Premium Resource - This is a legacy feature available with Premium Editions of LabKey Server.
Learn more or
contact LabKey.
The expression matrix assay ties expression-level information to sample and feature/probe information. After appropriate files are loaded into the system, users can explore microarray results by building queries and visualizations based on feature/probe properties (such as genes) and sample properties.
Expression data may be manually extracted from
Gene Expression Omnibus (GEO), transformed, and imported to LabKey Server.
Tutorial steps:
Files loaded include:
- Metadata about features/probes (typically at the plate level)
- Sample information
- Actual expression data (often called a "series matrix" file)
Review File Formats
In order to use the assay, you will need three sets of data: a run file, a sample type, and a feature annotation file.
The run file will have one column for probe ids (ID_REF) and a variable number of columns named after a sample found in your sample type. The ID_REF column in the run file will contain probe ids that will be found in your feature annotation file, under the Probe_ID column. All of the other columns in your run file will be named after samples, which must be found in your sample type.
In order to import your run data, you must first import your sample type and your feature annotation set. Your run import will fail if we are unable to find a match for your ID_REF value for a sample in your sample type. If you don't have current files, you can use these small example files:
Set Up the Folder
- Log in to your server and navigate to your "Tutorials" project. Create it if necessary.
- If you don't already have a server to work on where you can create projects, start here.
- If you don't know how to create projects and folders, review this topic.
- Create a new folder named "Expression Matrix Tutorial". Choose the folder type "Assay."
- Select (Admin) > Folder > Management and click the Folder Type tab.
- Check the box for Microarray and click Update Folder.
- Add a Sample Types web part on the left.
- Add a Feature Annotation Sets web part, also on the left.
Define Sample Type
- Click New Sample Type.
- On the Create Sample Type page, name your sample type. Here we use ExpressionMatrixSamples.
- In the Naming Pattern field, enter the following, which means use the ID_REF column as the unique identifier for samples:
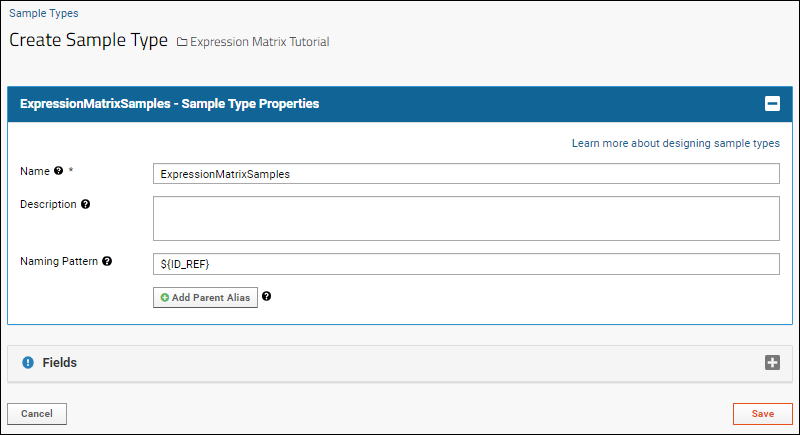
- Click the Fields section to open it.
- Click Manually Define Fields.
- Use Add Field to create all the fields that will match your spreadsheet.
- For each field enter a name without spaces and select a data type. For our sample, enter:
- ID_REF - Text
- SampleA - Integer
- SampleB - Integer

Import Sample Information
Now that you have created the new sample type, you will see the name in the
Sample Types web part.
- Click ExpressionMatrixSamples to open it.
- Click Import More Samples.
- Into the Data area, paste in a TSV of all your samples (or you can click Upload file... and upload the file directly).

Add Feature Annotation Set
- Return to the main folder pageby clicking the Expression Matrix Tutorial link near the top.
- In the Feature Annotation Sets web part, click Import Feature Annotation Set.
- Enter the Name: Feature Annotations 1
- Enter the Vendor: Vendor 1
- For Folder: Select the current folder.
- Browse to select the annotation file. (Or use the provided file sample_feature_annotation_set.txt.) These can be from any manufacturer (i.e. Illumina or Affymetrix), but must be a TSV file with the following column headers:
Probe_ID
Gene_Symbol
UniGene_ID
Gene_ID
Accession_ID
RefSeq_Protein_ID
RefSeq_Transcript_ID
- Click Upload.
Create an Expression Matrix Assay Design
- In the Assay List web part, click New Assay Design.
- Click the Specialty Assays tab.
- Under Use Instrument Specific Data Format, choose Expression Matrix.
- Select the Assay Location (for our samples, use the current folder).
- Click Choose Expression Matrix Assay.
- Name your assay, adjust any fields if needed.
- Click Save.
Import a Run
Runs will be in the TSV format and have a variable number of columns.
- The first column will always be ID_REF, which will contain a probe id that matches the Probe_ID column from your feature annotation set.
- The rest of the columns will be for samples from your imported sample type (ExpressionMatrixSamples).
An example of column headers:
ID_REF GSM280331 GSM280332 GSM280333 GSM280334 GSM280335 GSM280336 GSM280337 GSM280338 ...
An example of row data:
1007_s_at 7.1722616266753 7.3191207236008 7.32161337343459 7.31420082996567 7.13913363545954 ...
To import a run:
- Navigate to the expression matrix assay you just created. (Click the name in the Assay List from the main page of your folder.)
- Click Import Data.
- Select the appropriate Feature Annotation Set.
- Click Choose File and navigate to your series matrix file (or use the provided example file series_matrix.tsv).
- Click Save and Finish to begin the import.
Note: Importing a run may take a very long time as we are generally importing millions of rows of data. The Run Properties options include a checkbox named Import Values. If checked, the values for the run are imported normally. If unchecked, the values are not imported to the server, but links between the series matrix, samples, and annotations are preserved.
View Run Results
After the run is imported, to view the results:
- Click the Assay ID (run or file name) in the runs grid.
There is also an alternative view of the run data, which is pivoted to have a column for each sample and a row for each probe id. To view the data as a pivoted grid:
- Select (Admin) > Go to Module > Query
- Browse to assay > ExpressionMatrix > [YOUR_ASSAY_NAME] > FeatureDataBySample.
- Click View Data.
- You can add this query to the dashboard using a Query web part.
Related Topics