
| TCR-alpha Constant Transmembrane Domain | TRACnst-TM |
| TCR-beta Leader | TRBLdr |
| TCR-beta Variable | TRBV |
| TCR-beta Constant Ig Domain | TRBCnst-Ig |
| TCR-beta Constant Connector | TRBCnst-Connector |
| TCR-beta Constant Transmembrane Domain | TRBCnst-TM |
| TCR-delta Leader | TRDLdr |
| TCR-delta Variable | TRDV |
Chain Format Syntax
1. Regions are separated by a semicolon. Optional regions are surrounded by braces {}.Example 1: A kappa light chain is specified as:{Ldr} ; KV ; KCnst-Ig ; {KCnst-Po}{Ldr | Unk<M>} ; (HV ; Lnk ; KV/LmdV | KV/LmdV ; Lnk ; HV){Ldr} ; HV ; IgG2:HCnst-Ig ; IgG2:Hinge ; IgG2:Fc-N ; IgG2:Fc-C ; IgG2:HCnst-Po{Ldr} ; HV ; IgG1:HCnst-Ig ; IgG1:Hinge<!113-123>{Ldr} ; HV ; IgG1:HCnst-Ig ; IgG1:Hinge ; IgG1:Fc-N ; IgG1:Fc-C<C15,W30> ; IgG1:HCnst-Po{Ldr} ; HV[3] ; IgG1:HCnst-Ig ; Unk<EPKSCD> ; Lnk ; HV[2] ; Unk<A> ; KCnst/LmdCnst ;
IgG1:Hinge<!1-107> ; IgG1:Fc-N ; IgG1:Fc-C<C15,W30> ; IgG1:HCnst-Po{Ldr} ; HV⊃VHH ; IgG1:Hinge ; IgG1:Fc-N ; IgG1:Fc-C ; IgG1:HCnst-PoStructure Formats
Structure formats are stored in the StructureFormats List where the only important information is the name and abbreviation. The more important table is ChainStructureJunction which describes the chain combinations that map to a structure format.For example, there are two chain combinations for an IgG1, 2 copies of a Kappa Light Chain + 2 copies of an IgG1 Heavy Chain or 2 copies of a Lambda Light Chain + 2 copies of an IgG1 Heavy Chain. In the ChainStructureJunction this is represented like this:| Structure Format | Chain Format | Combination | Stoichiometry | Num Distinct* | Fv Num Overrides** |
|---|---|---|---|---|---|
| IgG1 | Kappa Light Chain | 1 | 2 | 1 | |
| IgG1 | IgG1 Heavy Chain | 1 | 2 | 1 | |
| IgG1 | Lambda Light Chain | 2 | 2 | 1 | |
| IgG1 | IgG1 Heavy Chain | 2 | 2 | 1 |
Related Topics
Molecules, Sets, and Molecular Species
Definitions
Molecules are composed of various components and have a set of physical properties that describe them. When a molecule is added to the registry, the following additional entities are calculated and added, depending on the nature of the molecule. These additional entities can include:- Molecular Species
- Molecule Sets
- Other Protein Sequences
Molecule Components
A molecule can be created (= registered) based on one or more of the following:- a protein sequence
- a nucleotide sequence
- other molecules
Rules for Entity Calculation/Creation
When the molecule contains only protein sequences, then:- A mature molecular species is created, consisting of the leaderless segments of the protein sequences, provided a leader portion is identifiable. The leader segment has to be:
- Have annotation start with residue #1
- The annotation Type is "Leader"
- The annotation Category is "Region"
- (If no leader portion is identifiable, then the species will be identical to the Molecule which has just been created.)
- Additionally, a mature desK molecular species is created (provided that there are terminal lysines on heavy chains).
- New protein sequences are created corresponding to any species created, either mature, mature des-K, or both. The uniqueness constraints imposed by the registry are in effect, so already registered proteins will be re-used, not duplicated.
- A molecule set is created, provided that the mature molecular species is new, i.e., is not the same (components and sequences) as any other mature molecular species of another molecule. If there is an already existing mature species in the registry, then the new molecule is associated with that set.
- Physical properties are not calculated.
- Molecular species are not created.
- A molecule set is created, which has only this molecule within it.
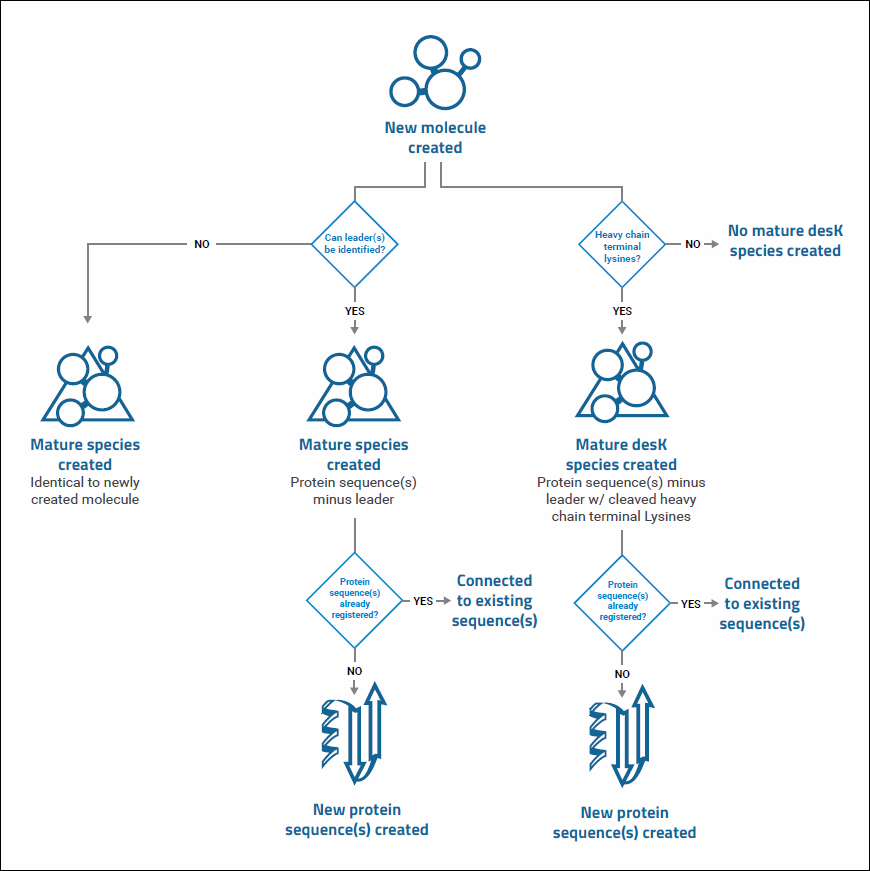
Aliases and Descriptions
When creating a Molecule, the auto-generated Molecule Set (if there is a new one) will have the same alias as the Molecule. If the new Molecule is tied to an already existing Molecule Set, the alias is appended with alias information from the new Molecule.When creating a molecule, the auto-generated molecular species (both mature and mature desK) should have the same alias as the molecule. Similarly, when creating a molecular species from a molecule (manually), the alias field will pre-populate with the alias from the molecule.For molecular species that are auto-generated, if it is creating new protein sequences (one or more) as the components of that molecular species, they have a Description:- Mature of “PS-15”
- Mature, desK of “PS-16”
- Mature of “PS-17”
- Mature, desK of “PS-18”
Related Topics
Register Molecules
Create a Molecule
To add a new molecule to the registry:- Open the Molecules creation wizard.
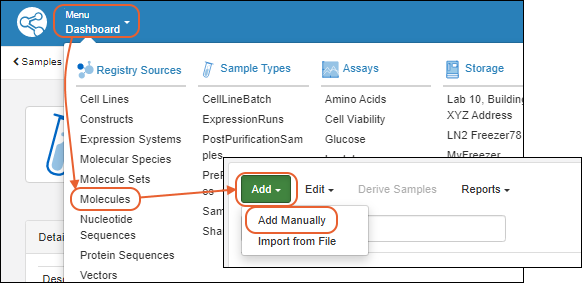
Add Details
On the first tab of the wizard, enter the following:- Name: Provide a name, or one will be generated for you. Hover to see the naming pattern
- Description: (Optional) A text description of the molecule.
- Alias: (Optional) Alternative names for the molecule.
- Common Name: (Optional) The common name of the molecule, if any.
- Molecule Parents: (Optional) Parent molecules for the new molecule.
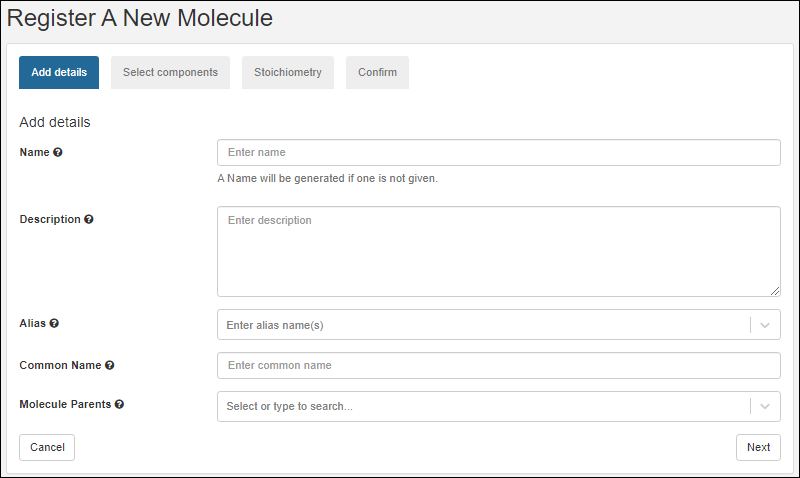 Click Next to continue.
Click Next to continue.
Select Components
- On the Select components tab, search and select existing components of the new molecule.
- After selecting the appropriate radio button, search for the component of interest.
- Type ahead to narrow the list.
- You will see a details preview panel to assist you.
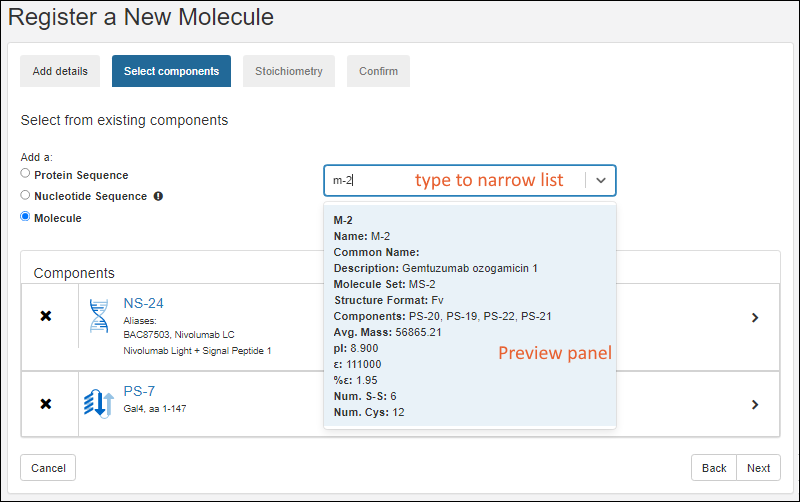
- Once you have added a component, it will be shown as a panel with entity icon. Click the to expand details.
Stoichiometry
LabKey Biologics will attempt to classify the structure format of the molecule's protein components, if possible. The structure format is based on the component protein chain formats.On the Stoichiometry pane enter:- Stoichiometry from each component
- Structure Format: Select a format from the pulldown list. The list is populated from the StructureFormat table.
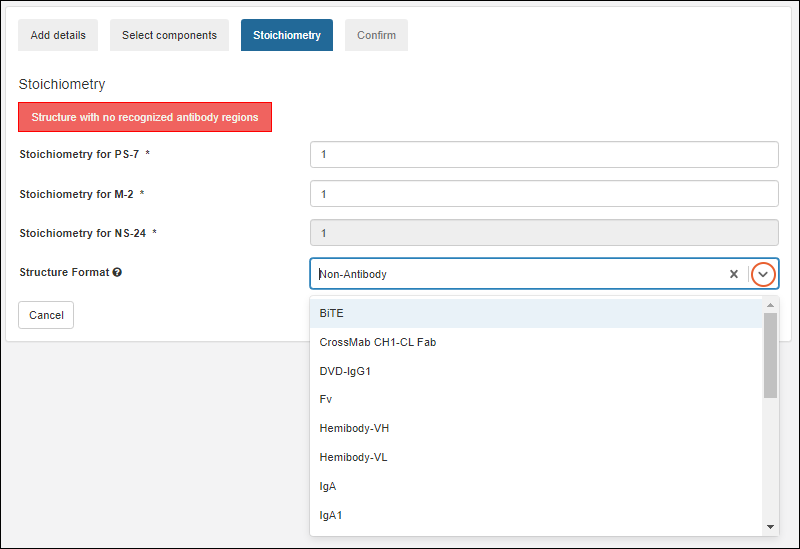 Click Next to continue.
Click Next to continue.
Confirm
On the final tab, confirm the selections and click Finish to add the molecule to the registry.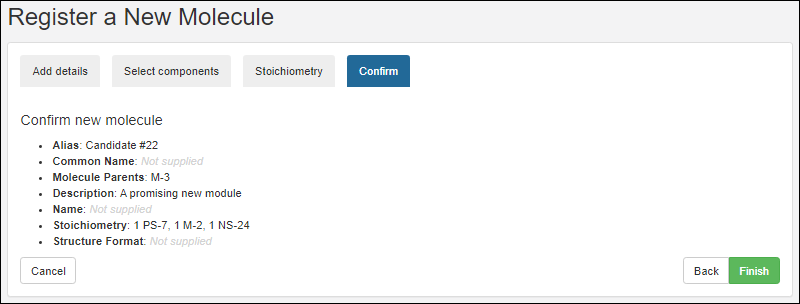 The new molecule will be added to the grid.
The new molecule will be added to the grid.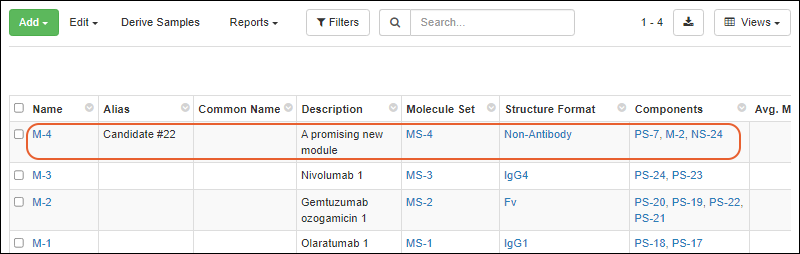
Related Topics
Molecular Physical Property Calculator
View Physical Property Calculator
From the grid of Molecules, select the Molecule of interest. Click the Physical Property Calculator tab.Calculation Inputs
In the Calculation Inputs panel, you'll enter the values to use in the calculation:- Num. S-S: Dynamically adjusted to be half of the "Num. Cys" value.
- Num Cys: A display only value derived from the sequence chosen.
- Sequence Scope: Select the desired scope. Sequence ranges will adjust based on your selection of any of the first three options. Use "Custom Range" for finer control of ranges.
- Full Protein Sequence: complete amino acid sequence of a protein, including all the amino acids that are synthesized based on the genetic information.
- Mature Protein Sequence: final, active form of the protein, after post-translational modifications and cleavages, or other changes necessary for the protein to perform its intended biological function.
- Mature Des-K Protein Sequence: mature protein sequence with the C-Terminal Lysine removed.
- Custom Range
- Sequence Ranges for each component sequence. These will adjust based on the range selected using radio buttons, or can be manually set using the "Custom Range" option.
- Use Range: Enter the start and end positions.
- Click View this sequence to open the entire annotated sequence in a new tab for reference.
- Stoichiometry
- Analysis Mode. Select from:
- Native
- Reduced
- Alkylated Cysteine (only available in "Reduced" mode). If available, select the desired value.
- Modifiers
- Pyro-glu: Check the box for "Cyclize Gln (Q) if at the N-terminus"
- PNGase: Check the box for "Asn (N) -> Asp (D) at N-link sites"
Properties
When you click Calculate, the right hand panel of Properties will be populated. You'll see both what the Classifier Generated value is (if any) and the Simulated value using the inputs you entered.Calculations are provided for:- Mass
- Average Mass
- Monoisotopic Mass
- Organic Average Mass
- pI - Isoelectric Point calculated by different methods:
- Bjellqvist
- EMBOSS
- Grimsley
- Patrickios (simple)
- Sillero
- Sillero (abridged)
- Other
- Chemical Formula
- Extinction Coefficient - ε
- Percent Extinction Coefficient
- Sequence Length
- Amino Acid (AA) composition
Export Calculations
To export the resulting calculated properties, click Export Data.The exported Excel file includes both the calculated properties and the inputs you used to determine them. For example, the export of the above-pictured M-17 calculation would look like this:Related Topics
Compounds and SMILES Lookups
SMILES Lookup Field
The Compound registry source type uses a custom SMILES field type only available in the Biologics module for this specific registry source. This datatype enables users to provide a SMILES string, e.g. "C1=CC=C(C=C1)C=O", that will return a 2D structure image, molecular weight and other computed properties.The SMILES string is used to search the Java library CDK ("Chemistry Development Kit") , a set of open source modular Java libraries for Cheminformatics. This library is used to generate the Structure2D image and calculate masses.Create/Import Compounds
New Compounds can be created manually or via file import. It's easier to get started understanding the lookup process by creating a single compound in the user interface.Create Compound: Carbon Dioxide
As an example, you could create a new compound, supplying the SMILES string "O=C=O" and the Common Name "carbon dioxide".Import File of SMILES Strings
When a file of SMILES strings is Created/imported, each is used to query for the respective 2D structure, molecular weight and set of computed properties. During the Create/Import operation if Name/ID isn’t specified, the SMILES string is used for Name.Related Topics
- Create Registry Sources
- Biologics: Bioregistry
- CDK (Chemistry Development Kit) : Open Source modular Java libraries for Cheminformatics
Entity Lineage
View Lineage
Any parents and children of an Registry Source or Sample can be viewed by clicking the Lineage tab on the details page. The main lineage dashboard shows a graphical panel on the left and details on the right.Two lineage views are available: (1) a graphical tree and (2) a grid representation.
The main lineage dashboard shows a graphical panel on the left and details on the right.Two lineage views are available: (1) a graphical tree and (2) a grid representation.
Lineage Graph
The graphical tree shows parents above and children below the current entity. Click any node in the tree to see details on the right. Hover for links to the overview and lineage of the selected node, also known as the seed.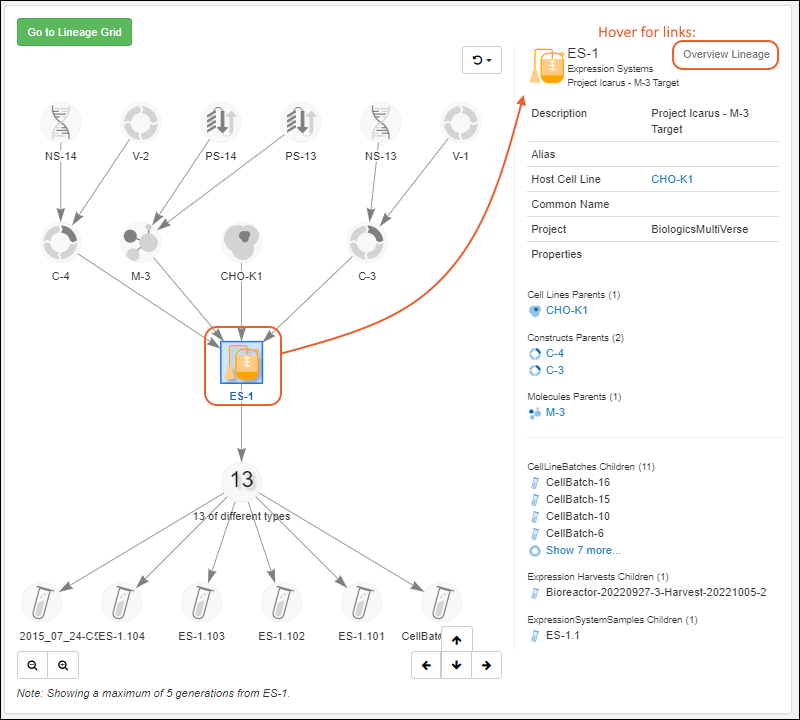
- Zoom out and in with the and buttons in the lower left.
- Step up/down/left and right within the graph using the arrow buttons.
- Up to 5 generations will be displayed. Walk up and down the tree to see further into the lineage.
- Click anywhere in the graph and drag to reposition it in the panel.
- Refresh the image using the button. There are two options:
- Reset view and select seed
- Reset view
Graph Filters for Samples
When viewing lineage for Samples, you will have a filter button. Use checkboxes to control visibility in the graph of derivatives, source parents, and aliquots.Lineage Grid
Click Go to Lineage Grid to see a grid representation of lineage. The grid view is especially helpful for viewing lengthy lineages/derivation histories. Control visibility of parents and children in the grid view using Show Parents and Show Children respectively.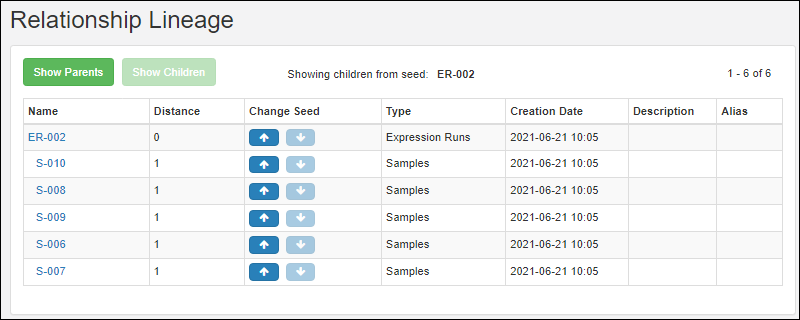 Items in the Name column are clickable and take you to the details page for that entity.Arrow links in the Change Seed column reset the grid to the entity in the Name column, showing you the parents or children for that entity.
Items in the Name column are clickable and take you to the details page for that entity.Arrow links in the Change Seed column reset the grid to the entity in the Name column, showing you the parents or children for that entity.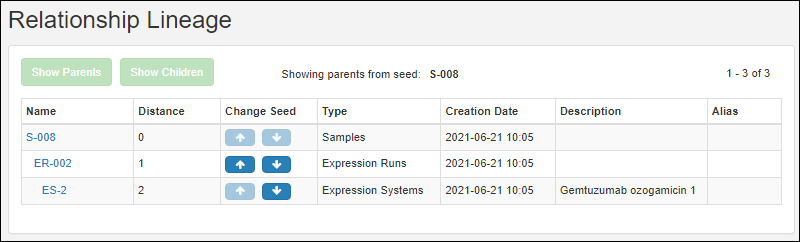
Troubleshooting
Note that if any entity names that just strings of digits, you may run into issues if those "number-names" overlap with row numbers of other entities of that type. In such a situation, when there is ambiguity between name and row ID, the system will presume that the user intends to use the value as the name.Related Topics
Customize the Bioregistry
- Definitions
- Edit Existing Registry Source Types
- Create New Registry Source Type
- Lineage for Custom Registry Source Types
- Delete Custom Registry Source Type
Definitions
The term Registry Source Type applies to all of the following entities in the Biologics application. These are two of the categories of Data Class available in LabKey Server. Within the Biologics application you cannot see or change the category, though if you access these structures outside the application you will be able to see the category.- Registry Source Types: (Category=registry)
- CellLine
- Construct
- ExpressionSystem
- MolecularSpecies
- Molecule
- MoleculeSet
- NucSequence
- ProtSequence
- Vector
- Media Types: (Category=media)
- Ingredients
- Mixtures
Edit Existing Registry Source Types
The default set of Registry Source and Media Types in Biologics are designed to meet a common set of needs for properties and fields for the Bioregistry and Media sections.If you want to customize these designs, you can edit them within the Biologics application following the same interface as used for Sample Manager "Sources".Open the desired type from the main menu. All entries under Registry Sources and Media are editable. Select Manage > Edit [Registry Source Type] Design.Create New Registry Source Type
From the main menu, click Registry Sources. Click Create Source Type.- For example, to use "DC-" followed by the date portion of when the entity is added, plus an incrementing number you could use:
DC-${now:date}-${genId}
Lineage for Custom Registry Source Types
The lineage relationships among the built-in Registry Source Types are pre-defined. Learn more in this topic: Biologics: TerminologyWhen you create your own Custom Registry Source Types, you can include Parent Aliases in the definition, making it possible to define your own lineage relationships.To add a new lineage relationship, either define it when you create the new Registry Source Type or edit it later.Delete Custom Registry Type
You cannot delete the built in Registry Source Types, but if you add a new custom one, you will have the ability to delete it. Before deleting the type, you may want to delete all members of the type. If any members are depended upon by other entities, deletion will be prevented, allowing you to edit to change the connections before deleting the type.- Select the custom type to delete from the Registry section of the main menu.
- Select Manage > Delete [Registry Source Type].
Related Topics
Bulk Registration of Entities
Supported File Formats
The file formats supported are listed in the file import UI under the drop target area.Tabular file formats are supported for all entity types:- Excel: .xls, .xlsx
- Text: .csv, .tsv
- GenBank: .genbank, .gb, .gbk
Assemble Bulk Data
When assembling your entity data into a tabular format, keep in mind that each Registry Source Type has a different set of required column headings.- Acquire a template Excel file for a given entity by clicking Template from the Registry Source Type overview page.
- Open the Excel file, and add data as appropriate under the column headers. See examples below.
Indicate Lineage Relationships
Lineage relationships (parentage) of entities can be included in bulk registration by adding "DataInputs/<DataClassType>" columns and providing parent IDs.For example, to include a Vector as a 'parent' for a bulk registered set of Expression Systems, after obtaining the template for Expression Systems, add a new column named "DataInputs/Vector" and provide the parent vector name for each row along with other fields defining the new Expression Systems.Bulk Upload Registry Source Data
After you have assembled your information into a table, you can upload it to the registry:- Go the Registry Source Type you wish to import.
- Select Add > Import from File.
- On the import page, you can download a template if you don't have one already, then populate it with your data.
- Confirm that the Source Type you want is selected, then drag and drop your file into the target area and click Import.
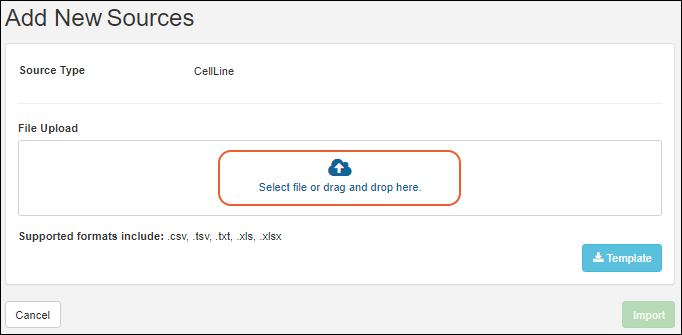
Bulk Data Example Files
Example Nucleotide Sequence File
Notes:- Annotations: Add annotation data using a JSON snippet, format is shown below.
| name | alias | description | flag | protSequences | sequence | annotations |
|---|---|---|---|---|---|---|
| NS-23 | Signal Peptide 1 | An important sequence. | FALSE | [{name: "PS-23"}] | CCCCTCCTTG GAGGCGCGCA ATCATACAAC CGGGCACATG ATGCGTACGC CCGTCCAGTA CGCCCACCTC CGCGGGCCCG GTCCGAGAGC TGGAAGGGCA | [ |
| name | alias | description | flag | protSequences | sequence | annotations |
|---|---|---|---|---|---|---|
| NS-100 | Signal Peptide 1 | some description | FALSE | [{name: "PS-100", nucleotideStart=1, nucleotideEnd=30, translationFrame=2}] | ATGGAGTTGGGACTGAGCTGGATTTTCCTTTTGGCTATTTTAAAAGGTGTCCAGTGT |
Example Protein Sequence File
Notes:- Organisms: A comma separated list of applicable organisms. The list, even if it has only one member, must be framed by square brackets. Examples: [human] OR [human, rat, mouse]
- ?: The column header for the extinction coefficient (ε).
- %?: The column header for the % extinction coefficient (%ε).
| Name | Alias | Description | Nuc Sequences | Chain Format | Avg. Mass | pI | ? | %? | Num. S-S | Num. Cys | Organisms | Sequence |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| PStest-150 | Test sequence for import | 1 | 13999.64 | 8.030 | 35500 | 2.54 | 1 | 2 | [mouse, rat] | EVQLVESGEL IVISLIVESS PSSLSGGLVQ GGGSLRLSCA ASGELIVISL IVESSPSSLS YSFTGHWMNW VRQAPGKGLE WVGIMIHPSD SETRYNQKFK DELIVISLIV ESSPSSLSIR FTISVDKSKN TLYLQMNSLR AEDTAVYYCA RIGIYFYGTT YFDYIWGQGT |
Mixtures and Batches
The text 'unknown' can entered for certain fields. For Mixtures, the Amount field; for Mixture Batches, the Amount and the RawMaterial fields.Mixture Bulk Upload| Type | Ingredient/Mixture | Amount Unit Type |
|---|---|---|
| Ingredient | I-2 | unknown |
| Ingredient | Amount Used | Raw Material Used |
|---|---|---|
| Sodium phosphate dibasic anhydrous | 5 | RawMat-1234 |
| Sodium Chloride | unknown | unknown |
| Potassium chloride | unknown | unknown |
Related Topics
Use the Registry API
Identity Service
When registering a sequence or molecule, use the identity of the sequence to determine uniqueness. External tools can use the following API to get the identity of a sequence prior to registration. To get the identity of a sequence use either the identity/get.api or identity/ensure.api. The ensure.api will create a new identity if the sequence hasn't been added yet.To get or ensure the identity of a single nucleotide sequence:LABKEY.Ajax.request({
url: LABKEY.ActionURL.buildURL("identity", "get.api"),
jsonData: {
items: [{
type: "nucleotide", data: "GATTACA"
}]
}
});LABKEY.Ajax.request({
url: LABKEY.ActionURL.buildURL("identity", "get.api"),
jsonData: {
items: [{
type: "nucleotide", data: "GATTACA"
},{
type: "protein", data: "ELVISLIVES"
}]
}
});LABKEY.Ajax.request({
url: LABKEY.ActionURL.buildURL("identity", "get.api"),
jsonData: {
items: [{
type: "molecule", items: [{
type: "protein", data: "MAL", count: 2
},{
type: "protein", data: "SYE", count: 3
]}
}]
}
});Manual Data Entry
You can enter molecules, sequences, cell lines, etc., using registration wizards within the Biologics application. For an example use of the wizard, see Register Nucleotide SequencesCut-and-Paste or Import from a File
The LabKey import data page can also be used to register new entities. For example, to register new nucleotide sequences:- Go to the list of all entity types (the DataClasses web part).
- Click NucSequence.
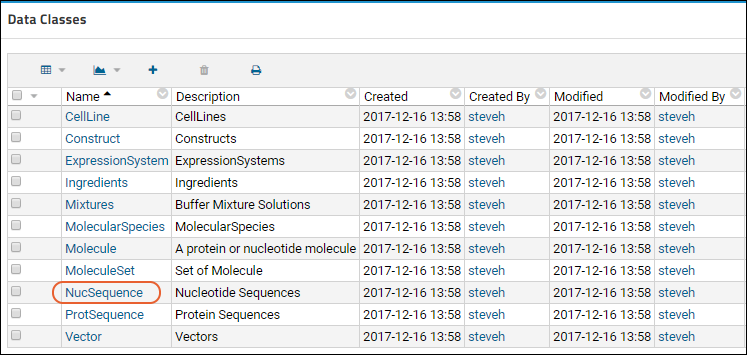
- In the grid, select > Import Bulk Data.
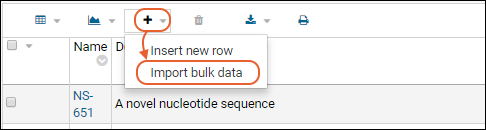
- Click Download Template to obtain the full data format.
- Upload an Excel file or paste in text (select tsv or csv) matching the downloaded template. It might be similar to:
| Description | protSeqId | translationFrame | translationStart | translationEnd | sequence |
|---|---|---|---|---|---|
| Anti_IGF-1 | PS-7 | 0 | 0 | 0 | caggtg... |
- Go to the list of all entity types
- Click ProtSequence
- In the grid, select > Import Bulk Data.
- Click Download Template to obtain a template showing the correct format.
- Paste in a TSV file or upload an Excel file in the correct format. It might be similar to:
| Name | Description | chainFormatId | organisms | sequence |
|---|---|---|---|---|
| PS-1 | PS1038-1 | 1 | ["human","mouse"] | MALWMRL... |
- Go to the list of all entity types
- Click Molecule
- In the grid, select > Import Bulk Data.
- Paste in data of the format:
| Description | structureFormatId | seqType | components |
|---|---|---|---|
| description | 1 | 1 | [{type: "nucleotide", name: "NS-1", stoichiometry: 3}] |
| description | 1 | 3 | [{type: "protein", name: "PS-1", stoichiometry: 2}, {type: "protein", ident: "ips:1234"}] |
| description | 1 | 3 | [{type: "chemical", name: "CH-1"}, {type: "molecule", name: "M-1"}] |
Register via Query API
The client APIs also can be used to register new entities.Register Nucleotide Sequence
From your browser's dev tools console, enter the following to register new nucleotide sequences:LABKEY.Query.insertRows({
schemaName: "exp.data",
queryName: "NucSequence",
rows: [{
description: "from the client api",
sequence: "gattaca"
},{
description: "another",
sequence: "cattaga"
}]
});Register Protein Sequence
To register new protein sequences:LABKEY.Query.insertRows({
schemaName: "exp.data",
queryName: "ProtSequence",
rows: [{
name: "PS-100",
description: "from the client api",
chainFormatId: 1,
sequence: "ML"
}]
});Register Molecule
To register new molecules:LABKEY.Query.insertRows({
schemaName: "exp.data",
queryName: "Molecule",
rows: [{
description: "from the client api",
structureFormatId: 1,
components: [{
type: "nucleotide", name: "NS-202"
}]
},{
description: "another",
structureFormatId: 1,
components: [{
type: "protein", name: "PS-1"
},{
type: "protein", name: "PS-100", count: 5
}]
}]
});Lineage, Derivation, and Samples
Parent/child relationships within an entity type are modeled using derivation. For example, the Lineage page for this nucleotide sequence (NS-3) shows that two other sequences (NS-33 and NS-34) have been derived from it.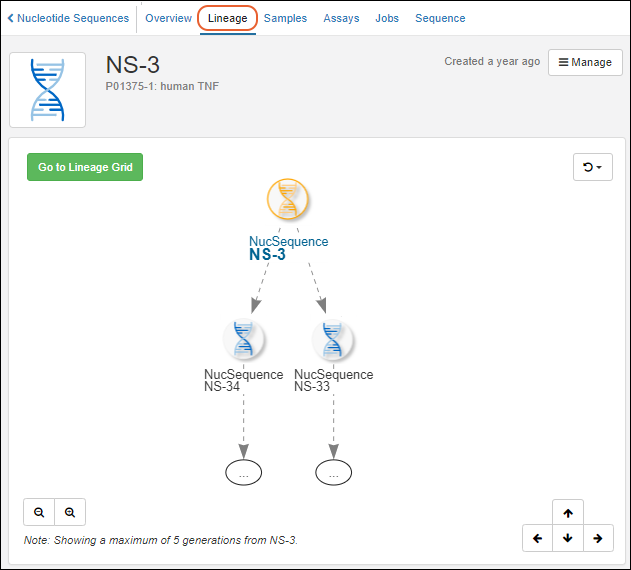 To create new children, you can use the "experiment/derive.api" API, but it is still subject to change. The dataInputs is an array of parents each with an optional role. The targetDataClass is the LSID of the entity type of the derived datas. The dataOutputs is an array of children each with an optional role and a set of values.
