Panorama is a web server database application for targeted proteomics assays that integrates into a Skyline proteomics workflow. The LabKey Panorama module supports management of targeted mass spectrometry data and integration with Skyline workflows (SRM-MS, filtering or MS2 based projects).
Panorama Folder Types
To begin working with Panorama, create a new folder, choosing
Panorama as the folder type. On the
Configure Panorama Folder page, select one of the available configurations. Each type in this list links to a set of documentation.
- Experimental data: A collection of published Skyline documents for various experimental designs.
- Quality control: System suitability monitoring of reagents and instruments.
- Chromatogram library: Curated precursor and product ion expression data for use in designing and validating future experiments.
- Check Rank peptides within proteins by peak area if your data contains relative peptide expression for proteins.
- Multi-attribute method (MAM): An experimental data folder variant with additional reporting for multi-attribute method analyses.
Create New Panorama Folder
- Create a new folder. Hover over your project name in the menu bar (On PanoramaWeb, right below the Panorama icon) and click on the New Subfolder icon circled in the image below.
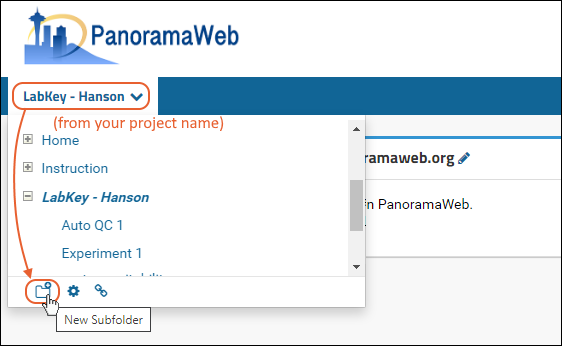
- Give the new folder a Name, then select the Panorama option under Folder Type. This is the folder type that should be selected for all workflows supported by Skyline (SRM-MS, MS1 filtering or MS2 based projects).
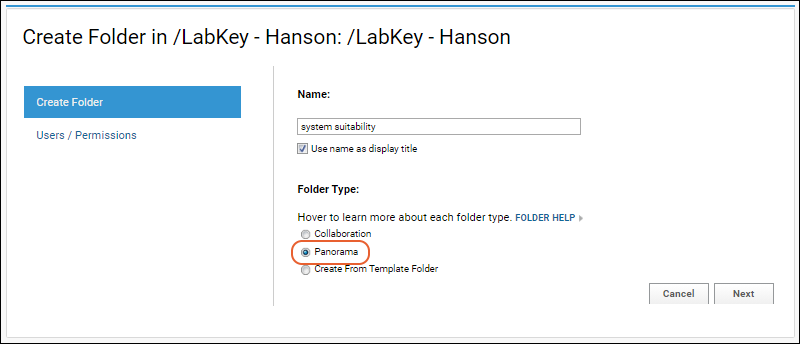
- On the Users / Permissions page, select one of the available options and click Next.
- You can also change permissions on a folder after it has been created.
- The next page, Configure Panorama Folder, asks you to choose the type of Panorama folder you would like to create, detailed above.
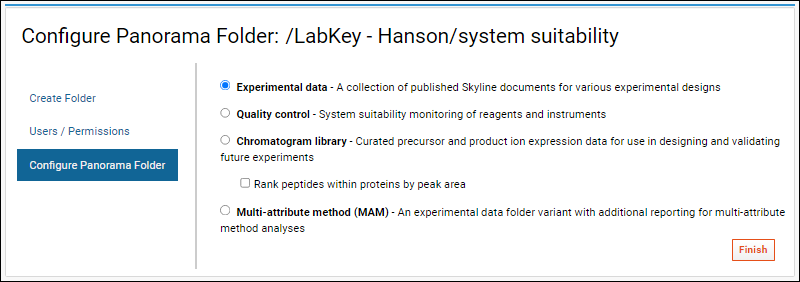

Topics for each Panorama Folder Type:


