The LabKey
data processing pipeline allows you to process and import data files with tools we supply, or with tools you build on your own. You can set a pipeline override to allow the data processing pipeline to operate on files in a preferred, pre-existing directory instead of the
file root, the directory where LabKey ordinarily stores files for a project. Note that you can still use the data processing pipeline without setting up a pipeline override if the system's default locations for file storage are sufficient for you.
A pipeline override is a directory on the file system accessible to the web server where the server can read and write files. Usually the pipeline override is a shared directory on a file server, where data files can be deposited (e.g., after MS/MS runs). You can also set the pipeline override to be a directory on your local computer.
Before you set the pipeline override, you may want to think about how your file server is organized. The pipeline override directory is essentially a
window into your file system, so you should make sure that the directories beneath the override directory will contain only files that users of your LabKey system should have permissions to see. On the LabKey side, subfolders inherit pipeline override settings, so once you set the override, LabKey can upload data files from the override directory tree into the folder and any subfolders.
Single Machine Setup
These steps will help you set up the pipeline, including an override directory, for usage on a single computer. For information on setup for a distributed environment,
see the next section.
- Select > Go to Module > Pipeline.
- Click Setup. (Note: you must be a Site Administrator to see the Setup option.)
- You will now see the "Data Processing Pipeline Setup" page.
- Select Set a pipeline override.
- Specify the Primary Directory from which your dataset files will be loaded.
- Click the Searchable box if you want the pipeline override directory included in site searches. By default, the materials in the pipeline override directory are not indexed.
- You may also choose to customize Pipeline Files Permissions using the panel to the right.
- Click Save.
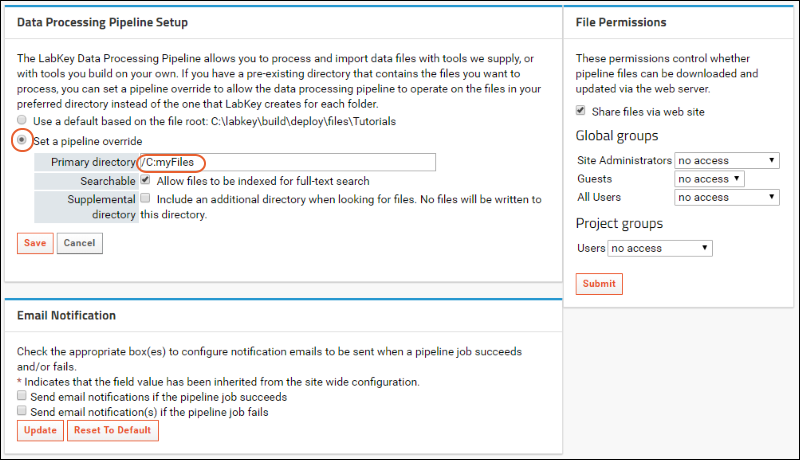
Notice that you also have the option to
override email notification settings at this level if desired .
Include Supplemental File Location (Optional)
Projects that set a pipeline override can specify a supplemental, read-only directory, which can be used as a repository for your original data files. If a supplemental directory is specified, LabKey Server will treat both directories as sources for input data to the pipeline, but it will create and change files only in the first, primary directory.
Note that UNC paths are not supported for pipeline roots here. Instead,
create a network drive mapping configuration via
> Site > Admin Console > Settings > Configuration > Files. Then specify the letter mapped drive path as the supplemental file location.
Set Pipeline Files Permissions (Optional)
By default, pipeline files are not shared. To allow pipeline files to be downloaded or updated via the web server, check the
Share files via web site checkbox. Then select appropriate levels of permissions for members of global and project
groups.
Configure Network Drive Mapping (Optional)
If you are running LabKey Server on Windows and you are connecting to a remote network share, you may need to
configure network drive mapping for LabKey Server so that LabKey Server can create the necessary service account to access the network share.
Setup for Distributed Environment
The pipeline that is installed with a standard LabKey installation runs on a single computer. Since the pipeline's search and analysis operations are resource-intensive, the standard pipeline is most useful for evaluation and small-scale experimental purposes.
For institutions performing high-throughput experiments and analyzing the resulting data, the pipeline is best run in a distributed environment, where the resource load can be shared across a set of dedicated servers. Setting up the LabKey pipeline to leverage distributed processing demands some customization as well as a high level of network and server administrative skill. If you wish to set up the LabKey pipeline for use in a distributed environment,
contact LabKey.
Related Topics