To create new children, you can use the "experiment/derive.api" API, but it is still subject to change. The dataInputs is an array of parents each with an optional role. The targetDataClass is the LSID of the entity type of the derived datas. The dataOutputs is an array of children each with an optional role and a set of values.LABKEY.Ajax.request({
url: LABKEY.ActionURL.buildURL("experiment", "derive.api"),
jsonData: {
dataInputs: [{
rowId: 1083
}],
targetDataClass: "urn:lsid:labkey.com:DataClass.Folder-5:NucSequence",
dataOutputs: [{
role: "derived",
values: {
description: "derived!",
sequence: "CAT"
}
}]
}
})LABKEY.Ajax.request({
url: LABKEY.ActionURL.buildURL("experiment", "derive.api"),
jsonData: {
dataInputs: [{
rowId: 1083
}],
targetSampleType: "urn:lsid:labkey.com:SampleType.Folder-5:Samples",
materialOutputs: [{
role: "sample",
values: {
name: "new sample!",
measurement: 42
}
}]
}
})Register Parents/Inputs
To indicate parents/inputs when registering an entity, use the columns "DataInputs/<DataClassName>" or "MaterialInputs/<SampleTypeName>". The value in the column is a comma separated list of values in a single string. This works for both DataClass and SampleType:LABKEY.Query.insertRows({
schemaName: "exp.data",
queryName: "MyDataClass",
rows: [{
name: "blush",
"DataInputs/MyDataClass": "red wine, white wine"
}]
});LABKEY.Query.insertRows({
schemaName: "samples",
queryName: "SamplesAPI",
rows: [{
name: "red wine"
},{
name: "white wine"
},{
name: "blush",
"MaterialInputs/SamplesAPI": "red wine, white wine"
}]
});Related Topics
Biologics: Plates
Plates Dashboard
Click Plates (under Sample Types) on the main menu. You can also go directly to Plate Sets or Plate Templates from here.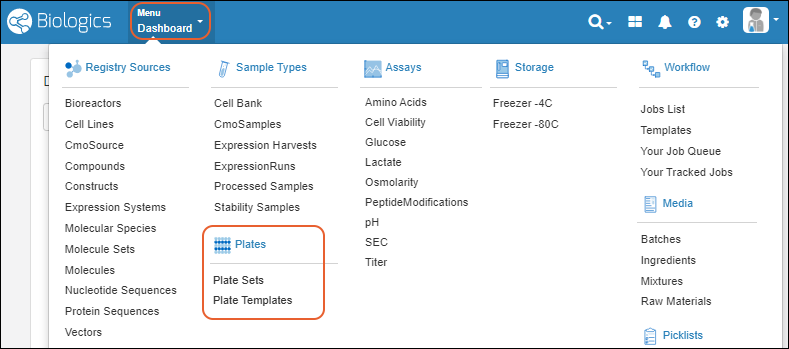
Plate Sets
- Create Plate Set
- Manage > Archive
- Manage > Plate Templates
Plate Templates
Plate templates let you set out some common features and well roles for given campaigns.Related Topics
Biologics: Assay Data
Documentation
Learn more about using the general assay framework in the Sample Manager documentation here:Additional Features with LabKey LIMS
Additional Features with Biologics LIMS
Biologics: Specialty Assays
Create a New Assay Design
To create a new assay design in Biologics LIMS, follow this process. Make sure to start from the container where you want to define the assay.- From the main menu, click Assays, then click Create Assay Design.
Select the Assay Type
- Select the Assay Type. For most cases, leave the Standard Assay tab selected.
- Learn more about using Specialty Assays in this topic: Describe Assay Data Structure.
- Click Choose Standard Assay.
- Creating a standard assay in Biologics is the same as described in the Sample Manager documentation here:
- Creating specialty assays follows a similar process. Specific properties of the specialty assays can be found in this section:
Related Topics
Biologics: Assay Integration
Connect Assays with Samples
To associate assay data with corresponding samples, include a field of type Sample in the run or result fields. You can name the field as you like and decide whether to allow it to be mapped only to a specific sample type, or to "All Samples".Learn more in the Sample Manager documentation.Connect Assays with Other Entities
To add Entity fields to an assay design, add a field of type Lookup that points to the desired Entity/DataClass. For example, the screenshot below shows how to link to Molecules.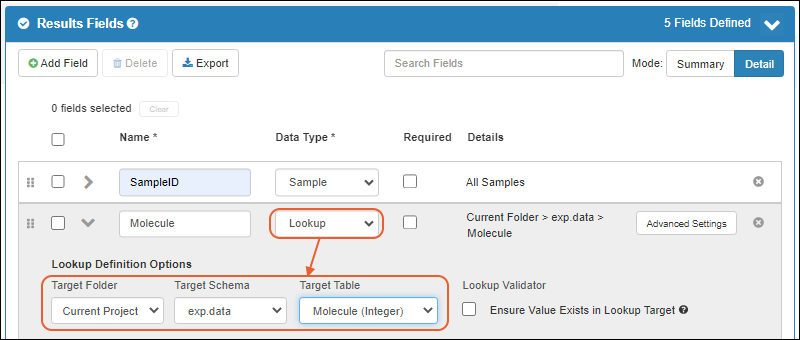
Create Integrated Grid Views
Once the assay design includes a lookup to a Sample or another Entity type (such as a Molecule), you can add other related fields to the assay design using Views > Customize Grid View.Learn about customizing and using grid views in the Sample Manager documentation.Related Topics
Biologics: Upload Assay Data
- Import Assay Data
- Import Assay Data Associated with Selected Samples
- Re-import a Run
- Delete Assay Runs
Import Assay Data
From the main menu, under Assays, click the name of the assay design you wish to import into.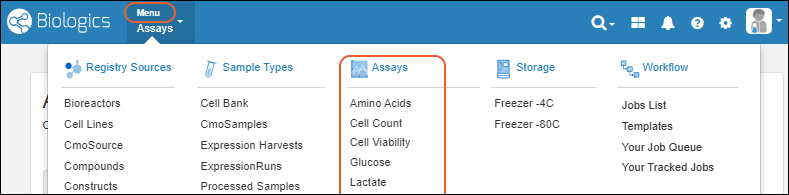 Follow the same general process as for importing assay data into Sample Manager described here. Some additional notes apply:
Follow the same general process as for importing assay data into Sample Manager described here. Some additional notes apply:
- Batch Details: If the assay design includes batch fields, you will be prompted to enter values for them.
- When importing Results, if your assay design includes File fields, you can simultaneously upload the referenced files in a second panel of the file upload tab.
Import Assay Data Associated with a Sample
An alternative way to import assay data lets you directly associate assay data with some sample(s) in the registry.From the Samples grid, select one or more samples, and then select Assay (on narrower browsers this will be a section of the More > menu). Expand the Import Assay Data section, then click the [Assay Design Name]. Scroll to select, or type into the filter box to narrow the list.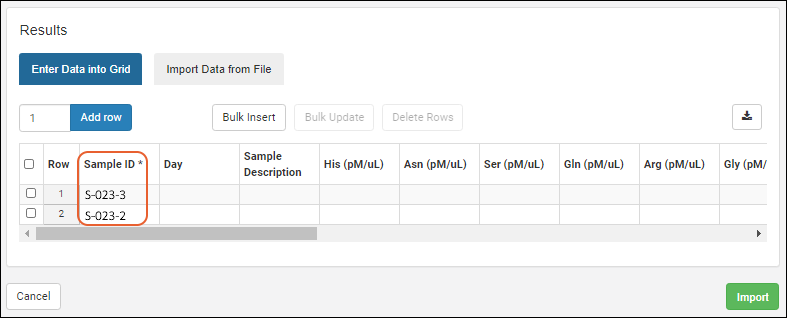 Enter the necessary values and click Import.
Enter the necessary values and click Import.
Re-Import Run
In cases where you need to revise assay data after uploading to the server, you can replace the data by "re-importing" a run.To re-import a run of assay data:- Go the details page for the target run. (For example, go to the runs table for an assay, and click one of the runs.)
- Click the Manage menu and select Re-import Run.
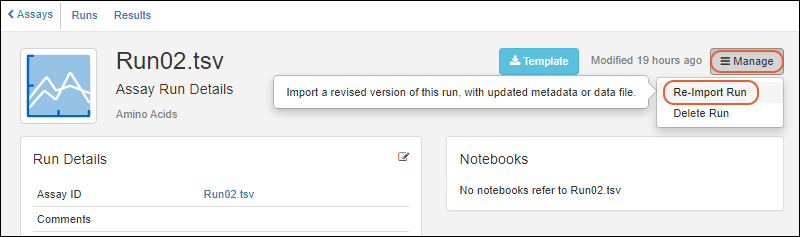
- For instrument-specific assay designs (such as NAb and Luminex), you will be directed to the specific assay’s import page in LabKey Server. Follow the procedure for re-import as dictated by the assay type: Reimport NAb Assay Run or Reimport Luminex Run.
- For general purpose assay designs, you will stay within the Biologics application:
- Revise batch and run details as needed.
- Enter new Results data. The first few lines of the existing results are shown in a preview. Choose Import Data from File or Enter Data Into Grid as when creating a new run.
- Click Re-Import.
- The data replacement event is recorded by the "Replaced By" field.
Delete Assay Runs
To delete one or more assay runs, you can:- Start from the run details and select Manage > Delete Run.
- From the grid of Runs for the assay, select the run(s) to delete, then choose Edit > Delete.
Related Topics
Biologics: Assay Batches and QC
Assay Batch Fields
Click the name of an Assay Design to see it's summary page, with several tabs, showing various scopes of the data. In addition to Runs and Results, assays in Biologics LIMS may also have fields (and a tab) for Batches of runs.Assay Quality Control
Visibility of assay data in Biologics can be controlled by setting quality control states. While an admin can edit assay designs to use QC states within the Biologics application, the actual states themselves cannot be configured in the application.Set Up QC States (Administrator)
To configure states, an admin must:- Open the Assay Design where you want to use QC. Select Manage > Edit Design.
- Confirm that the QC States checkbox is checked. If not, check it and click Finish Updating [Assay Design Name].
- Use > LabKey Server > [current Biologics folder name] to switch interfaces.
- Configure the available QC states in the system, using this topic: https://www.labkey.org/Documentation/wiki-page.view?name=assayQCconfig.
- Select > Manage Assays.
- Click the name of your assay design.
- Select QC State > Manage states.
- Define the states you want to use.
- Once complete, use > LabKey Biologics > [current Biologics folder name] to return to Biologics.
Use QC States (Users)
Once QC states have been configured in the system, users (with the appropriate roles) can assign those states to assay run data. Users can update QC states in the following cases:- When a user has both Reader and QC Analyst roles
- When a user has either Folder-, Project-, or Site Administrator roles
- Navigate to the assay run view page. (Cell Viability runs are shown below.)
- Select one or more runs.
- Select Edit > Update QC States.
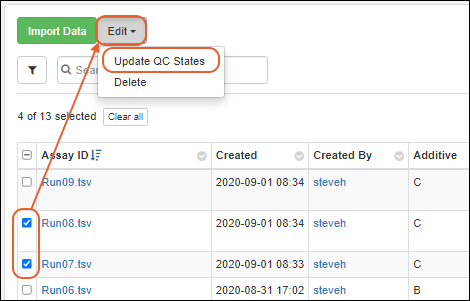
- In the popup dialog, use the dropdown menu to select the QC state. This state will be applied to all of the runs selected.
- Optionally add a comment.
- Click Save changes.
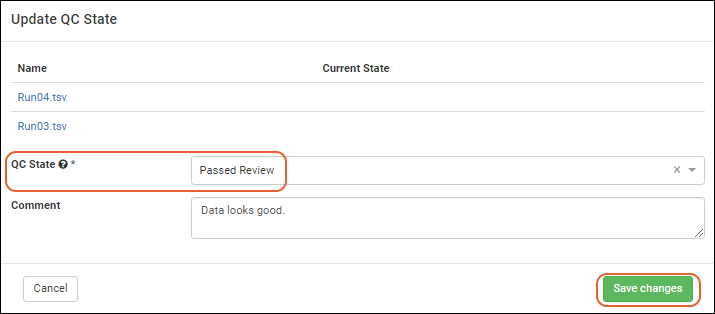
- The QC states will be reflected in the runs data grid. Admins and QC Analysts will see all of the runs, as shown below.
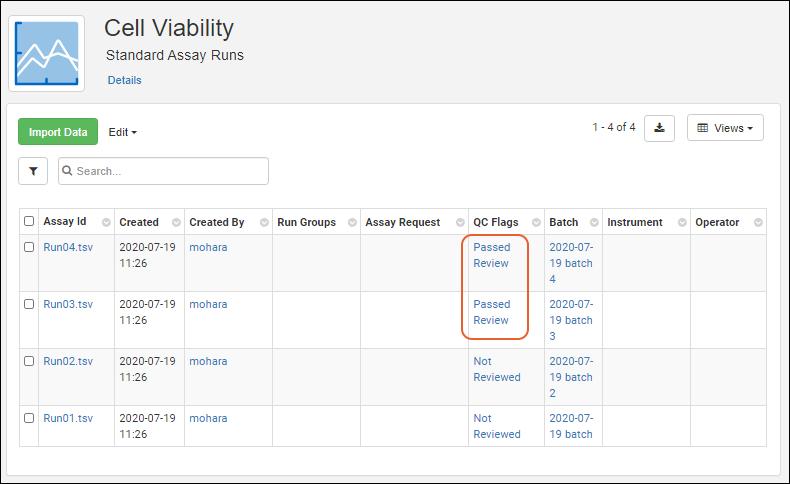
- It should be noted that if some of the QC states are not marked as "Public Data", they will not be included in the grid when viewed by non-admin users. For example, if "Not Reviewed" were marked as not public, the reader would see only the runs in public states:
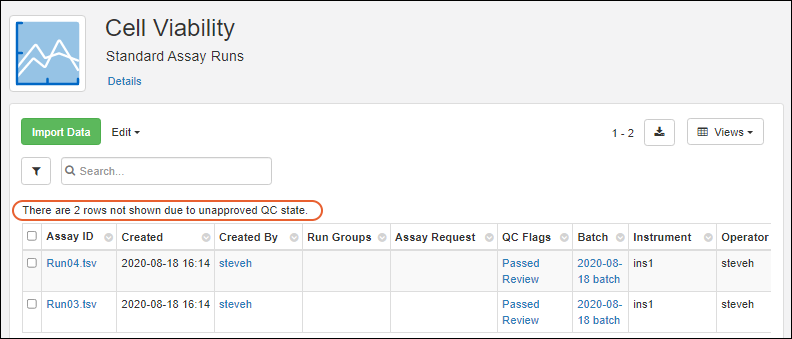
Related Topics
Biologics: Media Registration
Definitions
- Ingredients are virtual entities in LabKey Biologics, and capture the fixed natural properties of a substance. For example, the Ingredient "Sodium Chloride" includes its molecular weight, melting and boiling points, general description, etc. You register an Ingredient like this only once. Ingredients are managed with an interface similar to Registry Sources.
- Raw Materials are the particular physical instantiations of an Ingredient as real "samples" or "bottles". You register multiple bottles of the Raw Material Sodium Chloride, each with different amounts, sources, lot numbers, locations, vessels, etc. Raw Materials are managed using the Sample Type interface.
- Mixtures are recipes that combine Ingredients using specific preparation steps. Mixtures are virtual entities in LabKey Biologics. Each Mixture is registered only once, but are realized/instantiated multiple times by Batches. Mixtures are managed with an interface similar to Registry Sources.
- Batches are realizations of a Mixture recipe. They are physically real formulations produced by following the recipe encoded by some Mixture. Multiple Batches of the same Mixture can be added to the registry, each with its own volume, weight, vessel, location, etc. Batches are managed using the Sample Type interface.
Deletion Prevention
Media entities of any type cannot be deleted if they are referenced in an Electronic Lab Notebook. On the details page for the entity, you will see a list of notebooks referencing it.Topics
Managing Ingredients and Raw Materials
Definitions
In the LabKey Biologics data model, "Ingredients" are definitions, "Raw Materials" are physical things.- Ingredients are virtual entities that describe the properties of a substance. For example, the Ingredient "Sodium Chloride" includes its molecular weight, melting and boiling points, general description, etc. You register an Ingredient like this only once. Defining and creating ingredients uses the Sources UI.
- Raw Materials are physical instances of an Ingredient, tangible things that have a location in storage, with specified amounts, sources, lot numbers, locations, vessels, etc. Defining and creating raw materials uses the Samples UI.
- Mixtures are definitions. Recipe definitions, instructions for combining Ingredients. Mixtures are defined using a wizard but otherwise similar in structure and menus to Sources.
- Batches are physical things, what you get when you combine Raw Materials according to some Mixture recipe. Batches are defined using a wizard but otherwise similar in structure and menus to Samples.
Ingredients
Clicking Ingredients brings you to a grid of available (previously created) ingredients. Click Template to obtain an Excel template with the expected columns. Use the Add > menu to create new ingredients, with options similar to those for Sources.Work with Ingredients
Other actions from the grid of all ingredients are:- Edit: Select one or more ingredient rows, then choose whether to:
- Edit in Grid
- Edit in Bulk
- Delete: Note that ingredients cannot be deleted if they have derived sample or batch dependencies, or are referenced in notebooks.
- Derive Samples:
- Select one or more parent ingredients, click Derive Samples, then choose which type of sample to create.
- Reports:
- Find Derivatives in Sample Finder: Select one or more ingredients and choose this report option to open the Sample Finder filtered to show samples with the selected ingredient components. From there you can further refine the set of samples.
Ingredient Details
Click the name of an ingredient to see the details page. On the overview you can see values set for properties of the ingredient, and if any ELNs reference this ingredient, you will see links to them under Notebooks.Using the Create menu, you can add new Mixtures or Raw Materials with this Ingredient as a parent.Raw Materials
The raw materials used in mixtures are listed on the Raw Materials tab.Aliquot Raw Materials
As for samples, you can create aliquots of Raw Materials, or use certain Raw Materials to create derived samples or pooled outputs. Select the desired parent material(s) and choose Derive > Aliquot Selected or Derive or Pool as desired.Learn more in the Sample Manager documentation here: When users are selecting raw materials from a dropdown, you can use identifying fields to customize what information is shown.Deletion Prevention
Media entities of any type cannot be deleted if they are referenced in an Electronic Lab Notebook. On the details page for the entity, you will see a list of notebooks referencing it.Related Topics
Registering Mixtures (Recipes)
- Registering Mixtures
- Register a Mixture: Wizard Steps
- Register a Mixture using 'Bulk' Ingredients Table
- Registering Mixtures with Unknown Ingredients, Amounts, or Concentrations
- Delete a Mixture
- Use the Recipe API to Update Mixtures
Definitions
- Mixtures are recipes that combine Ingredients using specific preparation steps. Mixtures are virtual entities in LabKey Biologics. Each Mixture is registered only once, but are realized/instantiated multiple times by Batches.
- Batches are realizations of a Mixture recipe. They are physically real formulations produced by following the recipe encoded by some Mixture. Multiple Batches of the same Mixture can be added to the registry, each with its own volume, weight, vessel, location, etc.
Registering Mixtures
The virtual recipes are listed in the Media > Mixtures grid. Find a given Mixture by filtering, sorting, or searching the grid.- Create a new mixture "from scratch"
- Create a new mixture starting from an existing mixture or ingredient
- Registering Mixtures with Unknown Ingredients, Amounts, or Concentrations.
- Registering Batches with Unknowns.
Start a New Mixture "From Scratch"
From the main menu, select Media, then Mixtures and then click Add above the grid of available mixtures.
Start from an Existing Ingredient or Mixture
Any of the registered ingredients or mixtures can be clicked for a detailed view. The details view includes general information. Mixture detail pages show the mixture’s included ingredients and preparation steps.Select Manage > Create Mixture. The Ingredients tab will be prepopulated.
Register a Mixture: Wizard Steps
The new mixture wizard adds a new mixture to the registry, and performs a check to ensure that the mixture name is unique in the registry. Duplicate mixture names are not allowed in the registry.Wizard Step 1: Details
The Details step asks for basic information about the mixture, including a type such as powder or solution. Once all required fields have been filled in, the Next button will become clickable.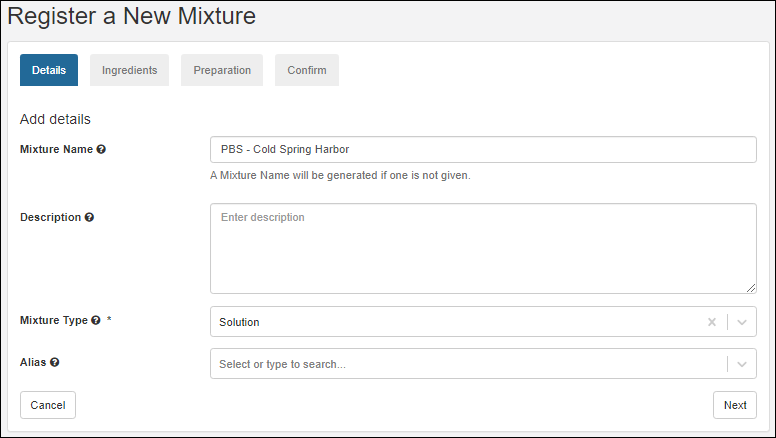 Upon clicking Next, the mixture name is checked against the registry to see if it is already in use. A warning displayed if a duplicate name is found.
Upon clicking Next, the mixture name is checked against the registry to see if it is already in use. A warning displayed if a duplicate name is found.
Wizard Step 2: Ingredients
The Ingredients step allows you to add single ingredients or existing mixtures to a new mixture recipe, as well as the required amounts and the amount unit. If you started the wizard from an ingredient or mixture it will be prepopulated here.Specify what Recipe Measure Type the mixture is using: Mass or Volume. This selection dictates what unit types are available for the ingredients below.- If Mass is selected, unit types will be: g/kg, g/g, mL/kg, mol/kg, mol/g, S/S
- If Volume is selected, unit types will be: g/kL, g/L, L/L, mL/L, mol/kL, mol/L, μL/L, μL/mL
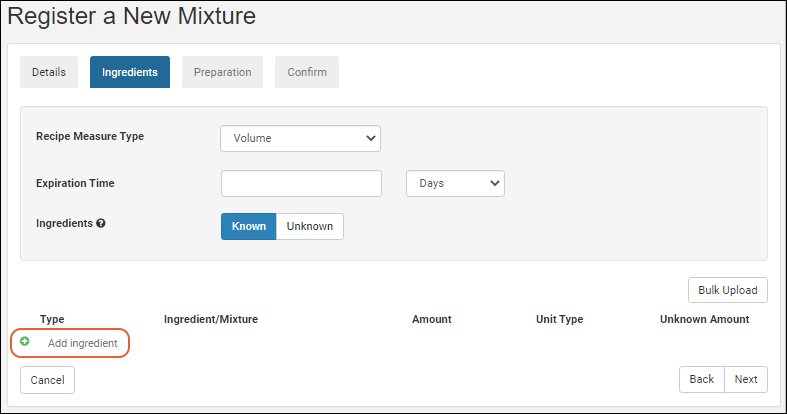 Under Type select either Ingredient or Mixture. The Ingredient/Mixture text boxes are 'type ahead' searches, that is, as you type, the registry will offer a filtered dropdown of options that contain your typed string. To control how many fields are shown with each option, you can provide a lookup view.
Under Type select either Ingredient or Mixture. The Ingredient/Mixture text boxes are 'type ahead' searches, that is, as you type, the registry will offer a filtered dropdown of options that contain your typed string. To control how many fields are shown with each option, you can provide a lookup view.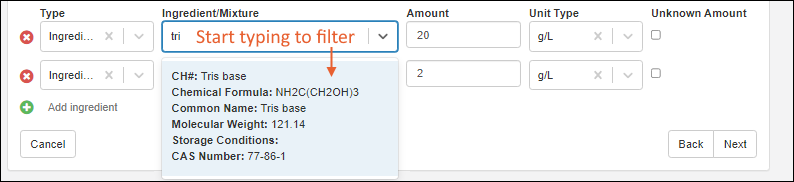 When at least one ingredient/mixture and the associated amount/amountUnit fields have been filled in, the Next button becomes enabled. If a selected Ingredient or Mixture is no longer desired, the red X button on the left can be clicked to remove it.
When at least one ingredient/mixture and the associated amount/amountUnit fields have been filled in, the Next button becomes enabled. If a selected Ingredient or Mixture is no longer desired, the red X button on the left can be clicked to remove it.
Wizard Step 3: Preparation
The preparation step lets you add one or more preparation instructions. Click Add Step to add one or more text boxes. When all instructions have been entered, click the Next button.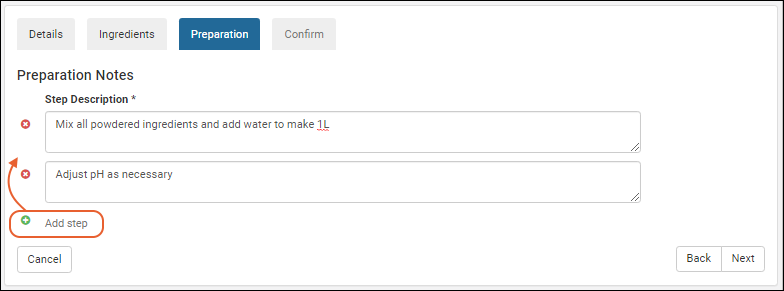
Wizard Step 4: Confirmation
The confirmation step summarizes all of the information entered so far.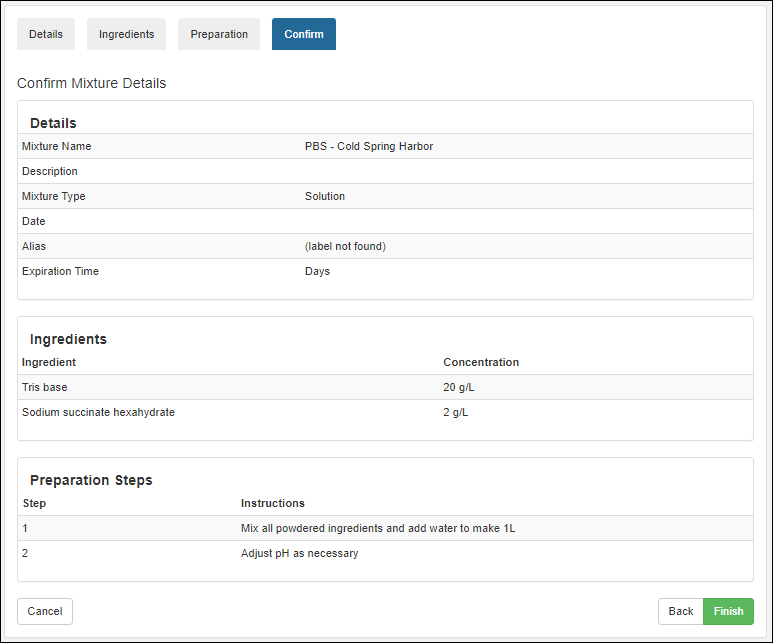 Click Finish to submit; the mixtures grid is shown with the new addition.
Click Finish to submit; the mixtures grid is shown with the new addition.
Register a Mixture using 'Bulk' Ingredients Table
This method of registering a mixture lets you enter the ingredients in a tabular format by copying-and-pasting from an Excel or TSV file.- Go to the Mixtures grid and click Add.
- On the Details tab, enter the Mixture Name and Mixture Type (and description and aliases if needed) then click Next.
- On the Ingredients tab, click Bulk Upload.
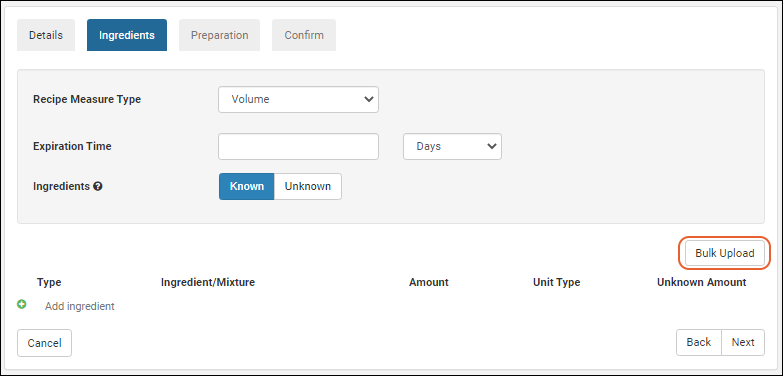
- A popup window appears showing the table headers which you can copy-and-paste into an empty Excel file.
- Fill out the table, adding separate lines for each ingredient in the mixture. Only the "Ingredient/Mixture" column is required. If a cell is left blank, you can complete the details of the mixture using the user interface.
- Type: (Optional) Specify either "Ingredient" or "Mixture".
- Ingredient/Mixture: (Required) The name of the ingredient or mixture which must be a pre-existing item already in the registry. Values from the Name or Scientific Name columns are supported.
- Amount: (Optional) A number that indicates the weight or volume of the ingredient.
- Unit Type: (Optional) Possible values are the same as when entering ingredients in the UI.
- Select whether to:
- Append new ingredients or
- Replace ingredients already listed.
- Copy-and-paste the table back into the popup window and click Add Ingredients.
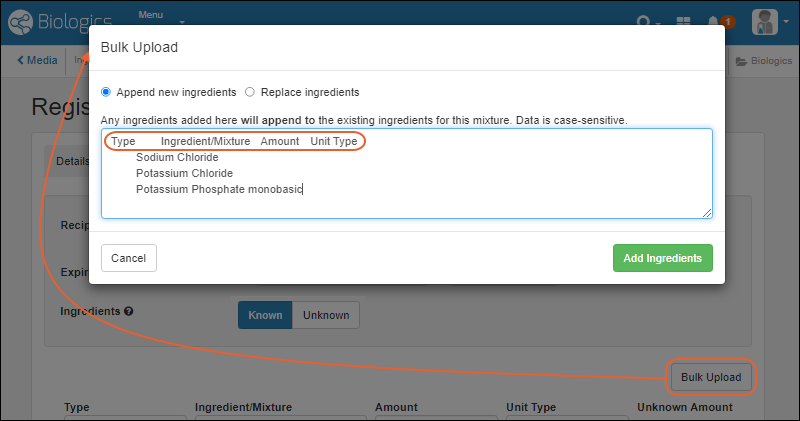
- Fill in any remaining fields, if necessary, and click Next.
- Complete the rest of the mixture wizard.
Registering Mixtures with Unknown Ingredients, Amounts, or Concentrations
In cases where you do not know the exact ingredients, amounts, or concentrations of a material, you can still add it to the registry. These scenarios are common when receiving a material from an outside vendor who does not disclose the exact formulation of the product.In such cases, the mixture registration process is largely the same, except for the Ingredients step, where you can toggle all ingredients and amounts as unknown, or toggle individual amounts as unknown.Clicking Unknown disables entry of any further details about the recipe: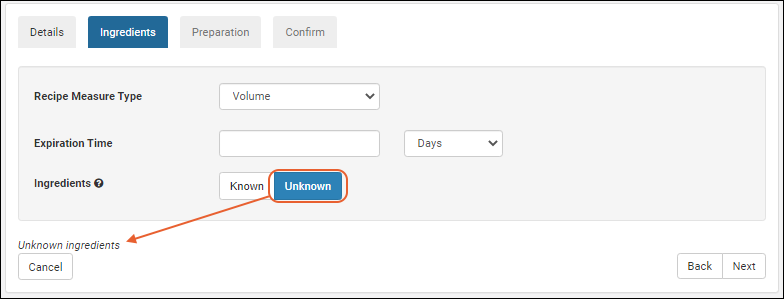 Selecting Known but checking an Unknown Amount box disables that specific ingredient's amount and unit type inputs (other ingredients may still have known amounts).
Selecting Known but checking an Unknown Amount box disables that specific ingredient's amount and unit type inputs (other ingredients may still have known amounts).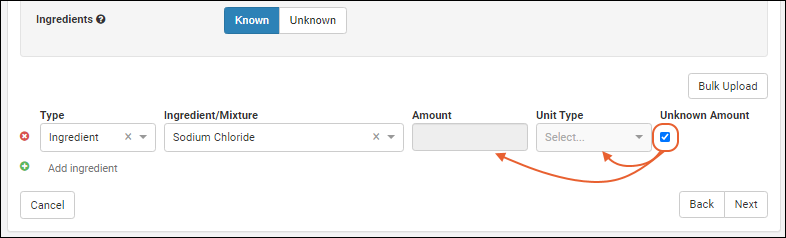 Note that you can register materials and mixtures without specifying ingredients or amounts, even when you do not explicitly check one of the 'unknown' boxes. Warning messages are provided to confirm that your registration is intentional.
Note that you can register materials and mixtures without specifying ingredients or amounts, even when you do not explicitly check one of the 'unknown' boxes. Warning messages are provided to confirm that your registration is intentional.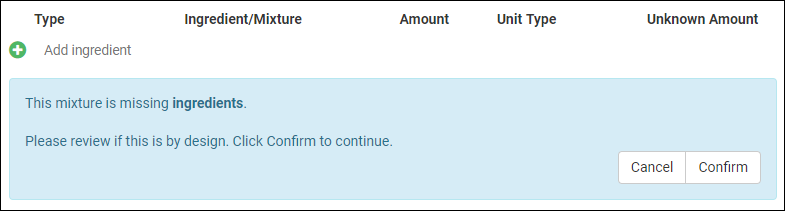 Once registered, mixtures with unknown ingredients and amounts behave just as any other mixture, with a slight change to their detail view to illustrate unknown amounts or ingredients.
Once registered, mixtures with unknown ingredients and amounts behave just as any other mixture, with a slight change to their detail view to illustrate unknown amounts or ingredients.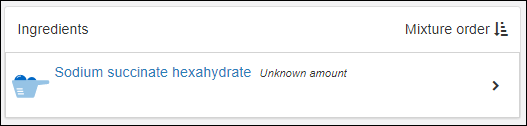
Bulk Registration of Mixtures That Include Unknowns
In bulk registration of Mixtures, some fields support the text "unknown" on import. For details, see Bulk Registration of Entities.Delete a Mixture
Each Mixture is composed of an entry in the Mixture DataClass table and a Protocol that defines the ingredients used and the steps. To delete the mixture completely, delete the Protocol and then you can delete the Mixture DataClass.- Switch to the LabKey Server interface via > LabKey Server > [Folder Name]
- Select > Go to Module > Experiment.
- Scroll down to the list of Protocols.
- Select the Protocol(s) to delete, and click to delete.
- You should now be able to delete the Mixture DataClass row.
Use the Recipe API to Update Mixtures
Mixtures can be updated after they are created by using the Recipe API. You can make these updates using PUT requests to recipe-recipe.api:- Mutation of ingredients and their associated metadata.
- Description can be changed on the underlying protocol.
- Recipe produces metadata.
- Recipe name
- Changing the recipe steps. Use the pre-existing recipe-steps.api endpoint if you're looking to edit these.
- Call recipe-read.api and receive the full recipe/mixture definition.
- Mutate the received object in the way you'd like (change ingredients, amounts, etc).
- Remove things you do not want to mutate (e.g. "produces", etc) or similarly copy to a new object only the properties you want to mutate.
- Call recipe-recipe.api and PUT the mutated object on the payload as the recipe.
Related Topics
- Managing Ingredients and Raw Materials: Register new ingredients and raw materials
- Registering Batches
Registering Batches
- Batch Design
- Batches
- Registering Batches with Unknowns
- Bulk Registration of Batches That Include Unknowns
- Aliquot Batches
- Add Detail to "Raw Materials Used" Dropdown
- Use the Recipe API to Update Batches
Batch Design
Batches represent a specific quantity of a given mixture. The mixture is a virtual entity, i.e. a recipe, used to create a real, physical batch of material. The design of the Batch, meaning the fields, properties, and naming pattern, is set by default to provide commonly used fields and defaults, but can be edited using Manage > Edit Batch Design.Batches
To create a new batch, click Add above the batches grid, or select Manage > Create Mixture Batch from any mixture detail page to include that mixture in your batch.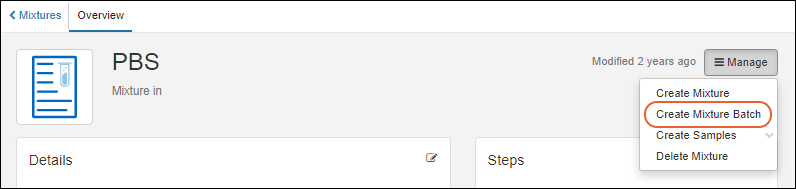 The steps in the batch creation wizard are similar to those in the mixture wizard.
The steps in the batch creation wizard are similar to those in the mixture wizard.
Details
On the Details tab, complete all required fields and any option fields desired. If the Batch Name is not provided, it will be automatically assigned a unique name using the naming pattern for Batches. Hover over the for any field to see more information.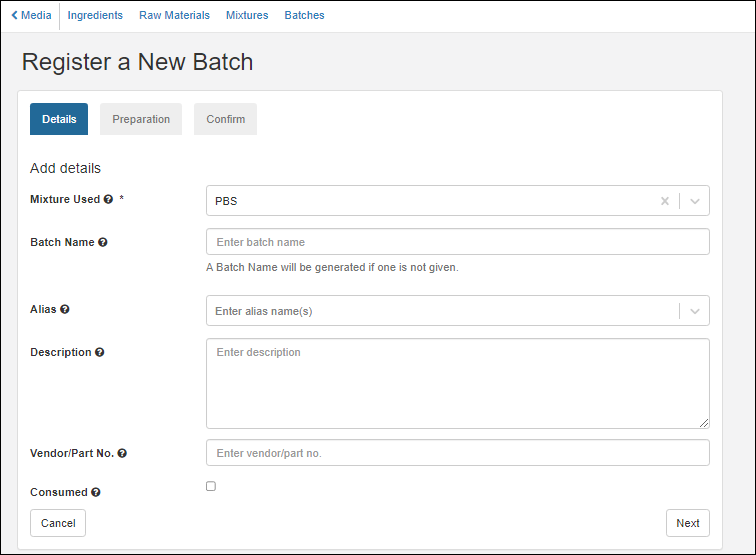 Click Next.
Click Next.
Preparation
On the Ingredients tab, each ingredient row is disabled until you add a Desired Batch Yield and Unit. Once selected, the rest of the page will become enabled and populate information.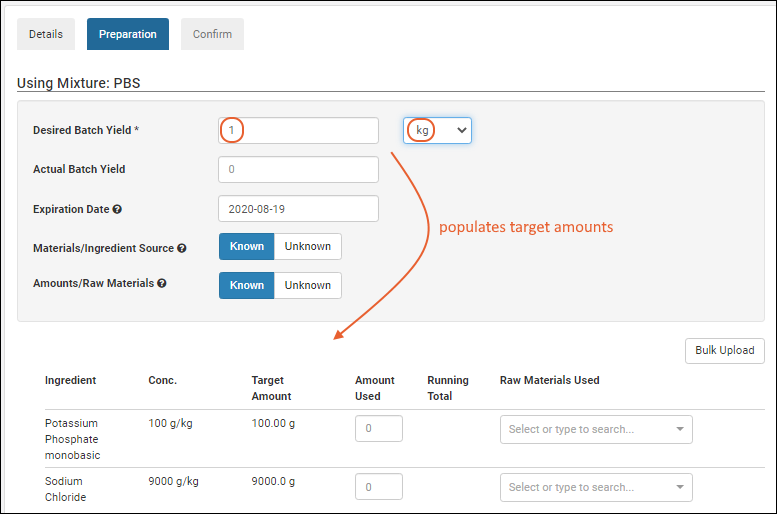 Enter the exact amounts of which raw materials were used to create the mixture recipe. (If ingredients and/or amounts are unknown, see options below.) For each raw material, specify a source lot by its id number. An administrator can enhance the details show to users in this dropdown or other dropdowns by using Identifying Fields.Each ingredient in the recipe may have one or more source lots of raw material This is useful when you exhaust one lot and need to use a second lot to complete the batch. For example, using 40g of Potassium Phosphate empties a lot, if the next lot contains 45, you would need an additional 15g from another lot to reach the target amount of 100g shown in the following. Add additional lots by clicking Add raw material for the specific ingredient.
Enter the exact amounts of which raw materials were used to create the mixture recipe. (If ingredients and/or amounts are unknown, see options below.) For each raw material, specify a source lot by its id number. An administrator can enhance the details show to users in this dropdown or other dropdowns by using Identifying Fields.Each ingredient in the recipe may have one or more source lots of raw material This is useful when you exhaust one lot and need to use a second lot to complete the batch. For example, using 40g of Potassium Phosphate empties a lot, if the next lot contains 45, you would need an additional 15g from another lot to reach the target amount of 100g shown in the following. Add additional lots by clicking Add raw material for the specific ingredient. Note that if one or more raw materials are indicated, the option to set an amount or raw material as unknown is no longer available.
You can also add ingredients (or other mixtures) as you register a batch. Click Add ingredient at the end of the ingredient list.
Note that if one or more raw materials are indicated, the option to set an amount or raw material as unknown is no longer available.
You can also add ingredients (or other mixtures) as you register a batch. Click Add ingredient at the end of the ingredient list.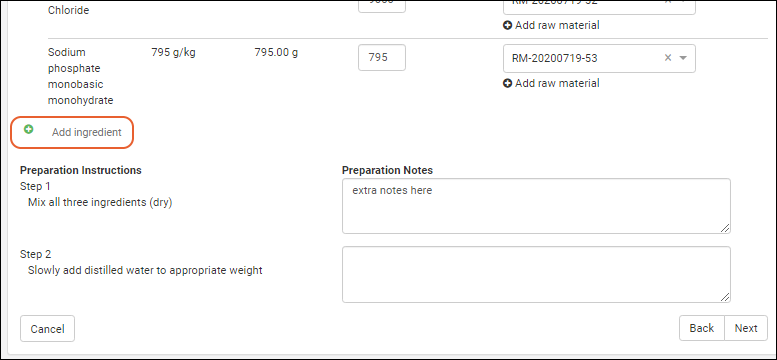 For each preparation step in the mixture recipe, the user can enter any preparation notes necessary in creating this particular batch. Notes are optional, but can provide helpful guidance if something unusual occurred with the batch.Once all required fields are filled, the Next button will become clickable.
For each preparation step in the mixture recipe, the user can enter any preparation notes necessary in creating this particular batch. Notes are optional, but can provide helpful guidance if something unusual occurred with the batch.Once all required fields are filled, the Next button will become clickable.
Confirmation
The confirmation step allows the user to view the information they have entered. If necessary, they can click back to return to previous tabs to make any updates. Preparation notes are collapsed, but can be expanded by clicking the right-arrow icon.Click Finish to register this batch and see the row as entered in the grid.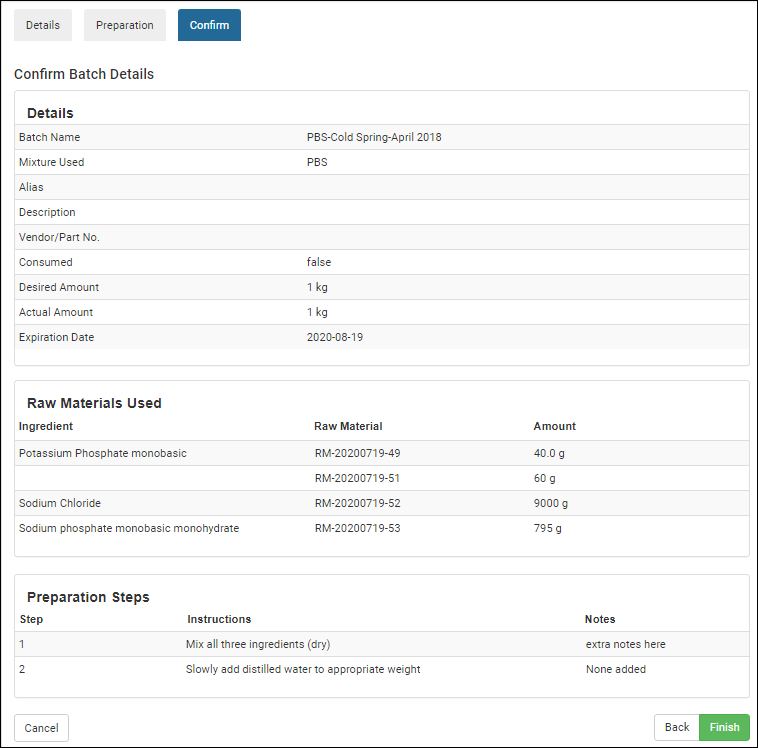 If multiple raw materials were used, the batch details panel will display material lot numbers and the amounts used.
If multiple raw materials were used, the batch details panel will display material lot numbers and the amounts used.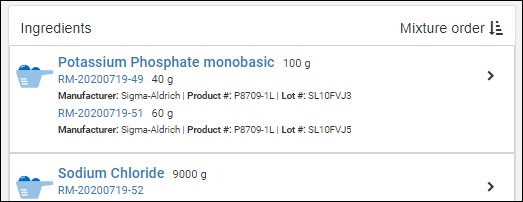 If additional ingredients were added to the batch, these modifications from the mixture recipe will be noted in a panel:
If additional ingredients were added to the batch, these modifications from the mixture recipe will be noted in a panel: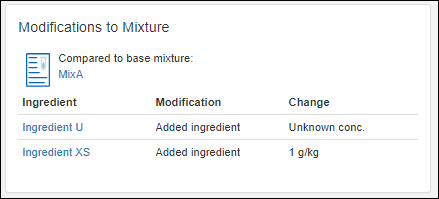
Amounts and Units
The amount, and units for that amount, are determined from the Batch Yield details provided on the Preparation tab. Three fields are stored and visible:- Recipe Amount (RecipeAmount) : The amount, i.e. "Desired Batch Yield". This value is stored as a property on the run that created the batch. Note that this field is not editable as it is not a property of the batch.
- Recipe Amount Units (Units and also RawUnits): The units value for the Recipe Amount field.
- Recipe Actual Amount (StoredAmount and also RawAmount): The "Actual Batch Yield".
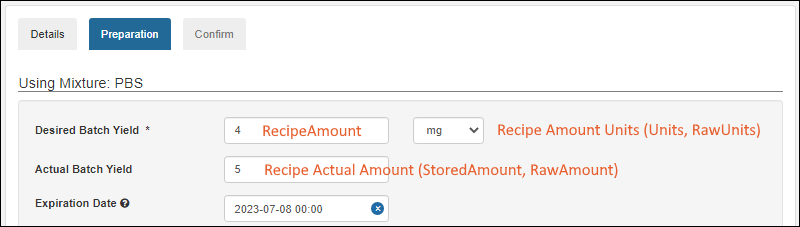 Note that if you have existing data prior to the addition of the Amount and Units default fields in version 23.4, the combined amount-with-units field will be parsed into the two fields. If we cannot parse that text field (such as in a case where the units value is not supported) the two fields will be left blank and need to be manually populated.
Note that if you have existing data prior to the addition of the Amount and Units default fields in version 23.4, the combined amount-with-units field will be parsed into the two fields. If we cannot parse that text field (such as in a case where the units value is not supported) the two fields will be left blank and need to be manually populated.
Registering Batches with Unknowns
When registering batches, the Ingredients step includes the option to mark:- all materials as unknown
- individual raw materials as unknown
- individual amounts as unknown
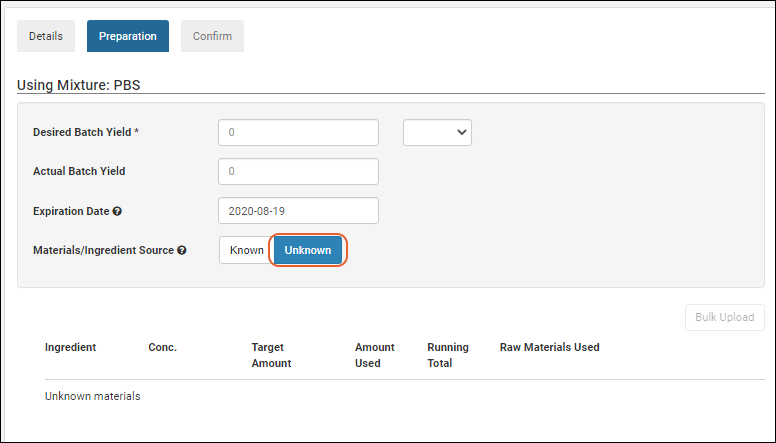 Confirmation warnings are provided if the user provides incomplete values and an 'unknown' box is not ticked.
Confirmation warnings are provided if the user provides incomplete values and an 'unknown' box is not ticked. Once entered into the registry, the unknown factors are reflected in the user interface.
Once entered into the registry, the unknown factors are reflected in the user interface.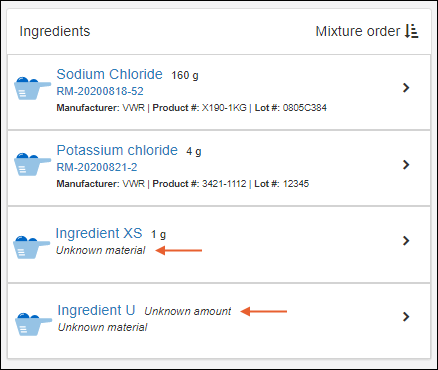
Bulk Registration of Batches That Include Unknowns
In bulk registration of Batches, some fields support the text "unknown" on import. For details, see Bulk Registration of Entities.Aliquot Batches
Instead of having to register sub-portions of mixture batches in advance, you can create large batches, then later create aliquots from them. You can specify the Aliquot Naming Pattern by editing the Mixture Batch Design. The default name of an aliquot is the name of the parent batch, followed by a dash and a counter for that parent. Learn more here: Select one or more Batches from the grid and choose Derive > Aliquot Selected.Add Detail to "Raw Materials Used" Dropdown
The "Name" of a Raw Material is it's generated ID; these are not particularly helpful to users selecting the raw materials for a batch. By customizing raw materials with Identifying Fields, administrators can provide their users with the necessary details when they select from the dropdown.For example, if you define "Product Number" and "Lot Number" as identifying fields, they will appear for users with the "Name" of the raw material.Use the Recipe API to Update Batches
Batches can be updated after they are created by using the Recipe API. You can make these updates using PUT requests to recipe-batch.api:- Mutation of materials and their associated metadata.
- Mutation of amount, amountUnits, and actualAmount of produces.
- Comments
- Changing which sample is produced.
- Changing the source recipe (mixture).
- Changing the batch steps. Use the pre-existing recipe-notes.api endpoint if you're looking to edit these.
- Call recipe-getBatch.api and receive the full mixture batch definition.
- Mutate the received object in the way you'd like (change materials, amounts, etc.).
- Remove things you do not want to mutate (e.g. "produces", etc.) or similarly copy to a new object only the properties you want to mutate.
- Call recipe-batch.api and PUT the mutated object on the payload as the batch.
Related Topics
Biologics Administration
Biologics Admin Topics
- Biologics: Detail Pages and Entry Forms
- Biologics: Protect Sequence Fields
- Manage Notebook Tags
- Biologics Admin: URL Properties
Related Topics
Biologics: Detail Pages and Entry Forms
Custom Details Pages
An administrator can create a special custom grid view to control the fields shown on an entity details page. If you create this special view using the Biologics interface, it will only show for your own user account. To make this change to the details view for all users, select > LabKey Server > [Your Biologics Project Name] to switch to the LabKey Server UI. Here you can save the "BIOLOGICSDETAILS" view and share it with all users.Customize the view for the entity type, then save it:- Uncheck "Make default view for all users"/"Default grid view for this page".
- Use the name "BIOLOGICSDETAILS".
- Check the box to "Make this grid view available to all users" (available only in the LabKey Server interface).
- Optionally check the box to "Make this grid view available in child folders."
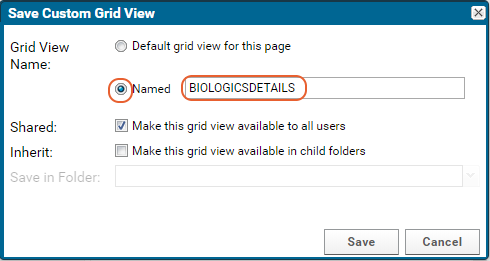
Entry Forms
To modify entry and update forms, modify the fields in the field designer. For example, to add a field "NIH Registry Number" to the Cell Line entity:- From the main menu, click Cell Lines.
- Select Manage > Edit Cell Lines Design.
- Click the Fields section to open it.
- Expand the desired field details and enter a Label if you want to show something other than the field name to users.
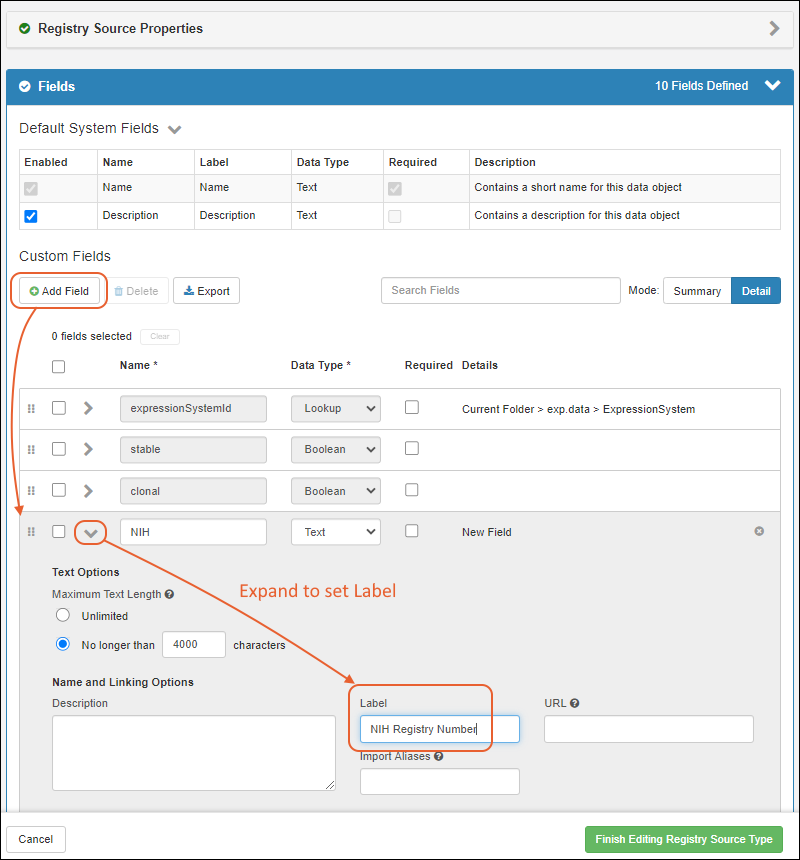
- Click Finish Editing Source Type.
- The field will appear in the entry form for Cell Lines when you Create a new one.
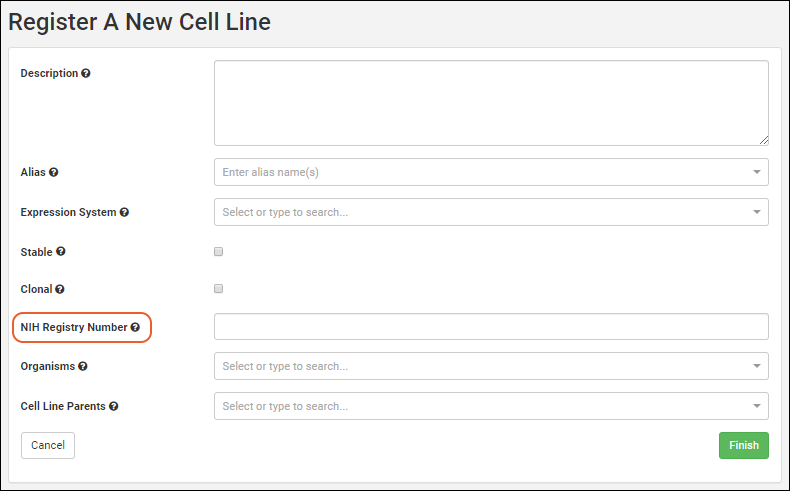 Follow similar procedures to delete or modify entry form fields for other entities.
Follow similar procedures to delete or modify entry form fields for other entities.
Customize Details Shown for Lookups
Lookup fields connect your data by letting a user select from a dropdown which joins in data from another table. By default, a lookup column will only show a single display field from the table, generally the first text field in the target table.Details for Registry Sources and Sample Types
By defining Identifying Fields for Registry Sources and Sample Types, including Media types, you can expose additional detail. Learn more here: As an example of how this can be used to make it easier for users to select the right raw materials for a batch, review this topic: Add Detail to "Raw Materials Used" Dropdown for Registering BatchesDetails for Lists
You can also customize the lookup view in the Sample Manager, LabKey LIMS, and Biologics LIMS for a list or other table by editing the metadata directly. For example, for dropdown targeting a list of Supplier names, it might also be helpful to users to show the state where that supplier is located. In this example, the "Suppliers" list would include the following XML metadata to set shownInLookupView to true for the desired field(s):<tables xmlns="http://labkey.org/data/xml">
<table tableName="Suppliers" tableDbType="NOT_IN_DB">
<columns>
<column columnName="State">
<shownInLookupView>true</shownInLookupView>
</column>
</columns>
</table>
</tables>
Related Topics
Biologics: Protect Sequence Fields
- Restrict Visibility of Nucleotide and Protein Sequences
- PHI/IP Protection Levels and Permissions
- Omit or Blank Protected Columns
- Conditional Viewing of Sequences
- Mark Other Fields as Protected
Restrict Visibility of Nucleotide and Protein Sequences
An administrator can enable Protected Data Settings within the Biologics application. Once enabled, a user (even an administrator) must have the "Restricted PHI Reader" role in the folder to view the protein sequences and nucleotide sequences themselves.- In the Biologics application, select > Application Settings.
- Scroll down to Protected Data Settings, then check the box to Require 'Restricted PHI' permission to view nucleotide and protein sequences.
- If you don't see this box, you do not have sufficient permissions to make this change.
PHI/IP Protection Levels and Permissions
There are four available 'levels' of protection, detailed in this topic: https://www.labkey.org/Documentation/wiki-page.view?name=phiLevels- Not PHI
- Limited PHI
- Full PHI
- Restricted PHI: the highest level of information to be protected. Sequences are protected at this level.
- Use the menu to select LabKey Server and your Biologics folder.
- Select > Folder > Permissions.
- Assign the Restricted PHI Reader role to any user who should be able to read Sequences.
- Note that this role is required to see Sequences, even for users with admin access to the Biologics application.
- Learn more about setting permissions here: https://www.labkey.org/Documentation/wiki-page.view?name=configuringPerms
- Save and Finish when permissions settings are complete.
Omit or Blank Protected Data
You can choose whether to show an empty column (the default) or omit the column entirely.- Select > LabKey Server > Your Biologics Folder.
- Select > Folder > Management.
- Click the Compliance tab.
- Confirm that Require PHI roles to access PHI columns is checked.
- Use the radio button to select either:
- Blank the PHI column. (Default) The column is still shown in grid views and is available for SQL queries, but it will be empty.
- Omit the PHI column. The column is completely unavailable to the user.
Conditional Viewing of Sequences
Once configured as described above, any user without 'Restricted PHI' permission will see that there is a Sequence column, but the heading will be shaded and no sequences shown. Hovering reveals the message "PHI protected data removed"Mark Other Fields as Protected
Other fields in Registry Source Types and Sample Types may also be protected using the PHI mechanism, when enabled. A user with lower than "Restricted PHI Reader" access will not see protected data. In the LabKey Server interface, they will see a banner to this effect.The mechanism of setting and user experience is slightly different for Registry Source Types and Samples.Protect Registry Source Type Fields
When a field in an entity Registry Source Type other than the Sequence fields is set to a higher level of PHI than the user is authorized to access, it will be hidden in the same way as the Sequence fields.To mark fields in Registry Source Type, such as registry entities including nucleotide and protein sequences, use the LabKey Server interface to edit these "Data Classes".- Use the menu to select LabKey Server and your Biologics folder.
- Click the name of the Data Class to edit, such as "NucSequence"
- Click Edit.
- In the Fields section, click to expand the desired field.
- Click Advanced Settings.
- Use the PHI Level dropdown to select the desired level.
- Click Apply.
- Click Save.
- Return to the Biologics application.
Protect Sample Fields
When a field in a Sample Type is protected at a higher level of PHI than the user can access, they will not see the empty column as they would for an entity field. To mark fields in Sample Types as PHI/IP, use the Biologics interface.- Click the name of your Sample Type on the main menu.
- Select Manage > Edit Sample Type Design.
- In the Fields section, click to expand the desired field.
- Click Advanced Settings.
- Use the PHI Level dropdown to select the desired level.
- Click Apply.
- Click Save.
Related Topics
Manage Notebook Tags
Manage Tags
From the main menu, click Notebooks, then select Manage > Tags to open the dashboard.Delete Unused Tag
To delete tags, an administrator can select one or more rows and click Delete. Tags in use for notebooks cannot be deleted.Add New Tag
To add a new tag, an administrator clicks Create Tag, provides a name and optional description, sets a tag color, then clicks Create Tag again in the popup.Require Tags
By default, new notebooks do not require a tag. An administrator can require a tag for every notebook by selecting > Application Settings and scrolling to the Notebook Settings section. Check the box to require a tag.Select Tag for Notebook
Notebook authors will see the colors and tags available when they create or edit notebooks:Related Topics
Biologics Admin: URL Properties
Last Page You Viewed
When you are in the LabKey Server interface, you can hand edit the URL to return to the last page you were viewing before switching to LabKey.Substitute the server and path for your biologics-app.view and append the lastPage property:https://example.lkpoc.labkey.com/Biologics%20Example/biologics-app.view?#/.lastPage
Reports Dashboard
To view the Reports dashboard, use "/reports":https://example.lkpoc.labkey.com/Biologics%20Example/biologics-app.view?#/reports
URL Redirects
The following redirects can be done directly to the URL for navigating a user programmatically. Note that these examples assume various rowID to assay mappings that may be different in your implementation.| URL ending like this | Can be represented by | Resolves to... |
|---|---|---|
| Any Biologics URL | #/.lastPage | The last page you were viewing |
| /assay-assayBegin.view?rowId=45 | #/assays/45 | #/assays/general/titer |
| /assay-assayResults.view?rowId=765 | #/assays/765 | #/assays/general/amino%20acids |
| /experiment-showDataClass.view?name=CellLine | #/rd/dataclass/cellline | #/registry/cellline |
| /experiment-showDataClass.view?name=Mixture | #/rd/dataclass/mixture | #/media/mixtures |
| /experiment-showData.view?rowId=3239 | #/rd/expdata/3239 | #/registry/molecularspecies/3239 |
| /experiment-showMaterials.view?rowId=3087 | #/rd/samples/3087 | #/samples/expressionsystemsamples/3087 |
| /list-grid.view?listId=18&pk=2 | #/q/lists/18/2 | #/q/lists/mixturetypes/2 |
| /reports | #/reports | Reports Dashboard |
| /assays | #/assays | Assay Dashboard |
| /query-begin.view | #/q | Schema/Query Browser |
| /list-begin.view | #/q/lists | Lists |